Research Article - (2024) Volume 3, Issue 2
Protein Structural and Sequence Analysis of Human ACE2 Using Prediction and Modeling Bioinformatics Tools for Diagnostics Biomarkers and Drug Design Features: an Opinion Study Path
2Department of Infection and Immunity, Allergy and Clinical Immunology, Clinical and Applied Virology, Luxembourg Institute of Health, Esih-sur-Alzette, Luxembourg
3Department of Biological Sciences, Ahmadu Bello University Zaria, Nigeria
4Department of Biochemistry, University of Benin, Benin city, Edo State, Nigeria, Department of Biomedical Engineering, University of Lagos, Lagos, Nigeria
5Department of Internal Medicine, Imo State University Owerri, Imo State, Nigeria; Ministry of Health, Saudi Arabia
6Department of Public Health, National Open University, Uromi, Edo State, Nigeria; Edo State University Ekpoma (Medical Laboratory Science), Nigeria
7Department of Biological Science, Abubakar Tafawa Balewa University ATBU, Bauchi, Bauchi State, Nigeria
8Center for Material Science, University of South Africa, UNISA Science Campus, South Africa
9Department of Medical Laboratory Science, Kampala International University, Kampala, Uganda
Received Date: Mar 18, 2024 / Accepted Date: Apr 08, 2024 / Published Date: Apr 22, 2024
Copyright: ©©2024 Ozurumba-Dwight Leo N, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Ozurumba-Dwight, L.N., Muller, C.P., Ndams, I.S., Avan, D.E., Enwere, O.O., et al. (2024). Protein Structural and Sequence Analysis of Human ACE2 Using Prediction and Modeling Bioinformatics Tools for Diagnostics Biomarkers and Drug Design Features: an Opinion Study Path. Curr Res Vaccines Vaccination, 3(2), 01-06.
Abstract
Angiotensin converting enzyme-2 receptor (ACE2) present on human cell membrane surfaces is a critical receptor for severe acute respiratory syndrome-coronavirus-2 (SARS-CoV-2) to bind onto and invade cells. Severe burden caused by the recent COVID-19 (coronavirus disease-2019) pandemic was of global public health and economic importance. ACE2 protein has multi-functional roles and its expression on cell membrane surfaces influences COVID-19 pathogenesis. Engagement of key bioinformatics tools that retrieves data from protein sequences can be compared with structure of ACE2 proteins from same individuals and patients of different clinical status for COVID-19 (for individuals that are non-infected, infected asymptomatic and symptomatic). This can either be from data obtained from disease states or from already deposited annotated data generated from past studies stored in protein databases. This can be geared towards investigating effectiveness of a pool of biological attributes such a structural neighbor profiles which help describe micro-environment of single amino polymorphisms (SAPs). Analysis can be done for SAPs (also known as non-synchronous single nucleotide polymorphism (nsSNPs). Then engage predictive and modeling tools to predict key mutational loci- particularly those involving SAPs and their functional significance in association with disease states in known and predicted protein sequences. Generated biological data can contribute to ontology and associated molecular pathways. Similar data can be obtained from in-vivo studies in SARS-CoV-2 infected animal models. This is an opinion study opinion that can open up paths to new designs and development protocols in therapeutics and diagnostics biomedicine. As the study moves on, based on observations in the various analyses, we expect possibilities for a few flexibilities to enhance getting optimized credible and impact oriented results.
Keywords
ACE2, Critical Receptor, Protein, Diagnostic, Structural, 13A0 Structural Neighbor Profile.
Introduction
There was a year 2002-2003 outbreak of a coronavirus scientifically named SARS-CoV (severe acute respiratory syndrome-coronavirus) and caused scare in some parts of the world, followed by entry of MERS-CoV (middle east respiratory syndrome-coronavirus) infection in 2009 but they were on each of these occasions, tamed before they could turn into global pandemic scales [12,14].
Lu et al reported in their study early in the COVID-19 pandemic period with data of 88% identity between 2019-nCoV (2019-coronavirus) genome sequence (obtained from broncho-alveolar lavage fluid isolated from 9 patients, 8 of who visited the Huanan sea food market in Wuhan China) and two bat derived coronaviruses [13]. A homology identity of 79% with its predecessor SARS-CoV coronavirus and 50% identity with MERS-CoV obviously are speculative of possibility that SARS-CoV-2 may have passed onto man from bats with an intermediate host such as sea foods as the mediator. However, if SARS-CoV-2 had been a direct evolution from its predecessor SARS-CoV, the level of mutations that evolved SARS-CoV-2 must have been notably high considering a homology identity of 79%. These are likelihoods.
Key genomic and protein structural features are avenues from which to explore options for deeper understanding of host-pathogen interactions at the fine specificities of molecular investigations [18,21,22]. This is one of the gateways to design and development of novel diagnostics and therapies. As such, the genome of SARS-CoV-2 was first sequenced in early January 2020 which invariably has been a plus to our searches for new diagnostics and therapeutics [23].
Genomics characterizations with bioinformatics platform analysis have been useful to health and biomedical sciences [7,12,17]. It has enabled specific attributes of importance to be observed such variants information to identify, track and disrupt their evolution and emergence [8]. In another study, emergence of omicron lineage in South Africa in 2021 influenced a change in character of severe acute respiratory syndrome coronavirus2 pandemic, showing elevated transmissibility and increased immune evasion [22]. Another study provided genomic sequence information and functional annotation of transcripts and proteins of I. ricinus genome with transcriptome and proteome of the unfed I. ricinus midgut, identifying putative genomes, genes and genome contigs screened for homology- annotated by mass spectrometry [5].
A study by Stewart-Jones et al evaluated domain based mRNA vaccine encoding a specific region of the spike protein receptor binding domain of SARS-CoV-2 delta virus and its omicron variants [21]. The mRNA-1283 designed has entered clinical tri8las. With the success of messenger RNA (mRNA) vaccines against coronaviruses in 2019, strategies can now focus on improving vaccine potency, breadth and stability [21,17].
About Angiotensin Converting Enzyme-2 Receptor (ACE2)
Angiotensin converting enzyme 2 receptor (ACE2) present on human cell membrane surfaces is a critical protein and receptor for SARS-CoV-2 to bind onto and gain entry into the cell [7,16]. On entry of SARSCoV-2 into the cell, ACE2 is internalized with the viral particles into endosomes and there are variants of ACE2 as they differ in how they affect given population’s susceptibility to SARS-CoV-2 [2,20].
Angiotensin converting enzyme-2 receptor (ACE2) is a critical receptor protein. Variants of ACE2 have been observed from studies which affect susceptibility to SARS-CoV-2 to depict its polymorphic nature. The discovery that genetic variations in ACE2 (particularly deleterious missense variants in its flexible regions) may affect its structure and function, and thus may affect affinity towards SARS-CoV-2 [20]. Further related to this is the emerged finding that there is an ACE2 gene which is highly polymorphic located on chromosome Xp22 [20,29]. ACE2 gene encode for ACE2 protein expression while ACE gene encode for ACE protein.
About Severe Acute Respiratory Syndrome-Coronavirus-2 (SARS-CoV-2)
SARS-CoV-2 is a single stranded RNA enveloped virus of the beta class RNA corona-virus. Studies using an RNA based metagenomic next generation sequencing platform on a strain of this virus revealed it has 29,881bp in its length [9]. Scientists in Wuhan obtained complete genome sequences from five patients infected with SARS-CoV-2 and observed that they share 75% sequence identity with SAR-CoV [23]. It indicated closeness and relationship between these viruses that plagued mankind at different periods. This is one of the evidences for the existence of SARS-CoV-2 and reality of the most recent coronavirus pandemic.
Management
Initially, treatments came through drugs originally developed and used against Pneumonia, HIV (human immunodeficiency virus) such as Azvudine, nucleotide analogues such as Remdesvir, convalescent plasma administration such as LES-untPLAZ- muh and malaria drugs for its co-management, supported with hyperbaric oxygen therapy for certain classes of severely affected patients [3,4,10,11,19,25,27]. Then onto immunotherapy and finally the successful vaccines that emerged one after another. Avenues for COVID-19 therapies have targeted various pathogenic mechanisms, namely the neutralization of ACE2 receptors or SARS-CoV-2 spike protein epitopes, and disruption of endocytic pathways, among others [10].
Justification
SARS-CoV-2 (severe acute respiratory syndrome-coronavirus-2), the causative agent of COVID-19 disease, created deep scare and threat to mankind through a pandemic which globally took over 5 million lives in death toll, commencing with the first officially recorded case infection in December, 2019 (Wuhan-Hu-1 strain), before therapeutic success started [10,12,26]. Gene sequencing techniques have helped to assemble gene and protein sequences, revealing accumulation of various types of mutations/ variants/amino acid substitutions. Their implications are subjects of continuous studies to better understand the implications, emerging trends and challenges these will pose to predicting their pathogenicity on host and use to do prediction of diseases progression and outcome. For instance, using bioinformatics and computational biology approaches, a study revealed that we can use a hybrid system of two classifiers for medical data analysis and elucidation of rare attributes of utilities for diagnosis and treatment [15].
Central Idea in this Perspective
A bioinformatics with computational biology approach as projected in this perspective will aim to utilize key molecular features such as, nearby functional Sites (NFS) and solvent accessibility from ACE2 protein sequences to obtain structural elucidations to determine their association with disease states in patients. These probes can be of diagnostic relevance.
To characterize and hypothesize variants of ACE2 protein through SNP and other mutation function(s), there are specific tools for achieving this, such as LS-SNP (large scale human SNP annotation (LS-SNP), SNP@promoter and PolyMAPr among others. Invariably, one of the most featured annotations of a SNP is identification of its location on a known or predicted protein structure.
![]() The two basic hypothesis suggested from the approach in this perspective are:
The two basic hypothesis suggested from the approach in this perspective are:
1- Clues from association between key structural and sequence features in ACE2 protein variants on cell surface of human cells of varying status of disease state can provide new avenues for diagnostics bio-markers and drug development against Sars-CoV-2 infections.
2- Using bioinformatics and computational biology platforms, we can further our knowledge on the mechanisms connecting molecular features in ACE2 protein sequence and structure with disease state (association), which when organized can contribute to our ontology and associated molecular pathways.
Associated Concept
These basic hypotheses stem from the emerging concept that rapid accumulation of single amino acid polymorphisms (SAP) (also known as non-synchronous single nucleotide polymorphism-nsSNPs), brings in avenues to explore to predict their association with specific disease conditions (28). Annotations of structure based features such as - solvent accessibility, C-beta density, number and types of variations inside and on surface of 3D protein structure, nearby functional sites (NFS) (like active site, trans-membrane regions and binding sites), number of amino acids in disordered regions in protein 3D-structure, secondary structure and other useful protein structure-function features can be determined from protein structure databases of annotated 3D-protein structures and from predicted 3D-protein structure of high quality. This is to be obtained from an inclusive approach in this presentation. This typically can use structure predictive bioinformatics tools like SIFT which uses conservation in multiple sequence alignment as sole feature and experimental mutations as its training data Single amino acid polymorphism prediction methods (SAPRED) which is a classifier based method, (http://sapred.cbi.pku.edu. cn/, http://sapred.cbi.pku.edu.cn/supp.do), PANTHER (protein analysis through evolutionary relationships), SNPs3D and Polyphen which includes protein structure data and other features (biotools:polyphen-2; http://genetics.bwh.harvard.edu/pph/). These can be used to determine association with disease severity, alongside benefits for diagnostics and disease management. Some of these structure predictive tools use experimental amino acids substitutions as training set, while others use substitution based on disease-associated human alleles.
Bioinformatics Platform Propelled Insight for this Perspective
Studies by Ye at al used support vector machine classifier (SVM) to annotate SNPs in an approach that started by compiling single amino acid polymorphism (SAP) dataset from Swiss-Prot variant pages using attributes of the wild-type and variant protein to derive several biologically informative attributes that included structural neighborhood profiles to elucidate the SAP’s micro-environment, nearby functional sites, likelihood properties of protein aggregation and disordered regions, that determines whether the SAP is located in structurally disordered regions [28]. It revealed that the 13A0 structural neighbor profile is of higher predictive power than > Nearby Functional sites which is in turn higher than > solvent accessibility in predictive power to associate disease states, with an 82.60% accuracy through a protein level 5-fold cross-validation [28]. The analysis obtained from disordered regions and aggregation properties fed into the overall accuracy of this prediction.
This Projected Study Technique Suggests this Approach
Step A Search for protein sequence and structures of ACE2 receptor of individuals that are not infected with SARS-CoV-2 (non-infected individuals) from sequence databases such as NCBI, Swiss-Prot, Protein data bank Japan (PDBj), UniProt and Conserved domains database, which all have protein molecular data unified across board.
Conduct ACE2 Protein Extraction and Purification from these Categories of Patients Outlined thus
I-From non-infected individuals and have their ACE2 protein analyzed for protein sequence, structure and functions annotation. II-From infected: individuals (patients) who have mild symptoms for COVID-19 and from those who completely recovered to have their ACE2 proteins extracted, sequenced and analyzed for their 3D structure.
III-From infected: who have severe (complicated) symptoms, and from those who completely recovered to be analyzed for their 3D structure.
IV-From infected: individuals who are asymptomatic for COVID-19 and from those who completely recovered to be analyzed for their 3D structure.
Then conduct comparative analyses of protein sequence of ACE2 proteins and 3D structure of ACE2 protein, using bioinformatics sequence and structure- multiple alignment and predictive analytical tools.
Data generated from features of sequence and 33D-structural analyses ofACE2 protein can be analyzed in relation to susceptibility to infection and associations with pathologies. Findings will have benefits for patient management, disease control and identification of key proteins (and amino acids), including features to target for drug design, development and clinical trials. Success of designed drugs often require determining optimal performance state to place the therapeutic compound, need for adjuvant or not, coupling with delivery nano-particle carriers such as liposomes, route of drug/ vaccine administration and safety levels for optimally efficient administration.
Step B Conduct and ensure ACE2 protein sequence annotation for identified features of importance in ACE2 sequence. It will be advantageous if the spike S Protein sequence and structure of SARS-CoV-2 in each of the categories of infected patients earlier listed are analyzed and kept for records. These can be analyzed for association with ACE2 binding domain of each patient and compared across other categories of patients. This will serve as an internal control to examine flexibilities or differences observed from ACE2 protein sequence analysis.
Step C Conduct and ensure ACE2 protein 3D-structure annotations for structural features of importance. Also, it will be advantageous if the spike S Protein sequence analysis of SARS-CoV-2 done for each of the categories of earlier listed patients are kept for records. These can be analyzed for association with ACE2 binding domain of each patient and compared across other categories of patients. This will serve as an internal control to examine flexibilities or differences observed from ACE2 protein structure analysis.
Step D For protein structure, the approach will include predictive modeling using bioinformatics tools such as SAPRED,, SIFT and Polyphen2, a further development of earlier version Polyphen and one other high performing structural predictive tool especially for ACE2 protein structure before viral entry using an approach in which we model the structure from its protein sequence generated from non-infected individuals or from closest homologues of the form of this protein at this stage [1]. This can be found on protein databases and compare with established ACE2 protein 3D-structure for the one found to be already deposited on protein databases.
From this point, annotations of structure based features from the 3D protein sequence structure such as - solvent accessibility; C-beta density; number and types of variations inside and on surface of 3D protein structure; nearby functional sites (NFS) (like active site, trans-membrane regions and binding sites), number of amino acids in disordered regions in protein 3D-structure; secondary structure and other useful protein structure-function features can be determined from generated 3D-protein structures of ACE2 and from the most likely structure in the predicted 3D-protein structures aided by SAPRED, SIFT and Polyphen predictive and structure modeling tools. Typically, structural neighbor profiles help describe the environment of single amino acid polymorphism’s (SAP’s) to indicate possible use for disease bio-marker. Accumulation of single amino acid polymorphisms (SAPs), also known as non-synonymous single nucleotide polymorphisms (nsSNPs), brings the opportunities and needs to understand and predict their disease association [28]. For instance, single amino acid polymorphisms (SAPs, conventionally known as non-synonymous SNPs or nsSNPs), which cause amino acid substitutions in protein sequences product account for a notable percentage of gene lesions known to be related to genetic diseases [28].
Finally, conduct structure based analysis from analyzed 3D structure using structural alignment and association tools to investigate for associations with the strong protein structural parameter of 13A0 structural neighbor profile, amino acids in disordered regions in protein, solvent accessibility; number and types of variations; nearby functional sites (NFS) parameters. This will strengthen the depth of data we generate.
Summarized Approach in a Two Streamed Path
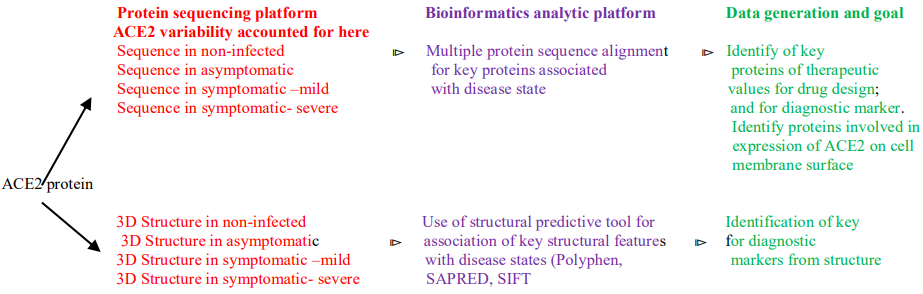

Future Direction of Study to Further the Finding
As part of other related paths to exploit, since genes normally regulate protein expression, we suggest that there are regulatory genes involved in expression of different ACE2 proteins and ACE2-compositte molecules in individuals and patients. If identification of gene regulators of ACE2 synthesis and expression of cell surface can be known, then we can from here screen, identify and develop RNA based vaccines. Different research groups have predicted the effect of various ACE2 variants on ACE2-SARSCoV-2 interaction and thus host susceptibility, with biochemical assays that confirm [20].
This perspective aims to produce an approach that opens-up some critical features through molecular predictive modeling. There is possibility of future follow-up by bio-engineering of specific molecules involved in the invasion, protein-protein interactive interfaces and conformational changes. This is directed to see the effects and areas that can be structurally harnessed and for design of new therapies against COVID-19.
Since no single study is intended as “an-only possible path to therapeutics and diagnostics, this proposed research path is not an all-inclusive in itself, but proposed paths on to a comprehensive multi-faceted approach for biomarkers and drug design and development.
This approach can possibly open up paths to insight oriented new protocols to screen for disease diagnostic makers, and to systematically screen, identify and develop new drugs to be developed and tested, of which includes vaccines. If successful, clues from this approach can be of benefit for therapeutics and bio¬markers in other diseases.
Acknowledgement
Immense gratitude to University of Manchester and its Institute of Biotechnology, Center for Bioinformatics at Peking University PKU Beijing China, The Open University of United Kingdom, UZH Zurich, Walden University MN and Luxembourg Institute of Health LIH.
Conflict of Interest
We declare none
References
- Adzhubei, I. A., Schmidt, S., Peshkin, L., Ramensky, V. E., Gerasimova, A., Bork, P., ... & Sunyaev, S. R. (2010). A method and server for predicting damaging missense mutations. Nature methods, 7(4), 248-249.
- Antony, P., & Vijayan, R. (2021). Role of SARS-CoV-2 and ACE2 variations in COVID-19. biomedical journal, 44(3), 235-244.
- Bender, A. T., Tzvetkov, E., Pereira, A., Wu, Y., Kasar, S., Przetak, M. M., ... & Okitsu, S. L. (2020). TLR7 and TLR8 differentially activate the IRF and NF-κB pathways in specific cell types to promote inflammation. Immunohorizons, 4(2), 93-107.
- Bose, S., Adapa, S., Aeddula, N. R., Roy, S., Nandikanti, D., Vupadhyayula, P. M., ... & Konala, V. M. (2020). Medical management of COVID-19: evidence and experience. Journal of clinical medicine research, 12(6), 329-343.
- Cramaro, W. J., Revets, D., Hunewald, O. E., Sinner, R., Reye, A. L., & Muller, C. P. (2015). Integration of Ixodes ricinus genome sequencing with transcriptome and proteome annotation of the naïve midgut. BMC genomics, 16, 1-15.
- Cao, B., Wang, Y., Wen, D., Liu, W., Wang, J., Fan, G., ...& Wang, C. (2020). A trial of lopinavir–ritonavir in adults hospitalized with severe Covid-19. New England journal of medicine, 382(19), 1787-1799.
- Hatmal, M. M. M., Alshaer, W., Al-Hatamleh, M. A., Hatmal, M., Smadi, O., Taha, M. O., ... & Plebanski, M. (2020). Comprehensive structural and molecular comparison of spike proteins of SARS-CoV-2, SARS-CoV and MERS-CoV, and their interactions with ACE2. Cells, 9(12), 2638.
- Hill, V., Du Plessis, L., Peacock, T. P., Aggarwal, D., Colquhoun, R., Carabelli, A. M., ... & Rambaut, A. (2022). The origins and molecular evolution of SARS-CoV-2 lineageB. 1.1. 7 in the UK. Virus Evolution, 8(2), veac080.
- Huang, Y., Yang, C., Xu, X. F., Xu, W., & Liu, S. W. (2020).Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19.Acta Pharmacologica Sinica, 41(9), 1141-1149.
- Iacob, S., & Iacob, D. G. (2020). SARS-coV-2 treatment approaches: numerous options, no certainty for a versatile virus. Frontiers in Pharmacology, 11, 557875.
- Kim, J., Jung, J., Kim, T. H., Kang, N., Choi, H., Oh, D.H., ... & Choi, J. P. (2021). Pneumonia-targeted lopinavir/ ritonavir-based treatment for patients with COVID-19: an early-period retrospective single center observational study. BMC Infectious diseases, 21, 1-8.
- Lekana-Douki, S. E., N'dilimabaka, N., Levasseur, A., Colson, P., Andeko, J. C., Zong Minko, O., ... & Lekana-Douki, J. B. (2022). Screening and whole genome sequencing of SARS-CoV-2 circulating during the first three waves of the COVID-19 pandemic in Libreville and the Haut-Ogooué Province in Gabon. Frontiers in Medicine, 9, 877391.
- Lu, R., Zhao, X., Li, J., Niu, P., Yang, B., Wu, H., ... & Tan, W.(2020). Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. The lancet, 395(10224), 565-574.
- Mostafa, A., Kandeil, A., Shehata, M., El Shesheny, R., Samy, A. M., Kayali, G., & Ali, M. A. (2020). Middle east respiratory syndrome coronavirus (mers-cov): State of the science. Microorganisms, 8(7), 991.
- Nalluri, M. R., Kannan, K., Manisha, M., & Roy, D. S. (2017). Hybrid disease diagnosis using multiobjective optimization with evolutionary parameter optimization. Journal of healthcare engineering, 2017.
- Nagesha, S. N., Ramesh, B. N., Pradeep, C., Shashidhara, K. S., Ramakrishnappa, T., Krishnaprasad, B. T., ... & Chandaragi,S. S. (2022). SARS-CoV 2 spike protein S1 subunit as an ideal target for stable vaccines: A bioinformatic study. Materials Today: Proceedings, 49, 904-912.
- Ozurumba-Dwight, L., Dalton, D., Ebenso, E., Tonukari, N., Ogbonna, C., Enwerre, O., ... & Osuya, I. (2023). Protein structural and sequence analysis of human ACE2 using prediction and modeling bioinformatics tools for diagnostics biomarkers and drug design features: an opinion study path.
- Senefeld, J. W., Franchini, M., Mengoli, C., Cruciani, M., Zani, M., Gorman, E. K., ... & Joyner, M. J. (2023). COVID-19 convalescent plasma for the treatment of immunocompromised patients: a systematic review and meta-analysis. JAMA network open, 6(1), e2250647-e2250647.
- Sodhi, P. V., Sidime, F., Tarazona, D. D., Valdivia, F., & Levano,K. S. (2022). A closer look at ACE2 signaling pathway and processing during COVID-19 infection: identifying possible targets. Vaccines, 11(1), 13.
- Stewart-Jones, G. B., Elbashir, S. M., Wu, K., Lee, D., Renzi,I., Ying, B., ... & Carfi, A. (2023). Domain-based mRNA vaccines encoding spike protein N-terminal and receptor binding domains confer protection against SARS-CoV-2. Science translational medicine, 15(713), eadf4100.
- Tsui, J. L. H., McCrone, J. T., Lambert, B., Bajaj, S., Inward,R. P., Bosetti, P., ... & Kraemer, M. U. (2023). Genomic assessment of invasion dynamics of SARS-CoV-2 Omicron BA. 1. Science, 381(6655), 336-343.
- Wang, H., Li, X., Li, T., Zhang, S., Wang, L., Wu, X., & Liu,J. (2020). The genetic sequence, origin, and diagnosis of SARS-CoV-2. European Journal of Clinical Microbiology & Infectious Diseases, 39, 1629-1635.
- Yang, Y., Peng, F., Wang, R., Guan, K., Jiang, T., Xu, G., ... & Chang, C. (2020). The deadly coronaviruses: The 2003 SARS pandemic and the 2020 novel coronavirus epidemic in China. Journal of autoimmunity, 109, 102434.
- Yang, N., & Shen, H. M. (2020). Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. International journal of biological sciences, 16(10), 1724.
- Yavarian, J., Nejati, A., Salimi, V., Shafiei Jandaghi, N. Z., Sadeghi, K., Abedi, A., ... & Mokhtari-Azad, T. (2022). Whole genome sequencing of SARS-CoV2 strains circulating in Iran during five waves of pandemic. PLoS One, 17(5), e0267847.
- Ye, Y. (2022). China approves first homegrown COVIDantiviral.
- Ye, Z. Q., Zhao, S. Q., Gao, G., Liu, X. Q., Langlois, R. E.,Lu, H., & Wei, L. (2007). Finding new structural and sequence attributes to predict possible disease association of single amino acid polymorphism (SAP). Bioinformatics, 23(12), 1444-1450.
- Zipeto, D., Palmeira, J. D. F., Argañaraz, G. A., & Argañaraz,E. R. (2020). ACE2/ADAM17/TMPRSS2 interplay may be the main risk factor for COVID-19. Frontiers in immunology, 11, 576745.