Research Article - (2022) Volume 3, Issue 3
Curcumin Induces MCF-7 Cells Pyroptosis via Autophagy/CTSB/NLRP3/ Caspase-1/GSDMD Signaling Pathway in Vitro and Vivo
2Department of Epidemiology and Health Statistics, Dalian Medical University, 9 Lvshun South Road, Dalian, 116044, PR, China
3Department of Occupational and Environmental Health, Dalian Medical University, 9 Lvshun South Road, Dalian, 116044, PR, China
4Department of Nutrition and Food Hygiene, Dalian Medical University, 9 Lvshun South Road, Dalian, 116044, PR, China
Received Date: Jul 29, 2022 / Accepted Date: Aug 04, 2022 / Published Date: Aug 18, 2022
Copyright: ©Copyright: ©2022 Xiaofeng Li, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Duan H,Jiang L, Sun X, Liu X, Yang G, Sun X, Cheng T, Ji Y, Zhang F, Du Y, Ou S, Ma R, Guan X, Teng N, Li X (2022). Curcumin Induces MCF-7 Cells Pyroptosis via Autophagy/CTSB/NLRP3/Caspase-1/GSDMD Signaling Pathway In Vitro And Vivo. J Vet heal Sci, 3(3), 250-261.
Abstract
Background: Curcumin, as a lipid-lowering drug, has been reported to be effective in the treatment of breast cancer. However, the underlying molecular mechanisms have not been completely investigated.
Methods: MTT assay was used to determine the effect of curcumin on survival rate of MCF-7 cells. The effects of curcumin on tumor growth were observed in animal models of breast cancer. The positive reactions of Caspase-1, IL-1β and IL-18 were detected by immunohistochemistry. LC3, p62, CTSB, ASC, Pro-Caspase-1, GSDMD, NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 were determined by Western blot in vitro and vivo. ELISA determined the release of extracellular IL-1β and IL-18. LDH release was measured. The expression level of CTSB in cytoplasm were determined by immunofluorescence assay. Cell proliferation, cell migration and tube formation assays were used to determine the abilities of cells. In this study, NLRP3 inflammasome inhibitor MCC950, cathepsin B inhibitor CA-074 ME and autophagy inhibitor 3-MA were used to act on cells to investigate the role of NLRP3 inflammasome, cathepsin B and autophagy in curcumin-induced pyro ptosis of MCF-7 breast cancer cells.
Results: In mouse model of breast cancer, we observed that curcumin treatment significantly induced cell autophagy and pyro ptosis. In human breast cancer MCF-7 cells, we found that curcumin induced pyrophoric cell death was dependent on the activation of NLRP3/Caspase-1/GSDMD signaling pathway, which was CTSB-dependent. In addition, curcumin-induced cell autophagy caused lysosomal rupture and CTSB release. Furthermore, NLRP3 inhibitor (MCC950) significantly suppressed curcumin-induced pyro ptosis, as well as CTSB inhibitor (CA074 Me) and autophagy inhibitor (3-MA). Besides, we also found that curcumin suppressed cell proliferation, cell migration and tube formation, which could be reversed by inhibitors.
Conclusions: In summary, our results demonstrated that curcumin induced MCF-7 cell pyro ptosis by the activation of autophagy/CTSB/NLRP3/Caspase-1/GSDMD signaling pathway. These findings offer novel insights into the potential molecular mechanisms of curcumin in treatment of breast cancer.
Keywords
Curcumin MCF-7 Pyro ptosis GSDMD NLRP3 Autophagy
Introduction
Breast cancer is the most common malignancy in females and ranks second among causes for cancer related death in women [1]. Lifestyle is considered an increasingly significant contribut-ing factor to breast cancer etiology. Obesity, overweight, hyper-cholesterolemia, metabolic syndrome and alcohol represent risk factors for breast cancer [2]. Chemotherapy has been used as a routine treatment for breast cancer and other cancers. However, chemotherapy is prone to drug resistance and has high side ef¬fects. For many years, clinicians and researchers have been ex¬ploring and examining various therapeutic modalities for breast cancer. Finding natural anticancer compounds with low toxicity and high selectivity has always been the mainstream direction of cancer research.
Curcumin, a lipid-lowering drug is the main component of tur-meric, derived from the roots of plant Curcuma longa [3]. It has been widely studied for its anti-cancer, antioxidant, anti-inflam-matory, anti-angiogenic and wound healing effects for its me¬dicinal properties in Chinese and Indian systems of medicine [4]. And the main mechanisms of action by which curcumin ex¬hibits its unique anticancer activity include inducing apoptosis and inhibiting tumor proliferation and invasion by suppressing various cellular signaling pathways [5]. Several studies reported that curcumin may regulate multiple signaling pathways, includ¬ing PI3K/AKT, nuclear factor (NF)-κB, MAPK and JAK/STAT [6]. Recent studies have shown that curcumin can also inhibit the phosphorylation of protein kinase B (Akt)/mammalian target of rapamycin (mTOR), decreased B-cell lymphoma 2 (BCL2) and promoted BCL-2-associated X protein (BAX) and cleavage of caspase 3, subsequently inducing apoptosis of breast cancer cells [7]. However, the mechanism of curcumin in breast cancer has not been totally investigated and lack of in vivo experiments.
Pyro ptosis, a proinflammatory form of regulated cell death that depends on the enzymatic activity of inflammatory proteases belong to the caspase family, especially caspase-1 [8]. Plas¬ma membrane rupture, cytoplasmic swelling, DNA cleavage, NLRP3 inflammasome activation and release of proinflammato¬ry cell contents generally accompany Pyroptosis [9]. Addition¬ally, it needs lipopolysaccharide (LPS) to upregulate, NLRP3 and pro-IL-1β are also essential. Inflammasomes are multi-pro-tein signaling complexes that trigger the activation of inflam¬matory caspases and the maturation of IL-1β [10]. Nucleotide oligomerization domain-like receptor proteins (NLRPs), espe¬cially NLRP3, interact with ASC and pro-caspase-1 to active caspase-1. Caspase-1 is activated within the inflammasome, and active caspase-1 processes adermin D (GSDMD) and cytokines such as pro- IL-1β and pro- IL-18. Upon permeabilization of the plasma membrane by GSDMD pores, cells undergo a lytic, pro-inflammatory cell death (pyro ptosis) that promotes the release of mature IL-1β and IL-18 [11]. Currently, it has been reported that the release of lysosomal cathepsin B induces the activation of NLRP3 inflammasome [12]. Cathepsin B (CTSB) is an in¬tracellular cysteine protease, mainly localized in the lysosome [13]. In addition, the level of CTSB in cytoplasm is related to autophagy flux. When autophagy level increases, autophagic ly¬sosomal degradation leads to cytoplasmic release of CTSB that related to activation of NLRP3 inflammasomes [14-16].
Autophagy, a lysosomal dependent catabolism process by eu¬karyotic cells degraded longevity proteins and their organelles, and is involved in the development, differentiation and homeo¬stasis of cells under various physiological and pathological conditions [17]. Autophagy plays a key role in cancer. In recent years, few studies have indicated that curcumin can induces au¬tophagy [18-20].
In our study, we hypothesized that curcumin might induce au¬tophagy, the cytosolic release of lysosomal contents (CTSB), activation of NLRP3 inflammasome and Caspase-1 and GSD¬MD-dependent pyro ptosis.
Materials & Methods
Cell Culture and Treatment
The MCF-7 cells were obtained from the ATCC cell bank. MCF-7 cells were cultured in DMEM medium (Gibco, 1853128) sup¬plemented with 10% fetal bovine serum (Biological Industries, 1707254) 1% penicillin/streptomycin in a humidified 5% CO2 at 37 °C incubator. Cells were passaged every 3 days at desired confluence. Curcumin was purchased from Sigma Aldrich (CAS No.458-37-7, assay≥94%) and dissolved in DMSO to make a stock solution of 50mM. MCF-7 cells were pretreated with 0.2μg/ml lipopolysaccharide (LPS, Sigma Aldrich, L4391) for 4h and then expose to curcumin (8μM) for 48h.This experiment involved pharmacological inhibitors. MCF-7 cells were pre¬treated with the addition of autophagy inhibitor 3-methylade-nine (3-MA, 5mM, Sigma, assay 98%), CTSB inhibitor CA-074 Me (10μM, AdooQ, A13256, assay>98%), NLRP3 inhibitor MCC950 (5Mm, MCE, HY-12815A, assay>98%) and 0.2μg/ml LPS for 4h, then to expose to curcumin (8μM) for 48h.
Animal Experiment
Ten six-year-old female BALB/c-Nude mice (weight: 20g) were purchased from Beijing Vital River, raised in the Institute of Ge¬nome Engineered Animal Models for Human Disease of Dalian Medical University (SPF level). The mice were kept under ster¬ile conditions and fed a sterilized mouse diet and water. All mice were anaesthetized via inhalation of isoflurane and a tumor cell suspension of 107 MCF-7 cells in 0.2 ml DMEM was injected subcutaneously into the inguinal of each mouse. When tumors reached about 30-60mm3 at 1 week, the mice were randomly separated into two groups (n = 5/group). The mice were treated with curcumin 200 ug/kg or saline (control) by intraperitone-al injections every day for 4 weeks. Tumor size was measured every week in two perpendicular dimensions with Vernier cal-ipers and converted to tumor volume using the formula: a*b*b (a: longer, b: shorter). At the end of the experimental period, all mice were euthanized and tumors were segregated and weighed.
Cell Viability Assays
The cytotoxicity of curcumin was measured by using the 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) assay. MCF-7 cells (5*104/ml) were seeded in 96-well plates and treated with 0,2,4,8,16,32,64,128μM for 24h or 48h.After treatment, MTT (5 mg/ml in sterile PBS, Solar bio, M8180) was added and the 96-well plates were incubated for 4h at 37â??.Thereafter the supernatant was removed and 100μl DMSO was added into each well for 30 min at 37â??.After that, the plates were shaken carefully until the blue formazan crystals were fully dissolved. Then measured the absorbance at 570 nm using a Bio-Rad Microplate Reader, and the cell viability (%) was calculated using the formula: (A570 of treated group/A570 of control) × 100%.
Western Blot Analysis
Proteins were extracted from the MCF-7 cells or from the tu-mors, completely lysed in the lysis buffer by using a nuclear and cytoplasmic protein extraction kit (Keygen Biotech, Chi-na). Then quantified with the BCA Protein Assay kit (Thermos, MK164230). An equal amount (30μg) of total proteins were loaded into different lanes and separated on 10%-15% SDS-Page gel, then blotted and transferred onto polyvinylidene di-fluoride (PVDF) membranes. Membranes were blocked for 1h with 10% skimmed milk by gentle shaking in a water bath shaker at 37 °C and then incubated with primary antibody LC3 (Abcam, ab192890, 1:1000), p62 (Protein tach, 18420-1-AP, 1:1000), CTSB (Wanleibio, WL01089, 1:500), ASC (Wanleibio, WL02462, 1:500), pro-Caspase-1 (Wanleibio, WL02996,1:500), GSDMD (Cell Signaling Technology, #93709, 1:500), NLRP3 (Wanleibio, WL02635, 1:1000), Caspase-1 (Affinity, AF4005, 1:500), GSDMD-N(Abcam, ab215203), IL-1β (Wanleibio, WL00891, 1:500), IL-18 (Wanleibio, WL01127, 1:1000), β-ac- tin (Cell Signaling Technology, #3700, 1:1000) at 4â?? overnight. After washing 3 times with PBS, membranes were incubated with the Secondary Goat anti-mouse IgG and Goat anti-rab-bit-IgG at room temperature for 1.5h. The bound antibody was visualized using the Super Signal West Pico Kit (Thermo Scien¬tific) and Bio-Rad ChemiDoc™ MP imaging system (Bio-Rad Laboratories), then analyzed by Image software. The experi¬ments above were independently repeated three times.
Immunohistochemically Assays
The tissue samples of the tumors, preserved in 2.5% glutaralde-hyde-polyoxymethylene solution, were dehydrated and embed¬ded in paraffin following routine methods. The paraffin sections were dewaxed and hydrated following routine methods. Rinsed the paraffin sections in PBS-T (3×5 min before each following steps), and then blocked with 3% peroxide-methanol at room temperature for endogenous peroxidase ablation. Afterwards, the sections were immersed in a boiling sodium citrate buffer for 15min, cooled down to room temperature. After that, incubated with blocking buffer (normal goat serum at room temperature for 20 min. Then incubated with primary antibody Caspase1 (Affinity, AF4005, 1:100), IL-1β (Wanleibio, WL00891, 1:100), IL-18 (Wanleibio, WL01127, 1:100) at 4â?? overnight. After that, the sections were incubated with the Secondary Goat anti-rab-bit-IgG at 37â?? for 30min.Then incubated with the S-A/ HRP at 37°C for 30 min. Colored with 3,3-diaminobenzidin (DAB), and kept at room temperature without light for 10 min. After rinsing adequately with water, the sections were stained with hematox-ylin for 5s, then dehydrated and sealed with neutral resins. We observed the sections under an upright microscope.
Immunofluorescence Staining
In order to observe distribution of CTSB in cytoplasm, MCF-7 cells were incubated on slides and treated with primary rab¬bit monoclonal antibodies against CTSB (Abcam, ab125067, 1:200) after fixation and blocking. Lysosomes were labeled with Lysotracker Red (Beyotime) and Alexa Fluor 488 secondary an¬tibodies (Proteintech, SA00006-2, 1:400) was added in the dark. We analyzed the 4, 6-diamidino-2-phenylindole (DAPI) coun-terstained slides under a fluorescence microscope (40 × 10).
Lactate Dehydrogenase (LDH) Release Assays
To access the toxic effect of curcumin, the LDH release of MCF-7 cells were measured using an LDH Cytotoxicity Assay Kit (Be-yotime, C0016). MCF-7 cells were seeded in 96 well plates to desired confluence and treated with inhibitor for 4h then treated with curcumin for 48h. One hour before the end of the treatment, we added 10% of the LDH release reagent and the original cul¬ture volume when the sample showed maximum enzyme activ¬ity. The culture supernatants were collected, and the absorbance was read at 490 nm with a microplate reader (Thermos Fisher Scientific). The percentage of LDH released was calculated as the percentage of the total release amount and considered to be the sum of the enzyme activity in the cell lysate and the enzyme activity in the medium.
Enzyme-Linked Immunosorbent Assays (ELISA)
IL-1β and IL-18 levels were measured by using ELISA kits (Lengton, BPE10083, BPE10092) according to the manufactur-er’s instructions. MCF-7 cells were seeded in 96-well plates to desired confluence and treated with inhibitor for 4h then treat¬ed with curcumin for 48h. The 96-well plates were centrifuged at 3000rpm for 20 min at 4 °C. Finally, the supernatants were collected. A total of 50μl of serially diluted samples and stan¬dard were added to the ELISA plate wells and incubated with horseradish peroxidase-conjugated specific antibodies for IL-1β and IL-18 for 60 min at 37 °C. The OD values were detect¬ed at 450nm by using microplate reader. The linear regression equation of the standard curve was calculated according to the concentration of the standard corresponding OD value. Finally, the sample concentration was calculated on the linear regression equation according to its OD value.
Cell Migration Assays
For the Trans well migration assays, MCF-7 cells were treated with various treatments. After indicated treatment, cells were suspended at serum-free medium then seeded onto the upper chambers with non-coated membrane (24-well insert; 8-mm pore size). After that, we filled the lower chambers with 600ml DMEM containing 10% FBS. After 24 hours of incubation, non-invading cells on the upper surface of the upper chamber were utterly removed by using cotton swabs. Other cells on the lower surface of filters were fixed with methanol for 30 min then stained with 0.1% crystal violet for 30 min. The number of in¬vaded cells was counted under an upright microscope.
Materiel Tube Formation Assay
Via a precooled pipette, 50 µL of freeze thawing liquid Materiel (Corning, 354248, USA) was embedded into a 96-well plate at 4 °C. HUVECs (2 × 104 cells per well) were seeded onto the solidified materiel 96-well plate and cultured for 12 h at 37°C in 5% CO2. After the HUVECs were overlaid, we replaced the culture medium of HUVECs with MCF-7 cell culture medium treated by curcumin and inhibitor. Capillary-like structures were evident and counted using a phase-contrast microscope and the networks formed by HUVECs were quantified with Image.
Plate Clone Formation Assay
MCF-7 cells were cultured and seeded onto 6-well plates (1000 cells per well) to prepare the cells for the plate clone formation assay. Different wells were treated variously. When the cells in 6-well plates growth to a density of 50% per cluster, which were washed with the phosphate-buffered saline (PBS) and fixed by 4% paraformaldehyde (Sigma-Aldrich, St. Louis, MO, USA). Giemsa stain (Sigma-Aldrich) was used to stain the cells. Even-tually, cell clones were counted and analyzed.
Statistical Analysis
All Data were expressed as means ± standard deviation (SD) from at least three independent experiments performed in trip-licates and analyzed by using the SPSS 20.0. Significance was determined using one-way analysis of variance (ANOVA) or t-test, and P value < 0.05 was considered statistically significant
Result
Curcumin Inhibited Proliferation and Induced Cell Death in MCF-7 Breast Cancer Cells
In order to assess the effects of curcumin on proliferation and cell death of MCF-7 cells, we performed in vitro assays. As shown in Fig. 1, the cell viability of MCF-7 cells was observed after incubation with different concentrations (0, 1, 2, 4, 8, 16, 32, 64, 128μM) of curcumin for 48h. At the concentrations above 8μM, curcumin restrained the growth of MCF-7 cells signifi¬cantly (P<0.05).
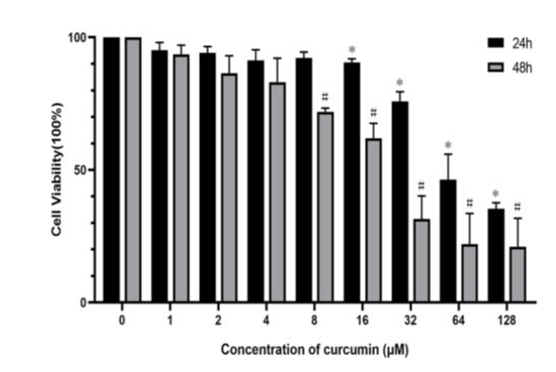
Figure 1: Effects of curcumin on cell viability in MCF-7 cells. Cells were treated with different concentrations of curcumin (from 0μM to 128μM) for 24 and 48 h. Cell viability was detected by MTT assays. Values were presented as the mean ± SEM of triplicate cell stimulation. (*P < 0.05 versus the 24h control group; #P < 0.05 versus the 48h control group)
Curcumin Inhibited the Tumor Growth in Mouse Mod¬el of Breast Cancer
In vivo assays, we established a mouse model of breast cancer, to comprehend the role of curcumin on tumors’ growth. Results obtained by detection of mice body indicated that curcumin has no toxicity effect on mice. As shown in Fig. 2, tumors’ volumes decreased significantly (P<0.05) after 4 weeks of curcumin treatment as compared with control.
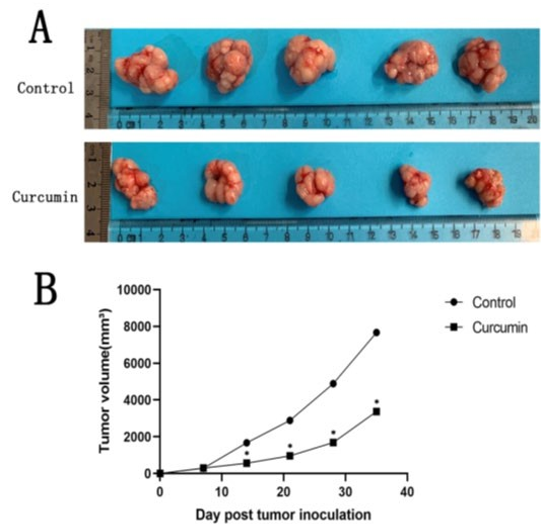
Figure 2: The condition and growth curve of tumours. (A) Image of MCF-7 breast cancer tumours. (B) The growth curve of the tumour. (*P < 0.05 versus the control group)
Curcumin Induced Pyro ptosis in Vitro and Vivo
To confirm the effect of curcumin in tumor, protein expression levels of LC3, p62, CTSB, ASC, pro-Caspase-1, GSDMD, NLRP3, Caspase-1, GSDMD-N, mature IL-1β and IL-18 were measured. As shown in Fig. 3AB, curcumin treatment (200ug/ kg) significantly (P<0.05) decreased the expression of p62 and increased the expression of LC3, CTSB, ASC, pro-Caspase-1, GSDMD, NLRP3, Caspase-1, GSDMD-N, mature IL-1β, ma¬ture IL-18 in tumors compared with control. Furthermore, re¬sult of immunohistochemically illustrated that tumors treated with curcumin displayed an increase in positive staining of Caspase-1, mature IL-1β and IL-18 compared with control (Fig. 3CD). In cell experiments (Fig. 4) we found that curcumin treat-ment(0~128μM) similarly decreased the expression of p62 and increased the expression of LC3, CTSB, ASC, pro-Caspase-1, GSDMD, NLRP3, Caspase-1, GSDMD-N, mature IL-1β and IL-18 compared with control in LPS-primed cells, with a dose-dependent manner. Notably, there was no significant dif¬ference between the LPS group and the control group. Together, these results suggest that curcumin is able to induce pyro ptosis in MCF-7 cells and tumors.
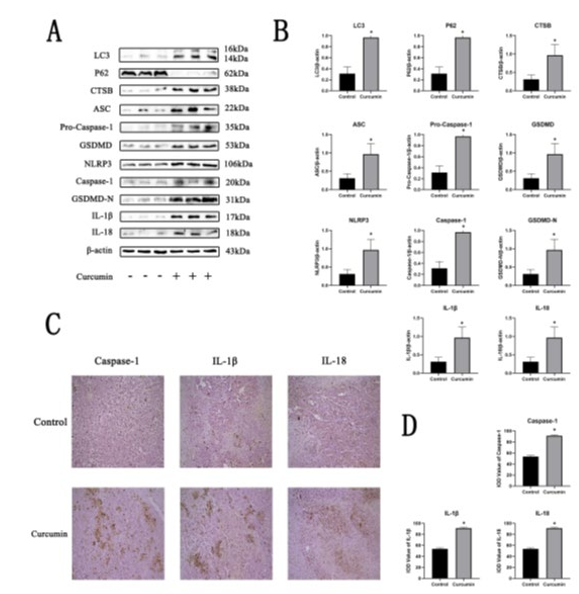
Figure 3: Curcumin induced tumour pyro ptosis in vivo. (A)(B) The protein expressions of LC3, P62, CTSB, ASC, Pro- Caspase-1, GSDMD, NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 in different treated tumour groups were detected by Western blotting. (C)(D) The positive reactions of Caspase-1, IL-1β and IL-18 in different tumour groups were detected by immunohistochemistry. (*P < 0.05 versus the control group
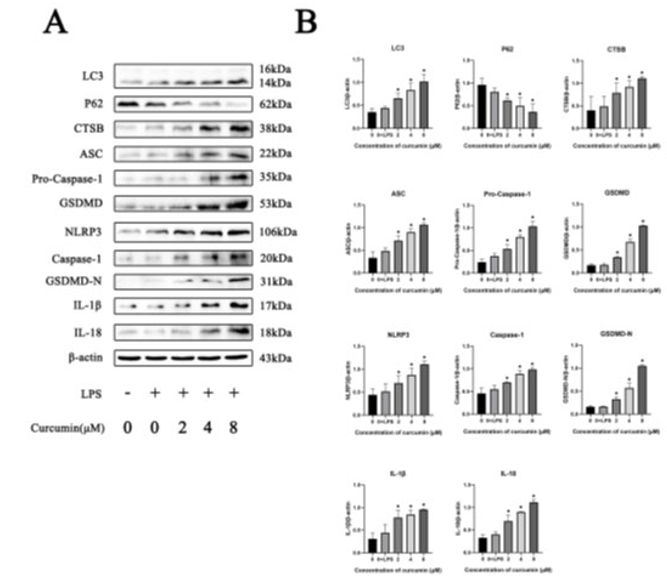
Figure 4: Curcumin induced cells pyro ptosis in MCF-7 cells. (A)(B) After pre-treated with or without 1 μg/ml LPS for 4 h and treated with different concentrations of curcumin for 48 h (from 0 μM to 8 μM), protein expression of LC3, P62, CTSB, ASC, Pro-Caspase-1, GSDMD, NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 were analysed by western blotting. (*P < 0.05 versus the control group
Curcumin-Induced MCF-7 Pyroptosis depends on Ac- tivation of the NLRP3 Inflammasome
To determine the role of NLRP3 inflammasome components in activation of caspase-1 under curcumin stress, we inhibited the expression of NLRP3 using NLRP3-specific inhibitor MCC950. The MCF-7 cells were pretreated with MCC950 (5 μM) and LPS (1 μg/ml) prior to treatment with 8 μM curcumin for 48 h. Be¬fore that, we detected the effect of curcumin on the viability of MCF-7 cells. As shown in Fig. 5AB, NLRP3, Caspase-1, GSD-MD-N, mature IL-1β and IL-18 expression were increased by curcumin treatment. However, MCC950 restrained Caspase-1, GSDMD-N expression and IL-1β, IL-18 production. Moreover, MCC950 inhibited the release of mature IL-1β, IL-18 and LDH caused by curcumin (Fig. 5CD). Besides, in functional experi¬ments, MCC950 reversed the amount of MCF-7 cell migration decreased by curcumin treatment (Fig. 5E), as well as the ma¬teriel tube formation (Fig. 5F) and plate clone formation (Fig. 5G). In all, these findings indicated that curcumin-induced pyro ptosis is dependent on NLRP3 inflammasome activation.
Figure 5: Curcumin-induced pyro ptosis is NLRP3 inflammasome-dependent in MCF-7 cells. (A)(B) MCF-7 cells were pre-treated with 1 μg/ml LPS and MCC950 (5 μM) for 4 h, and the expression of NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 were analysed by Western blotting. (C) IL-1β and IL-18 levels from different treated groups of supernatants were measured by using ELISA kits. (D) The LDH release of different treated groups of MCF-7 cells were measured by using an LDH Cytotoxicity Assay Kit. (E) Ability of migration of MCF-7 cells was detected after the exposure to MCC950 (5 μM) and curcumin (8 μM). (F) Ability of tube formation of MCF-7 cells was detected after the exposure to MCC950 (5 μM) and curcumin (8 μM). (G) Ability of plate clone formation of MCF-7 cells was detected after the exposure to MCC950 (5 μM) and curcumin (8 μM). (*P < 0.05 versus the control group; #P < 0.05 versus the 8 μM group)
Curcumin-Induced NLRP3 Inflammasome Activation and Pyro Ptosis Are Mediated By Cytoplasmic CTSB
It has been indicated that NLRP3 inflammasome activation was associated with a variety of upstream factors, including the re¬lease of lysosomal CTSB. As shown, CTSB expression was up regulated in cells and tumors after curcumin treated. Therefore, we pretreated cells with CA-074 Me, an inhibitor of CTSB, to measure the role of CTSB in curcumin-induced activation of the NLRP3 inflammasome. As shown in Fig. 6AB, the increased ex¬pression level of CTSB, NLRP3, Caspase-1, GSDMD-N, mature IL-β, IL-18 proteins caused by curcumin were inhibited by CA-074 Me compared with control. Besides, CA-074 Me inhibited the release of mature IL-1β, IL-18 and LDH caused by curcumin (Fig. 6CD). Moreover, in functional experiments, CA-074 Me reversed the amount of MCF-7 cell migration decreased by cur¬cumin treatment (Fig. 6E), as well as the materiel tube formation (Fig. 6F) and plate clone formation (Fig. 6G). As observed by microscopy, immunofluorescence results showed that curcumin treatment could cause MCF-7 intracellular lysosomes to rupture and release CTSB into the cytoplasm, and CA074 Me treatment could significantly reverse this situation (Fig. 6H). Together, the results indicated that curcumin-induced NLRP3 inflammasome activation and cell pyro ptosis was CTSB-dependent.
Figure 6: Curcumin-stimulated NLRP3 inflammasome activation and pyro ptosis are mediated by CTSB. (A)(B) MCF-7 cells were pre-treated with 1 μg/ml LPS and CA074 Me (10 μM) for 4 h, and the expression of CTSB, NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 were analysed by Western blotting. (C) IL-1β and IL- 18 levels from different treated groups of supernatants were measured by using ELISA kits. (D) The LDH release of different treated groups of MCF-7 cells were measured by using an LDH Cytotoxicity Assay Kit. (E) Ability of migration of MCF-7 cells was detected after the exposure to CA074 Me (10 μM) and curcumin (8 μM). (F) Ability of tube formation of MCF-7 cells was detected after the exposure to CA074 Me (10 μM) and curcumin (8 μM). (G) Ability of plate clone formation of MCF-7 cells was detected after the exposure to CA074 Me (10 μM) and curcumin (8 μM). (H) Detection of cytosolic and lysosomal CTSB levels after pre-treatment with CA074 Me (10 μM) for 4 h and treatment with curcumin (8 μM) for 48 h. (*P < 0.05 versus the control group; #P < 0.05 versus the 8 μM group)
Curcumin Induced Up-Regulation of MCF-7 Autophagy Then Induced the Release of CTSB, the Activation of NLRP3 Inflammasome and Pyro Ptosis through Au¬tophagy
The increase in cytosolic CTSB levels is associated with lyso-somal degradation. The results show an increase in autophag¬ic flux in MCF-7 cells. To confirm the change of autophagic flux, we used the autophagy inhibitor 3-MA in MCF-7 cells. As shown in Fig. 7AB, the up-regulation of LC3 and the downreg-ulation of p62 by curcumin were reversed by 3-MA since 3-MA inhibited the fusion of auto phagosome and autolysosome. The protein LC3 could be transformed from a soluble form (LC3-I) to a lipidzed form (LC3-II) when autophagy was activated. In addition, p62 is a multifunctional protein regulated by the bal¬ance between its transcriptional regulation and post-translation- al autophagy degradation [21]. The LC3 conversion and p62 protein are regard as the most credible biochemical markers of autophagy [22]. These results suggested that curcumin induced up-regulation of MCF-7 autophagy. In addition, the expression of cytosolic CTSB, NLRP3, Caspase-1, GSDMD-N, mature IL-1β, IL-18 were up regulated by curcumin and down regulated by 3-MA (Fig. 7AB). Moreover, 3-MA inhibited the release of ma¬ture IL-1β, IL-18 and LDH caused by curcumin (Fig. 7CD). Fur-thermore, in functional experiments, 3-MA reversed the amount of MCF-7 cell migration decreased by curcumin treatment (Fig. 7E), as well as the materiel tube formation (Fig. 7F) and plate clone formation (Fig. 7G). These results demonstrated that cur¬cumin induced up-regulation of MCF-7 autophagy, which was associated with the increase in cytosolic CTSB, activation of NLRP3 inflammasome and pyrophoric cell death.
Figure 7: Curcumin induced autophagy in MCF-7 cells. (A)(B) MCF-7 cells were pre-treated with 1 μg/ml LPS and 3- MA (5 μM) for 4 h, and the expression of NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 were analysed by Western blotting. (C) IL-1β and IL-18 levels from different treated groups of supernatants were measured by using ELISA kits. (D) The LDH release of different treated groups of MCF-7 cells were measured by using an LDH Cytotoxicity Assay Kit. (E) Ability of migration of MCF-7 cells was detected after the exposure to 3-MA (5 μM) and curcumin (8 μM). (F) Ability of tube formation of MCF-7 cells was detected after the exposure to 3-MA (5 μM) and curcumin (8 μM). (G) Ability of plate clone formation of MCF-7 cells was detected after the exposure to 3-MA (5 μM) and curcumin (8 μM). (*P < 0.05 versus the control group; #P < 0.05 versus the 8 μM group)
Discussion
The breast cancer has still been the most common cancer among women worldwide [23]. Recently study indicated that it has a younger trend [24]. Curcumin, a polyphenol extracted from Cur-cuma longa in 1815, has gained attention from scientists world¬wide for its anticancer potential [25]. The phosphoinositide 3-kinase (PI3K)/Akt signaling pathway has always been a focus of interest in breast cancer. However, deregulation of the PI3K/ Akt signaling pathway including PIK3CA activating mutation is frequently present in breast cancer [26]. Besides, the mecha¬nism of curcumin in the treatment of breast cancer has not been completely investigated. In the present study, we investigated the effect on pyro ptosis in curcumin-treated cells. It is worth mentioning that our study provided the first evidence for cur¬cumin induce cell pyro ptosis by activating autophagy/NLRP3/ Caspase-1/GSDMD pathway in MCF-7 breast cancer cells. Our study demonstrated that curcumin upregulated the level of auto¬phagy and triggered NLRP3 inflammasome activation, leading to pyrophoric cell death. In addition, curcumin-induced pyro-phoric cell death dependent upon the CTSB-mediated activation of NLRP3 inflammasome and curcumin-induced autophagy was implicated in the curcumin-induced release of CTSB.
To date, pyro ptosis is a type of programmed cell death mediat¬ed by the formation of plasma membrane pores by members of the adermin protein family, characterized by the swelling and lysis of cells, and release of many proinflammatory factors [27]. Caspase-1, as a statically determinate proenzyme, is an import¬ant component of the inflammasome and is formed by different PRRs through (or without) ASC and Caspase1 under the stimu¬lation of specific PAMPs and DAMPs [28]. The inflammasome contains an NLR and adaptor protein ASC, which interacts with procaspase-1 through caspase recruitment domain, and then self-cleavage forms active caspase-1. Caspase-1 mediated the maturation and secretion of proinflammatory (IL-1β and IL-18) and lead to pyro ptosis [29]. Adermin D (GSDMD) is a direct substrate of caspase-1, which can be specially cleaved by in¬flammatory caspases and plays a key role in the downstream of inflammatory caspases in pyro ptosis [30]. Inflammatory caspases cleaved GSDMD to form GSDMD-N, which is capa-ble of forming pore like structures in lipid membranes and thus constitutes the direct and unique effector of pyro ptosis [31]. GSDMD-N forms pores in the plasma membrane leading to membrane defects and cytoplasmic protein release [32]. In our study, treatment with curcumin elevated the levels of Caspase-1, GSDMD-N, IL-1β, IL-18 both in vitro and in vivo. Furthermore, the immunohistochemically results of Caspase-1, IL-1β and IL-18 in the tumor tissues of mice were significantly positive compared with control. In addition, the release of IL-1β, IL-18 and LDH in the cytoplasm were also increased. These results demonstrated that curcumin induced Caspase-1 and GSDMD dependent pyro ptosis in MCF-7 breast cancer cells.
Inflammation is involved in the development and progression of tumors, as well as in the antitumor response to treatment. Among the inflammasomes family, NLRP3 inflammasome is the most characterized. As we known that NLRP3 is deubiq-uitinated and associates to ASC, which then associates to pro- caspase-1 to form a large multimedia polyprotein complex. Pro-caspase-1 undergoes auto proteolytic cleavage, possibly due to proximal-induced multimerization, resulting in the active form of caspase-1 [33]. The activation of NLRP3 inflammasome in¬duces two primary effects, programmed cell death known as pyro ptosis, and/or proinflammatory responses caused by the release of inflammatory cytokines IL-1βand IL-18 [34]. Previ¬ous studies have found that curcumin induced NLRP3 inflam-masome priming and caspase-1 activation [35]. Interestingly, however, few studies have shown that curcumin suppresses IL-1β secretion through Inhibition of the NLRP3 Inflammasome [36]. In our study, we reported that treatment with curcumin el¬evated the levels of NLRP3, Caspase-1, IL-1β, IL-18, GSDMD and GSDMD-N, which was reversed using the NLRP3 inhibitor MCC950 both in vitro and in vivo. These results indicated that curcumin caused activation of the NLRP3 inflammasome and lead to pyrophoric cell death finally.
Previous study confirmed that three signaling pathways of the NLRP3 inflammasome activation involving potassium efflux, generation of reactive oxygen species, and cathepsin B release [37]. Cathepsin B is a cysteine protease involved in the regula¬tion of metalloproteinase, intracellular communication, autoph¬agy induction and immune resistance, and plays an important role in cancer progression and anticancer therapy [38]. Recent¬ly study indicated that cathepsin B was required for caspase-1 activation induced by many different NLRP3 inflammasome activators [39]. In our study, the similar results were observed. Treatment with curcumin elevated the levels of CTSB, NLRP3, Caspase-1, GSDMD-N, IL-1β and IL-18 both in vitro and in vivo that could reverse by the cathepsin B inhibitor CA074-Me. Furthermore, the result also showed that CA074 ME reduced cy¬toplasmic cathepsin B in immunofluorescence assay. Consistent with others these results demonstrated that the activation of the NLRP3 inflammasome was mediated by cathepsin B [40].
Autophagy is a process by which proteins and organelles are phagocytosed in autophagosomal vesicles and delivered to the lysosome/vacuole for degradation [41]. Autophagy is a tightly coordinated process that isolates misfolded proteins, damaged or aged organelles, and mutated proteins in double membrane vesicles called auto phagosomes that eventually fuse into lyso¬somes, leading to the degradation of the isolated components [42]. During autophagy, cytoplasmic LC3 protein is processed and recruited to the autophagosomal membranes; the auto pha¬gosome then fuses with the lysosome to form the autolysosome. Cytoplasmic LC3-I binds to phosphatidylethanolamine to form LC3-II. LC3-II was then incorporated into the auto phagosome membrane [43]. Ubiquitin-associated protein p62 is a classic autophagy receptor that binds to LC3 and recruits the select¬ed cargo to mature auto phagosomes. A lapidated form of LC3, LC3-II, p62 has been shown to be an autophagosomal marker in mammals [44]. In our study, treatment with curcumin elevat¬ed the levels of LC3, CTSB, NLRP3, Caspase-1, GSDMD-N, IL-1β, IL-18 and decreased the level of p62 both in vitro and in vivo, indicating that the degradation of auto phagosomes leads to the release of CTSB into the cytoplasm to activate the NLRP3 inflammasome. Besides, the increased level of CTSB subsequent activation of NLRP3 inflammasome and Caspase-1 and GSDMD-dependent pyrophoric cell death induced by cur¬cumin were reversed by the autophagy inhibitor 3-MA obvious¬ly. These results are consistent with the recent studies [45, 46]. Notably, curcumin did not increase NLRP3 protein expression after treatment with the autophagy inhibitor 3-MA, suggested that curcumin could not directly induce the activation of NLRP3 inflammasome, which was consistent with the anti-inflammato-ry properties of curcumin. Taken together, these results demon¬strate that curcumin induces autophagy in MCF-7 breast cancer cells and then activates the NLRP3 inflammasome.
In conclusion, our study indicates that curcumin activates the NLRP3 inflammasome by inducing autophagy in MCF-7 breast cancer cells, thus causing pyrophoric cell death via Autophagy/ CTSB/NLRP3/Caspase-1/GSDMD signaling pathway. We also demonstrate that the molecular mechanism of NLRP3 inflam¬masome activation underlies the autophagy and the release of CTSB from autolysosome caused by curcumin treatment. In addition, this study provides a new perspective on curcumin in the treatment of breast cancer. This knowledge may complement the treatment strategy for breast cancer and further confirm the bright future of curcumin in cancer treatment.
Conclusion
Curcumin induces MCF-7 breast cancer cells pyroptosis de-pendent on caspase-1 and GSDMD. Curcumin-induced pyro-ptotic cell death in MCF-7 breast cancer cells depends on the activation of NLRP3 inflammasome. The activation of NLRP3 inflammasome induces by curcumin is mediated by cytoplasmic CTSB. Curcumin induces autophagy in MCF-7 breast cancer cells, which then induces inflammation and eventually leads to pyroptosis.
Declarations
Ethics approval and consent to participate: The animal studies covered in this manuscript have been approved by the Ethics Committee of Dalian medical university.
Consent for publication
Not applicable
Availability of Data and Materials
Not applicable
Competing Interests
The authors declare that they have no competing interests
Funding
This work is supported by the following grant: Foundation of Liaoning Province Education Administration (No. LJKZ0849)
Authors’ Contributions
HD was a major contributor in the manuscript. All authors read and approved the final manuscript.
Acknowledgements
Not applicable
Authors’ Information (optional)
Not applicable
List Of Abbreviations
MCF-7: Michigan Cancer Foundation - 7
NLRP3: NOD-like receptor protein 3
CTSB: Cathepsin B
IL-1β: Interleukin-1β
IL-18: Interleukin-18
DMSO: Dimethyl sulphoxide
3-MA: 3-methyl-adenine
DMEM: Dulbecco’s modified Eagle’s medium
MTT: 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide
GSDMD: gasdermin D
LC3: miâ?crotubuleassociatedprotein1lightchain 3
P62: protein 62
ASC: Apoptosis-associated speck-like protein containing a CARD
LPS: Lipopolysaccharide
PBS: phosphate belanced solution
References
- Fahad Ullah, M. (2019). Breast cancer: current perspectives on the disease status. Breast Cancer Metastasis and Drug Resistance, 51-64.
- Garcia-Estevez, L., & Moreno-Bueno, G. (2019). Updating the role of obesity and cholesterol in breast cancer. Breast Cancer Research, 21(1), 1-8.
- Sinha, D., Biswas, J., Sung, B., B Aggarwal, B., & Bishay-ee, A. (2012). Chemo preventive and chemotherapeutic potential of curcumin in breast cancer. Current drug targets, 13(14), 1799-1819.
- Xu, P., Yao, Y., Guo, P., Wang, T., Yang, B., & Zhang, Z. (2013). Curcumin protects rat heart mitochondria against anoxia–deoxygenation induced oxidative injury. Canadian journal of physiology and pharmacology, 91(9), 715-723.
- Kunnumakkara, A. B., Bordoloi, D., Padmavathi, G., Mon-isha, J., Roy, N. K., Prasad, S., & Aggarwal, B. B. (2017). Curcumin, the golden nutraceutical: multitargeting for multiple chronic diseases. British journal of pharmacology, 174(11), 1325-1348.
- Nagaraju, G. P., Aliya, S., Zafar, S. F., Basha, R., Diaz, R., & El-Rayes, B. F. (2012). The impact of curcumin on breast cancer. Integrative Biology, 4(9), 996-1007.
- Hu, S., Xu, Y., Meng, L., Huang, L., & Sun, H. (2018).Curcumin inhibits proliferation and promotes apoptosis of breast cancer cells. Experimental and therapeutic medicine, 16(2), 1266-1272.
- Walle, L. V., & Lamkanfi, M. (2016). Pyroptosis. Current Biology, 26(13), R568-R572.
- Man, S. M., Karki, R., & Kanneganti, T. D. (2017). Molecular mechanisms and functions of pyroptosis, inflammatory caspases and inflammasomes in infectious diseases. Immu-nological reviews, 277(1), 61-75.
- Jo, E. K., Kim, J. K., Shin, D. M., & Sasakawa, C. (2016).Molecular mechanisms regulating NLRP3 inflammasome activation. Cellular & molecular immunology, 13(2), 148-159.
- Broz, P., Pelegrin, P., & Shao, F. (2020). The gasdermins, a protein family executing cell death and inflammation. Nature Reviews Immunology, 20(3), 143-157.
- Wang, Y., Jia, L., Shen, J., Wang, Y., Fu, Z., Su, S. A., &Xiang, M. (2018). Cathepsin B aggravates coxsackievirus B3-induced myocarditis through activating the inflammasome and promoting pyro ptosis. Plops pathogens, 14(1), e1006872.
- Wu, Q. Q., Xu, M., Yuan, Y., Li, F. F., Yang, Z., Liu, Y., &Tang, Q. Z. (2015). Cathepsin B deficiency attenuates cardiac remodeling in response to pressure overload via TNF-α/ ASK1/JNK pathway. American Journal of Physiology-Heart and Circulatory Physiology, 308(9), H1143-H1154.
- Ozaki, E., Campbell, M., & Doyle, S. L. (2015). Targeting the NLRP3 inflammasome in chronic inflammatory diseases: current perspectives. Journal of inflammation research, 8, 15.
- Gerónimo-Olvera, C., Montiel, T., Rincon-Heredia, R., Castro-Obregón, S., & Massieu, L. (2017). Autophagy fails to prevent glucose deprivation/glucose reintroduction-in-duced neuronal death due to calpain-mediated lysosomal dysfunction in cortical neurons. Cell death & disease, 8(6), e2911-e2911.
- Wang, D., Zhang, J., Jiang, W., Cao, Z., Zhao, F., Cai, T., & Luo, W. (2017). The role of NLRP3-CASP1 in inflammasome-mediated neuroinflammation and autophagy dysfunction in manganese-induced, hippocampal-dependent impairment of learning and memory ability. Autophagy, 13(5), 914-927.
- Klionsky, D. J., & Emr, S. D. (2000). Autophagy as a regulated pathway of cellular degradation. Science, 290(5497), 1717-1721.
- Lee, J. E., Yoon, S. S., & Moon, E. Y. (2019). Curcumin-induced autophagy augments its antitumor effect against A172 human glioblastoma cells. Biomolecules & Therapeu-tics, 27(5), 484.
- Zhang, G., Cao, J., Yang, E., Liang, B., Ding, J., Liang, J., & Xu, J. (2018). Curcumin improves age-related and surgically induced osteoarthritis by promoting autophagy in mice. Bioscience reports, 38(4).
- Heebkaew, N., Rujanapun, N., Kunhorm, P., Jaroonwitcha-wan, T., Chaicharoenaudomrung, N., Promjantuek, W., & Noisa, P. (2019). Curcumin induces neural differentiation of human pluripotent embryonal carcinoma cells through the activation of autophagy. BioMed Research International, 2019.
- Puissant, A., Fenouille, N., & Auberger, P. (2012). When autophagy meets cancer through p62/SQSTM1. American journal of cancer research, 2(4), 397.
- Jiang, P., & Mizushima, N. (2015). LC3-and p62-based biochemical methods for the analysis of autophagy progression in mammalian cells. Methods, 75, 13-18.
- Shapira, N. (2017). The potential contribution of dietaryfactors to breast cancer prevention. European Journal of Cancer Prevention, 26(5), 385.
- Johnson, R. H., Anders, C. K., Litton, J. K., Ruddy, K. J., & Bleyer, A. (2018). Breast cancer in adolescents and young adults. Pediatric blood & cancer, 65(12), e27397.
- Giordano, A., & Tommonaro, G. (2019). Curcumin and cancer. Nutrients, 11(10), 2376.
- Verret, B., Cortes, J., Bachelot, T., Andre, F., & Arnedos,M. (2019). Efficacy of PI3K inhibitors in advanced breast cancer. Annals of Oncology, 30, x12-x20.
- Wang, Y., Gao, W., Shi, X., Ding, J., Liu, W., He, H., &Shao, F. (2017). Chemotherapy drugs induce pyroptosis through caspase-3 cleavage of a gasdermin. Nature, 547(7661), 99-103.
- Xia, X., Wang, X., Cheng, Z., Qin, W., Lei, L., Jiang, J., & Hu, J. (2019). The role of pyroptosis in cancer: pro-cancer or pro-“host”?. Cell death & disease, 10(9), 1-13.
- Jorgensen, I., & Miao, E. A. (2015). Pyroptotic cell death defends against intracellular pathogens. Immunological reviews, 265(1), 130-142.
- Shi, J., Zhao, Y., Wang, K., Shi, X., Wang, Y., Huang, H., & Shao, F. (2015). Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature, 526(7575), 660-665.
- Gan, J., Huang, M., Lan, G., Liu, L., & Xu, F. (2020). High glucose induces the loss of retinal pericytes partly via NL-RP3-Caspase-1-GSDMD-mediated pyroptosis. BioMed Research International, 2020.
- Karmakar, M., Minns, M., Greenberg, E. N., Diaz-Aponte, J., Pestonjamasp, K., Johnson, J. L., & Pearlman, E. (2020). N-GSDMD trafficking to neutrophil organelles facilitates IL-1β release independently of plasma membrane pores and pyroptosis. Nature communications, 11(1), 1-14.
- Meyers, A. K., & Zhu, X. (2020). The NLRP3 inflammasome: metabolic regulation and contribution to inflam-maging. Cells, 9(8), 1808.
- Hamarsheh, S. A., & Zeiser, R. (2020). NLRP3 inflammasome activation in cancer: a double-edged sword. Frontiers in immunology, 11, 1444.
- Miller, J. M., Thompson, J. K., MacPherson, M. B., Beus-chel, S. L., Westbom, C. M., Sayan, M., & Shukla, A. (2014). Curcumin: A Double Hit on Malignant Mesothelio-maCurcumin Induces Pyroptosis in Mesothelioma. Cancer Prevention Research, 7(3), 330-340.
- Yin, H., Guo, Q., Li, X., Tang, T., Li, C., Wang, H., &Peng, J. (2018). Curcumin suppresses IL-1β secretion and prevents inflammation through inhibition of the NLRP3 inflammasome. The Journal of Immunology, 200(8), 2835-2846.
- Jin C, Flavell RA., (2010). Molecular mechanism of NLRP3 inflammasome activation. J Clin Immunol, 30(5)-628-31.
- Mijanovic, O., Brankovic, A., Panin, A. N., Savchuk, S., Ti-mashev, P., Ulasov, I., & Lesniak, M. S. (2019). Cathepsin B: A sellsword of cancer progression. Cancer letters, 449, 207-214.
- Chevriaux, A., Pilot, T., Derangère, V., Simonin, H., Martine, P., Chalmin, F., & Rébé, C. (2020). Cathepsin B is required for NLRP3 inflammasome activation in macro-phages, through NLRP3 interaction. Frontiers in Cell andDevelopmental Biology, 8, 167.
- Jia, C., Zhang, J., Chen, H., Zhuge, Y., Chen, H., Qian, F., & Chu, M. (2019). Endothelial cell pyroptosis plays an important role in Kawasaki disease via HMGB1/RAGE/ cathespin B signaling pathway and NLRP3 inflammasome activation. Cell death & disease, 10(10), 1-16.
- Behrends, C., Sowa, M. E., Gygi, S. P., & Harper, J. W. (2010). Network organization of the human autophagy system. Nature, 466(7302), 68-76.
- Levy, J. M. M., Towers, C. G., & Thorburn, A. (2017). Targeting autophagy in cancer. Nature Reviews Cancer, 17(9), 528-542.
- Pugsley, H. R. (2017). Assessing autophagy flux by measuring LC3, p62, and LAMP1 co-localization using multispec-tral imaging flow cytometry. JoVE (Journal of Visualized Experiments), (125), e55637.
- Tanida, I., Ueno, T., & Kominami, E. (2004). LC3 conju-gation system in mammalian autophagy. The international journal of biochemistry & cell biology, 36(12), 2503-2518.
- Zhang, G., Cao, J., Yang, E., Liang, B., Ding, J., Liang, J., & Xu, J. (2018). Curcumin improves age-related and surgically induced osteoarthritis by promoting autophagy in mice. Bioscience reports, 38(4).
- Lee, J. E., Yoon, S. S., & Moon, E. Y. (2019). Curcumin-induced autophagy augments its antitumor effect against A172 human glioblastoma cells. Biomolecules & Therapeu-tics, 27(5), 484.


