Research Article - (2023) Volume 1, Issue 1
The Unique Deletion, Insertion and Point Mutation in The Upstream of Nucleocapsid (N) Gene of Different Covid-19 Variants Suggests Differential Translational Mechanisms of N-Protein
Received Date: Apr 13, 2023 / Accepted Date: May 20, 2023 / Published Date: Jul 12, 2023
Copyright: ©Â©2023 Asit Kumar Chakraborty. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Chakraborty, A. K. (2023). The Unique Deletion, Insertion and Point Mutation in The Upstream of Nucleocapsid (N) Gene of Different Covid-19 Variants Suggests Differential Translational Mechanisms of N-Protein. Transl Med OA, 1(1), 39-47.
Abstract
The nucleocapsid protein (N protein; 219 AAs) of SARS-CoV-2 was synthesized from terminal part of the ~30kb genome and a 31ERS three AAs deletion was reported in all Omicron variants but not in Alpha, Beta, Gamma and Delta variants. We found that upstream of N-gene which also a part of ORF8 gene has different deletion, insertion and point mutations in different corona virus lineages. As for example, 119DF two amino acids (5’-28243GATTTC-3’) deletion was found in ORF8 protein of Delta variants located adjacent to the N-gene upstream. Similarly, four nucleotides (5’-AACA-3’) insertion at 28262 position was found in the Gamma (P.1) variant only. Whereas all Omicron variants (BA.1/BA.2/BA.4/ BA.5) and subvariants (XBB.1.5, BQ.1.1) which were spreading now with mild diseases, have no such deletion or inser- tion. Some point mutations T>C at 28247 position were found in Alpha variants in that small region of N-gene upstream. Moreover, “GAT” sequence at 28279 position in Gamma variant similar to Wuhan and Beta (B.1.351), was changed to “CTA” in Alpha variant (B.1.1.7) at the N-gene upstream. The extra “A” at 28273 position in Alpha variant (also found in Wuhan and Gamma) was equally distributed (“AAA” vs “AAAA”) as judged by BLAST-N search but it was not instrumental if there was a sequence error. The differential hairpin structures were obtained with upstream of Delta and Gamma but no such difference was noticed with hairpin structures of different Omicron N-gene upstream. We suggest that hairpin structures with higher δG specifically in Gamma variant (P.1) may reduce the translation of N-protein and may be the one of the causes of extinct of that lineage of corona virus.
Keywords
COVID-19, N-Gene Upstream, Translation Control, Hairpin Structure, Orf8 Gene, ERS N-Protein Deletion.
Introduction
Corona virus pathogenesis has rippled this Earth with few million deaths worldwide [1]. COVID-19 was detected in March-2019 and whole genome sequencing was available from December, 2019 onwards [2]. During three years period many mutations in the diverse variants of COVID-19 genomes were reported in the NCBI SARS-CoV-2 Database [3]. However, related SARS and MERS RNA viruses were known since 2003 and considerable molecular biology of such viruses have been reported in the PubMed [4,5]. Most astonishing fact was large polyprotein (7096 AAs) synthesis in the infected cells and such proteins were proteolytically cleaved into 16 polypeptides with important biological functions [6]. The Nsp1 protein is 180aa (regulatory factor), nsp2 is 638aa (RNA topoisomerase), nsp3 is ~1945aa (C3 protease), nsp4 is 500aa (membrane factor), nsp5 is ~305aa (C5 protease), nsp6 is 290aa (membrane factor), nsp7 is 183aa (accessory protein to replication), nsp8 is 198aa (accessory protein to replication), nsp9 is 113aa (RNA binding factor), nsp10 is 139aa (RNA binding factor), nsp11 is only 13aa (unknown function), nsp12 is 918aa (RNA-dependent RNA polymerase), nsp13 is 601aa (RNA helicase-capping methyltransferase), nsp14 is 527aa (exoribonuclease-methyltransferase) , nsp15 is 346aa (endoribonuclease-recombinase), nsp16 is 298aa (2’-O Uridine rRNA methyltransferase) [9-22]. On the country, structural spike protein is 1273aa long and other structural proteins (M, N, E) of corona virus are relatively very small [24-29]. The regulatory proteins like orf3a, orf7a, orf7b, orf8 and orf10 were also characterized having interacted with many cellular proteins [30-35]. The N-protein is a strong RNA-binding protein (419aa) and requires long structured RNA as substrate [36]. Importantly, a 31ERS deletion of the protein was found in all omicron viruses including BA.2.75, BF.7, BQ.1, BQ.1.1, BQ.1.1.1 and XBB.1.5 sub subvariants [34,37]. N-protein of SARS-CoV-2 has 91% and 49% similarity to SARS-CoV and MERS-CoV respectively and is predicted to be predominantly a basic nuclear protein [38]. The N-protein has a modular structure with an N-terminal RNA-binding domain (RBD) and a C-terminal dimerization domain (CTD), including three well characterized intrinsically disordered regions (IDRs) between the RBD and CTD [39- 41]. The protein oligomerizes through its CTD and disordered C-terminal tail whereas undergoes liquid-liquid phase separation with RNA mediated by its RBD and central disordered region [42]. Cells transfected with N-protein showed a G1/S phase block with an increased expression of tubulin isomers like TUBA1C and TUBB6 [41]. The twelve phosphorylated sites and nine potential protein kinase sites in N-protein suggested as promising targets for drug discovery and development for of a recombinant virus vaccine [38,43]. The R203K and G204R mutations in the N-protein destabilized and decreased overall structural flexibility [36]. The N-protein (419aa) is required for viral life cycle and strongly binds to viral RNA regulating its replication as well as transcription of sub-genomic RNAs. A novel DNA-aptamers binding to the SARS-CoV-2 nucleocapsid protein and its variants, Delta and Omicron was discovered recently improving diagnostic tools for SARS-CoV-2 [44,45]. The replication of COVID-19 virus has been investigated but translational mechanisms were largely unknown [46]. We have reported here the sequence variation in the small upstream of N-gene of different corona virus variants
Figure.1: Structure of COVID-19 and similarity between promoters of spike protein and N-protein. The deletions in different omicron lineages were also seen and such viruses like BQ.1, XBB.1.5 and BF.7 were less pathogenic.
Methods
We searched PubMed to get idea on published papers on N-gene upstream and also searched SARS-CoV-2 NCBI database using BLAST-N and BLAST-X search methods to get sequences. Multi-alignment of protein was done by MultAlin software [47,48] and multi-alignment of DNA by CLUSTAL-Omega software, EMBL-EBI [49,50]. The ORF1ab mutants was obtained by BlastN search of deletion boundary of 60-100nt sequence and then analyzing the sequences with 95-100% similarities. The Hairpin structure of ~ 120-200nt sequence was done by OligoAnalyzer 3.1 software (Integrated DNA Technologies) [51,52]. The protein 3-D structure of N-protein was determined by SWISS-Model software [53,54].
Results
BLAST search and Multi-alignment demonstrated a variation in the upstream of N-protein among Wuhan, Beta, Gamma, Delta corona virus sequences (figure-2). But all omicron variants N-gene upstream sequences were found homogeneous and no 5’-AAGC-3’ sequence insertion at 28262 position as found in Gamma (P.1) variant. Moreover, in Alpha (B.1.1.7) variant N-protein upstream, some mutations were found. Further, in Delta variant, a “DF” two amino acid deletions (5’-28243GATTTC-3’) in the small ORF8 protein carboxy-terminal was prominent at that locus (figure-3). Previously, we demonstrated hotspot deletion sites in the ORF7a gene and three TAA termination codons creating short chimera proteins [32-34]. Thus, sequence variation in that region of 3’-end of the SARS-CoV-2 may be one of the reasons of viral extinction or slower replication and pathogenicity and N-protein was implicated in replication and pathogenicity. Indeed, downstream of N-gene, another 26nt deletion at the 3’-UTR was prominent in most omicron corona viruses [34]. Surely, such big deletion in the 3’-UTR slower viral replication and viral titre.
In Table-1, we designed many oligonucleotides to check the abundance of different N-gene upstream mutants in the SARS- CoV-2 Database. The Alpha variant sequence (5’-cgt gtt gta gat ctg ttc tct aaa cga aca aac-3’) at 28233 position at the upstream of ORF8 termination codon was justified and “tt” insertion- like alignment was true but point mutation. The Delta variant sequence (5’-gtt cgt gtt tta atc taa acg aac aaa c-3’) was also justified. Similarly, Gamma “aaac” insertion in Gamma variant COVID-19 sequence (5’-atc taa acg aac aaa caa act aaa atg tct gat aat gga c-3’) at 28253 position (accession no. NC_045512.2) was true (>998 sequences). The Alpha variant variants sequence (5’-atc taa acg aac aaa cta aat gtc tct aaa tgg ac-3’) was also justified (>5000 sequences). But one nucleotide “A” addition abolished the specificity of Alpha whereas similar addition of one “A” to other positions did retain specificity (>5000). On the contrary, further deletion of “AA” abolished the Wuhan and Beta specificities at the both positions (Table-1). It appeared that “AAA” vs “AAAA” at the middle of N-gene upstream equally distributed in the database (>5000 sequences). Thus, if it is a sequencing error or indeed “AAA” was the mutant form was not clear as “AAAA” at that position found in Wuhan (B.0) as well as Gamma and Alpha variants but not valid in Delta variant (preference of “AAA”). Moreover, “GAT” sequence at 28279 position in Gamma variant similar to Wuhan and Beta (B.1.351) variants, was changed to “CTA” in Alpha (B.1.1.7) variant at the N-gene upstream. We concluded that the upstream sequences of the N-protein were varied in different Wuhan, Beta, Alpha and Delta variants. Such changes may increase or decrease the production of N-protein but such data is very limited in the PubMed. The N-protein binds to corona virus RNA genome and regulates the replication as well as viral mRNA synthesis and expected its expression will be tightly regulated in host cells [55,56].
We performed the hairpin secondary structure of the N-gene upstream of different corona virus variants and detected minor changes in nob-like structures with different δG (figure-4). Indeed, we found higher ÃÂ?G in Alpha variant (-7.9 Kcal/mole) and to some extent in Delta variant (-9.1 Kcal/mol; Table-2). However, “AAGA” insertion in Gamma variant did not change the overall secondary stem-loop structure of N-gene upstream sequences (-9.8 Kcal/mol) as demonstrated in figure-4.
In figure-5, we demonstrated the similarity between promoters of Spike protein verses N-protein. But such similarity was not detected in the promoter of ORF1ab or M proteins (data not shown). We found the 30nt deletion in the 3’-UTR of Omicron corona viruses (BA.2, BA.4, BA.5, BF.7, BQ.1 and XBB.1.5 etc.). Such deletions including other spike deletions (24LPP and 69HV) and also spike insertions (215EPE and 249 RWMD) may be significant to lower the coronaviruses pathogenicity [32-34]. Overall, transcriptional differences among the deadly coronavirus B.1.1.7 and B.1.617.2 lineages as well as in recent omicron sub subvariants (XBB.1.5, BQ.1.1.1) never been investigated before. We have first shown here the minor but important variations among the VOC of coronaviruses with possible difference in transcriptional regulations.
Figure.2: Multi-alignment of SARS-CoV-2 Delta, Alpha, Beta, Iota, Zeta and Gamma sequences showing different deletions and insertions surrounding N-gene. A “GATTTC” sequence variation was important between Alpha and Delta Variants. A two nucleotides (TT) insertion was found upstream of ORF8 termination codon of Alpha variant and four nucleotide (AAAC) insertion was found upstream of N-protein in Gamma variant. Moreover, “GAT” was changed into “CTA” in Alpha variant than all other variants (Wuhan, Delta, Beta, Zeta and Iota). Further, a single nucleotide “a” deleted in Delta, Alpha and Iota variant as compared to Wuhan virus including Beta and Zeta variants.
Figure.3: The N-upstream of omicron corona viruses (BA.1, BA.2, BA.4 and BA.5 subvariants) have similar N-upstream as compared to Alpha, Gamma and Delta variants. The “GATTTC” deletion in Delta variant and the ”AACA” insertion in Gamma variant as well as in Alpha variant were also shown.
|
No. |
Oligonucleotides sequences and position (NC_045512) |
Population |
Variant |
|
P1 |
(28233)5’-cgtgttgttgtagatctgttctctaaacgaacaaac-3’ |
>5000 |
Alpha |
|
P2 |
(28233)5’-cgtgttgttgtagatctgctctaaacgaacaaac-3’ |
33 |
Alpha* |
|
P3 |
(28233)5’-cgtgttgttgtagatctgatctaaacgaacaaac-3’ |
0 |
Alpha** |
|
P4 |
(28253)5’-cgtgttgttgtagatctgttatctaaacgaacaaac-3’ |
1 |
Alpha*** |
|
P5 |
(28233)5’-cgtgttgttgtagatctgttctctaaaacgaacaaac-3’ |
0 |
Alpha**** |
|
P6 |
(28256)5’-gttcgtgttgttttaatctaaacgaacaaac-3’ |
>5000 |
Delta |
|
P7 |
(28256)5’-gttcgtgttgttttattatctaaacgaacaaac-3’ |
0 |
Delta* |
|
P8 |
(28256)5’-gttcgtgttgtttaatctaaacgaacaaac-3’ |
6 |
Delta** |
|
P9 |
(28256)5’-gttcgtgttgttttaatctaaaacgaacaaac-3’ |
0 |
Delta*** |
|
P10 |
(28253)5’-atctaaacgaacaaacaaactaaaatgtctgataatggac-3’ |
>998 |
Gamma |
|
P11 |
(28253)5’-atctaaacgaacaaacaaactaaatgtctgataatggac-3’ |
20 |
Gamma* |
|
P12 |
(28253)5’-atctaaacgaacaaacaactaaaatgtctgataatggac-3’ |
0 |
Gamma** |
|
P13 |
(28253)5’-atctaaacgaacaaacaaactaaaatgtctataatggac-3’ |
0 |
Gamma*** |
|
P14 |
(28253)5’-atctaaacgaacaaactaaatgtctctaaatggac-3’ |
>5000 |
Alpha |
|
P15 |
(28253)5’-atctaaacgaacaaaactaaatgtctctaaatggac-3’ |
0 |
Alpha* |
|
P16 |
(28253)5’-atctaaacgaacaaactaaatgtctctaaaatggac-3’ |
0 |
Alpha** |
|
P17 |
(28253)5’-atctaaacgaacaaactaaaatgtctctaaatggac-3’ |
>4987 |
Alpha*** |
|
P18 |
(28253)5’-atctaaacgaacaaactaaaatgtctgataatggac-3’ |
>4999 |
Wuhan/Beta |
|
P19 |
(28253)5’-atctaaacgaacaaactaaatgtctgataatggac-3’ |
>5000 |
Wuhan* |
|
P20 |
(28253)5’-atctaaacgaacaactaaatgtctgataatggac-3’ |
2 |
Wuhan** |
|
P21 |
(28253)5’-atctaaacgaacaaactaatgtctgataatggac-3’ |
110 |
Wuhan*** |
Table-1: Oligonucleotides used to BLAST-N search for the identification of different COVID-19 variants with mutated N-gene upstream sequences
|
Variant |
DNA sequences |
Delta G (Kcal/mole) |
|
Wuhan |
5’cgttctatgaagactttttagagtatcatgacgttcgtgttgttttagatttcatctaaacgaacaaa ctaaaatgtctgataatggaccccaaaatcagcgaaatgcaccccg-3’(114nt) |
-9.896 |
|
Alpha |
5’cgttctatgaagactttttagagtatcatgacgttcgtgttgttgtagatctgttctctaaacgaacaa actaaatgtctctaaatggaccccaaaatcagcgaaatgcaccccg-3’(115nt) |
-7.929 |
|
Gamma |
5’cgttctatgaagactttttagagtatcatgacgttcgtgttgttttagatttcatctaaacgaacaaac aaactaaaatgtctgataatggaccccaaaatcagcgaaatgcaccccg-3’(118nt) |
-9.896 |
|
Delta |
5’cgttctatgaagactttttagagtatcatgacgttcgtgttgttttaatctaaacgaacaaactaaatg tctgataatggaccccaaaatcagcgaaatgcaccccg-3’ (107nt) |
-9.147 |
|
Omicron |
5’cgttctatgaagactttttagagtatcatgacgttcgtgttgttttagatttcatctaaacgaacaaa cttaaatgtctgataatggaccccaaaatcagcgaaatgcaccccg-3’(114nt) |
-9.896 |
Table-2: Delta G of Hairpin structures of N-gene upstream regulating translation of N-protein
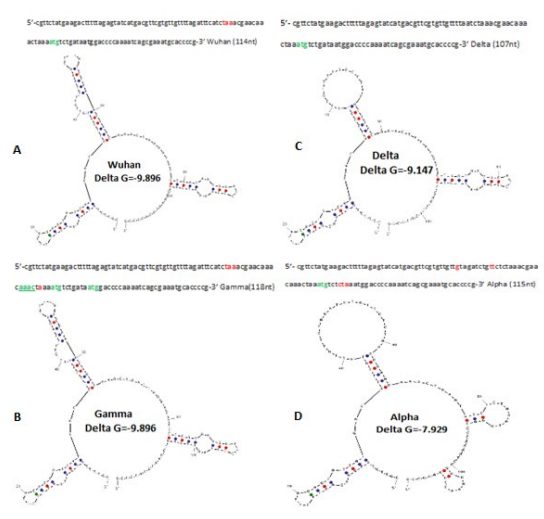
Figure.4: Hairpin structures of N-gene upstream of different corona virus variants. The hairpin structure of omicron lineages was similar to Gamma variant and was not shown here.
Figure.5: Blast-2 similarity between Spike gene and N-gene upstream showing similarity. It was shown a 5’-AACTAAA-3’ insertion in the ATG codon upstream in N-gene. The canonical TATATAAT promoter sequence was not detected. The SP3 (GGCCGGG), Myc (CACGTG), Oct1 (ATGCAAAT) eukaryotic promoter sequences were not detected
Discussion
In this communication, we reported the sequence variation among the different corona virus variants at the N-protein upstream as well as coding region. We recently reported the variations (Deletions, Insertions and Point Mutations) in the ORF1ab, ORF7a and ORF8 proteins [32-34]. Similarly, Spike protein deletions and huge mutations in omicron variants were well documented [25]. As for example, 24LPP deletion was found in most omicron corona viruses and 69HV spike deletion was found in omicron BA.1, BA.4 and BA.5 variants but not in omicron BA.2 variant. Similarly, 69HV deletion was first detected in B.1.1.7 lineages but not in B.1.617.2 lineages. Instead in B.1.617.2 lineages 157FR spike deletion was prominent. On the other hand, in omicron BA.1 lineage, both 69HV, 143VYY, 212L deletions as well as 215EPE insertion in the spike was documented [26]. Special D614G, N501Y, E484K, and L452R point mutations were instrumental in different corona virus variants and such mutations increased transmission and immune-escape [3,57]. The N501Y was found in B.1.1.7 but also transmitted in Beta, Gamma and Omicron variants but D614G mutation was found in all variants since its appearance in March, 2020. The E484K point mutation was prominent in Beta (B.1.351), Gamma (B.1.1.28.1) and Iota (B.1.526) variants as well as Mu (B.1.621) while at the same locus E484Q was reported in Kappa (B.1.617.1) variant and E484A in Omicron BA.1 (B.1.1.529) variant. The L452R point mutation was found in many COVID-19 variants like Delta (AY.103 and B.1.617.2) and Epsilon (B.1.429) variants while at the same locus L452Q point mutation was reported in Lambda (C.37) variant [6,23]. Thus, mutational landscapes continued since appearance of coronaviruses in 2003 and surely such minor variances in the subvariant populations determine its pathogenicity and omicron sub-lineages are very mild pathogenic [58].
Importantly, the deletions and mutations in the ORF1ab polyprotein (7092-7096 AAs) were reported at low rate compare to spike protein (1268-1273 AAs). The most important P4715L mutation in the RdRp was very dominant and T95I nsp2 RNA topoisomerase mutation was reported in Delta, Iota and few Omicron variants. It was a debate as D614G spike mutation increased 80% transmission in presence or absence of P4715L mutation in the RNA-dependent RNA polymerase. The3675SGF deletion in the ORF1ab was very prominent in most corona virus variants except Delta variant and 141KSF deletion was favored in omicron BA.4 variant and subvariants while the 82GHVMV locus deletions at the upstream of 141KSF, were reported in few Omicron variants recently [8].
This lineage specific changes were not new and our finding of N-gene upstream variation in sequence was important as many point mutations and deletions were found drastically enhanced viral transmission with increased pathogenicity. Obviously, characterization of corona virus sequences was done worldwide to tract specific VOC which may need extra caution to neighbor countries to control epidemic spread. The vaccination so far saved the world against Alpha and Delta variants but >30 mutations in the spike caused some doubt on early spike gene based genetic vaccines [60]. Recently, many laboratories worldwide engaged with yeast cells-expressed protein vaccine based on altered spike protein of Omicron corona viruses. However, adeno- vector based DNA-vaccine and mRNA vaccine were found good based on spike protein [59,60] as well as other proteins like RNA topoisomerase (nsp2) and uridine methyl transferases (nsp16) [9,18]. The co-infection of multidrug resistant bacteria in patients were reported and thus infections were very serious as most antibiotics failed to cure such pan-drugs resistant infections [61,62]. We recently found root and bark herbal extracts from Suregada multiflora (Gelonium multiflorum) and Cassia fistula (gloden showers) could combat such MDR bacteria [63-65]. Apart from N-protein, other small protein like nsp1 (180AAs) has profound role controlling virus replication and pathogenicity [66,67].
Conclusion
The genetic changes in RNA viruses have been well documented and ubiquitous. While the changes in N-gene upstream in different corona virus variants may throw some insights into extinct of specific lineage with time. Undoubtedly, we first highlights the possible differential regulation of N-gene in different SARS-CoV-2 variants.
Acknowledgement
The author thanks NCBI, USA for free distribution of SARS- CoV-2 database. The free distribution of CLUSTAL-Omega software is also acknowledged. AKC is a retired professor of Biochemistry.
Conflict of interest
The author declares no competing interest and the data discussed here was computer generated.
References
- Cui, J., Li, F., & Shi, Z. L. (2019). Origin and evolution of pathogenic coronaviruses. Nature reviews microbiology, 17(3), 181-192.
- Wu, F., Zhao, S., Yu, B., Chen, Y. M., Wang, W., Hu, Y., ... &Zhang, Y. Z. (2020). Complete genome characterisation of a novel coronavirus associated with severe human respiratorydisease in Wuhan, China. BioRxiv, 2020-01.
- Chakraborty, A.K. (2023). Highly infectious, less pathogenic and antibody resistant Omicron XBB. 1, XBB.1.5 and XBB. 1.5. 1-XBB. 1.5. 39 subvariant Coronaviruses do not produce ORF8 protein due to 8th codon GGA= TGA Termination Codon Mutation. Research Square (Preprint).
- Rota, P. A., Oberste, M. S., Monroe, S. S., Nix, W. A.,Campagnoli, R., Icenogle, J. P., ... & Bellini, W. J. (2003). Characterization of a novel coronavirus associated with severe acute respiratory syndrome. science, 300(5624), 1394-1399.
- Akaishi, T. (2022). Insertion-and-deletion mutations between the genomes of SARS-CoV, SARS-CoV-2, and bat coronavirus RaTG13. Microbiology spectrum, 10(3), e00716-22.
- Chakraborty, A.K., & Chanda, A. (2021). New Biotechnological Exploration on COVID-19 Proteins: Functions, Mutational Profiles and Molecular Targets for Drug Design. Sun Text Rev Virol. 2(1):115
- Clark, L. K., Green, T. J., & Petit, C. M. (2021). Structure of nonstructural protein 1 from SARS-CoV-2. Journal of Virology, 95(4), e02019-20.
- Chakraborty AK. (2023). The 82GHVMV and 141KSF deletions in the Nsp1 protein of ORF1ab polyprotein favour the creation of immune-weak SARS-CoV-2. SunText Rev Virol. ;3(2):137.
- Chakraborty, A.K. (2020). Coronavirus Nsp2 protein homologies to the bacterial DNA topoisomerase I and IV suggest Nsp2 protein is a unique RNA topoisomerase with novel target for drug and vaccine development. Virol Mycol. 9:185
- Xu, Z., Choi, J. H., Dai, D. L., Luo, J., Ladak, R. J., Li,Q., ... & Sonenberg, N. (2022). SARS-CoV-2 impairs interferon production via NSP2-induced repression of mRNA translation. Proceedings of the National Academy of Sciences, 119(32), e2204539119.
- Bacha, U., Barrila, J., Gabelli, S. B., Kiso, Y., Mario Amzel, L., & Freire, E. (2008). Development of broad-spectrum halomethyl ketone inhibitors against coronavirus main protease 3CLpro. Chemical biology & drug design, 72(1), 34-49.
- Azizogli, A. R., Pai, V., Coppola, F., Jafari, R., Dodd-o, J. B., Harish, R., ... & Kumar, V. A. (2023). Scalable Inhibitors of the Nsp3–Nsp4 Coupling in SARS-CoV-2. ACS omega, 8(6), 5349-5360.
- Liu, Y., Qin, C., Rao, Y., Ngo, C., Feng, J. J., Zhao, J., ...& Feng, P. (2021). SARS-CoV-2 Nsp5 demonstrates two distinct mechanisms targeting RIG-I and MAVS to evade the innate immune response. MBio, 12(5), e02335-21.
- Benvenuto, D., Angeletti, S., Giovanetti, M., Bianchi, M., Pascarella, S., Cauda, R., ... & Cassone, A. (2020). Evolutionary analysis of SARS-CoV-2: how mutation of Non-Structural Protein 6 (NSP6) could affect viral autophagy. Journal of Infection, 81(1), e24-e27.
- Reshamwala, S. M., Likhite, V., Degani, M. S., Deb, S. S., & Noronha, S. B. (2021). Mutations in SARS-CoV-2 nsp7 and nsp8 proteins and their predicted impact on replication/ transcription complex structure. Journal of medical virology,93(7), 4616-4619.
- Biswal, M., Diggs, S., Xu, D., Khudaverdyan, N., Lu, J., Fang, J., ... & Song, J. (2021). Two conserved oligomer interfaces of NSP7 and NSP8 underpin the dynamic assembly of SARS-CoV-2 RdRP. Nucleic acids research, 49(10), 5956-5966.
- Littler, D. R., Gully, B. S., Colson, R. N., & Rossjohn, J. (2020). Crystal structure of the SARS-CoV-2 non-structural protein 9, Nsp9. Iscience, 23(7), 101258.
- Chakraborty, A. K. (2020). Ribosomal proteins homologies and methylation of mitoribosome by Corona virus Nsp9, Nsp10 and previously described Nsp13-16 proteins suggested their roles in the inhibition of host protein synthesis: Discovery of new therapeutic targets against Corona virus infections. 10 th May, 2020.
- Gao, Y., Yan, L., Huang, Y., Liu, F., Zhao, Y., Cao, L., ...& Rao, Z. (2020). Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science, 368(6492), 779-782.
- Chakraborty, A. K. (2020). Multi-Alignment Compariso n of Coronavirus Non-Structural Proteins Nsp13-Nsp1 6 with Ribosomal Proteins and other DNA/RNA Modifyin g Enzymes Suggested their Roles in the Regulation of Hos t Protein Synthesis. Int J Clin Med Info, 3(1), 7-19.
- Rona, G., Zeke, A., Miwatani-Minter, B., de Vries, M., Kaur, R., Schinlever, A., ... & Pagano, M. (2022). The NSP14/NSP10 RNA repair complex as a Pan-coronavirus therapeutic target. Cell Death & Differentiation, 29(2), 285- 292.
- Saramago, M., Costa, V. G., Souza, C. S., Bárria, C.,Domingues, S., Viegas, S. C., ... & Matos, R. G. (2022). The nsp15 nuclease as a good target to combat SARS- CoV-2: Mechanism of action and its inactivation with FDA- approved drugs. Microorganisms, 10(2), 342.
- Chakraborty,A. K. (2020). Coronavirus ORF1ab polyprotein associated nsp16 protein is a RlmE Methyltransferase and may methylate 21S mitochondrial rRNA of host cells inhibiting protein synthesis. 2020040213. (Preprint)
- Zhang, Y., Chen, Y., Li, Y., Huang, F., Luo, B., Yuan, Y., ... & Zhang, H. (2021). The ORF8 protein of SARS-CoV-2 mediates immune evasion through down-regulating MHC-Ι. Proceedings of the National Academy of Sciences, 118(23), e2024202118.
- Chakraborty AK (2022) Hyper-Variable Spike Protein of Omicron Corona Virus and Its Differences with Alpha and Delta Variants: Prospects of RT-PCR and New Vaccine. J Emerg Dis Virol 7(1).
- Chakraborty, A.K. (2023). The 249RWMD spike protein insertion in Omicron BQ. 1 subvariant compensates the 24LPP and 69HV deletions and may cause severe disease than BF. 7 and XBB. 1 subvariants. Research Square (Preprint).
- Harvey, W. T., Carabelli, A. M., Jackson, B., Gupta, R. K.,Thomson, E. C., Harrison, E. M., ... & Robertson, D. L. (2021). SARS-CoV-2 variants, spike mutations and immune escape. Nature Reviews Microbiology, 19(7), 409-424.
- Zhang, Z., Nomura, N., Muramoto, Y., Ekimoto, T., Uemura, T., Liu, K., ... & Shimizu, T. (2022). Structure of SARS-CoV-2 membrane protein essential for virus assembly. Nature Communications, 13(1), 4399.
- Mandala, V. S., McKay, M. J., Shcherbakov, A. A., Dregni,A. J., Kolocouris, A., & Hong, M. (2020). Structure and drug binding of the SARS-CoV-2 envelope protein transmembrane domain in lipid bilayers. Nature structural & molecular biology, 27(12), 1202-1208.
- Freundt, E. C., Yu, L., Goldsmith, C. S., Welsh, S., Cheng, A., Yount, B., ... & Lenardo, M. J. (2010). The open reading frame 3a protein of severe acute respiratory syndrome- associated coronavirus promotes membrane rearrangement and cell death. Journal of virology, 84(2), 1097-1109.
- Timilsina, U., Umthong, S., Ivey, E. B., Waxman, B., & Stavrou, S. (2022). SARS-CoV-2 ORF7a potently inhibits the antiviral effect of the host factor SERINC5. Nature Communications, 13(1), 2935.
- Chakraborty, A. K. (2022). Dynamics of SARS-Cov-2 ORF7a Gene Deletions and Fate of Downstream ORF7b and ORF8 Genes Expression. Sun Text Rev Biotechnol. 3(1):142.
- Chakraborty, A. K. (2022). SARS-CoV-2 ORF8 gene CAA= TAA and AAA= TAA Termination Codon Mutations found mostly in B. 1.1. 7 Variants was Independent of Popular L84S Point Mutations. Int J Clin Med Edu Res, 1(6), 192- 208.
- Chakraborty, A.K. (2023). Highly infectious, less pathogenic and antibody resistant Omicron XBB. 1, XBB.1.5 and XBB. 1.5. 1-XBB. 1.5. 39 subvariant Coronaviruses do not produce ORF8 protein due to 8th codon GGA= TGA Termination Codon Mutation. Research Square (Preprint).
- Li, X., Hou, P., Ma, W., Wang, X., Wang, H., Yu, Z., ...& He, H. (2022). SARS-CoV-2 ORF10 suppresses the antiviral innate immune response by degrading MAVS through mitophagy. Cellular & molecular immunology, 19(1), 67-78.
- Rahman, M. S., Islam, M. R., Alam, A. R. U., Islam,I., Hoque, M. N., Akter, S., ... & Hossain, M. A. (2021). Evolutionary dynamics of SARS-CoV-2 nucleocapsid protein and its consequences. Journal of medical virology, 93(4), 2177-2195.
- Ye, Q., Lu, S., & Corbett, K. D. (2021). Structural basis for SARS-CoV-2 nucleocapsid protein recognition by single- domain antibodies. Frontiers in Immunology, 12, 719037.
- Carlson, C. R., Asfaha, J. B., Ghent, C. M., Howard, C.J., Hartooni, N., Safari, M., ... & Morgan, D. O. (2020). Phosphoregulation of phase separation by the SARS-CoV-2 N protein suggests a biophysical basis for its dual functions. Molecular cell, 80(6), 1092-1103.
- Cubuk, J., Alston, J. J., Incicco, J. J., Singh, S., Stuchell- Brereton, M. D., Ward, M. D., ... & Holehouse, A. S. (2021). The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nature communications, 12(1), 1936.
- Wu, C., Qavi, A. J., Hachim, A., Kavian, N., Cole, A. R., Moyle, A. B., ... & Leung, D. W. (2021). Characterization of SARS-CoV-2 nucleocapsid protein reveals multiple functional consequences of the C-terminal domain. Iscience, 24(6), 102681.
- Gao, T., Gao, Y., Liu, X., Nie, Z., Sun, H., Lin, K., ... &Wang, S. (2021). Identification and functional analysis of the SARS-COV-2 nucleocapsid protein. BMC microbiology, 21(1), 1-10.
- Poolsup, S., Zaripov, E., Hüttmann, N., Minic, Z., Artyushenko, P. V., Shchugoreva, I. A., ... & Berezovski,M. V. (2023). Discovery of DNA aptamers targeting SARS- CoV-2 nucleocapsid protein and protein-binding epitopes for label-free COVID-19 diagnostics. Molecular Therapy- Nucleic Acids, 31, 731-743.
- Dutta, N. K., Mazumdar, K., Lee, B. H., Baek, M. W., Kim,D. J., Na, Y. R., ... & Park, J. H. (2008). Search for potential target site of nucleocapsid gene for the design of an epitope- based SARS DNA vaccine. Immunology letters, 118(1), 65- 71.
- Kang, S., Yang, M., Hong, Z., Zhang, L., Huang, Z., Chen, X., ... & Chen, S. (2020). Crystal structure of SARS-CoV-2 nucleocapsid protein RNA binding domain reveals potential unique drug targeting sites. Acta Pharmaceutica Sinica B, 10(7), 1228-1238.
- Ge, C., Feng, J., Zhang, J., Hu, K., Wang, D., Zha, L., ...& Li, R. (2022). Aptamer/antibody sandwich method for digital detection of SARS-CoV2 nucleocapsid protein. Talanta, 236, 122847.
- Cong, Y., Ulasli, M., Schepers, H., Mauthe, M., V’kovski, P., Kriegenburg, F., ... & Reggiori, F. (2020). Nucleocapsid protein recruitment to replication-transcription complexes plays a crucial role in coronaviral life cycle. Journal of virology, 94(4), e01925-19.
- Corpet, F. (1988). Multiple sequence alignment with hierarchical clustering. Nucleic acids research, 16(22), 10881-10890.
- Wallace, I. M., Blackshields, G., & Higgins, D. G. (2005). Multiple sequence alignments. Current opinion in structural biology, 15(3), 261-266.
- Sievers, F., Wilm, A., Dineen, D., Gibson, T. J., Karplus, K., Li, W., ... & Higgins, D. G. (2011). Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Molecular systems biology, 7(1), 539.
- Altschul, S. F., Gish, W., Miller, W., Myers, E. W., & Lipman, D. J. (1990). Basic local alignment search tool. Journal of molecular biology, 215(3), 403-410.
- Roy, A., Kucukural, A., & Zhang, Y. (2010). I-TASSER: a unified platform for automated protein structure and function prediction. Nature protocols, 5(4), 725-738.
- Chou, P. Y., & Fasman, G. D. (1974). Prediction of protein conformation. Biochemistry, 13(2), 222-245.
- Bienert, S., Waterhouse, A., De Beer, T. A., Tauriello, G., Studer, G., Bordoli, L., & Schwede, T. (2017). The SWISS- MODEL Repository—new features and functionality. Nucleic acids research, 45(D1), D313-D319.
- Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., ... & Schwede, T. (2018). SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic acids research, 46(W1), W296-W303.
- Nguyen, T. T., Pathirana, P. N., Nguyen, T., Nguyen, Q. V. H., Bhatti, A., Nguyen, D. C., ... & Abdelrazek, M. (2021).Genomic mutations and changes in protein secondary structure and solvent accessibility of SARS-CoV-2 (COVID-19 virus). Scientific Reports, 11(1), 3487.
- Telwatte, S., Martin, H. A., Marczak, R., Fozouni, P., Vallejo- Gracia, A., Kumar, G. R., ... & Yukl, S. A. (2022). Novel RT-ddPCR assays for measuring the levels of subgenomic and genomic SARS-CoV-2 transcripts. Methods, 201, 15- 25.
- Chakraborty, A.K. (2022). A Method of Identification of SARS-CoV-2 Variant Using NCBI BLAST-2 100% Homology Search with Specific Oligonucleotides Selected at the Deletion Boundaries of S, N, ORF7a, ORF8 and ORF1ab Proteins. Research Square (Preprint).
- Gordon, D. E., Hiatt, J., Bouhaddou, M., Rezelj, V. V., Ulferts, S., Braberg, H., ... & Dorward, M. (2020). Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms. Science, 370(6521), eabe9403.
- Zhu, F. C., Guan, X. H., Li, Y. H., Huang, J. Y., Jiang,T., Hou, L. H., ... & Chen, W. (2020). Immunogenicity and safety of a recombinant adenovirus type-5-vectored COVID-19 vaccine in healthy adults aged 18 years or older: a randomised, double-blind, placebo-controlled, phase 2 trial. The Lancet, 396(10249), 479-488.
- Anderson, E. J., Rouphael, N. G., Widge, A. T., Jackson, L. A., Roberts, P. C., Makhene, M., ... & Beigel, J. H. (2020). Safety and immunogenicity of SARS-CoV-2 mRNA-1273 vaccine in older adults. New England Journal of Medicine, 383(25), 2427-2438.
- Cantón, R., Gijón, D., & Ruiz-Garbajosa, P. (2020).Antimicrobial resistance in ICUs: an update in the light of the COVID-19 pandemic. Current opinion in critical care, 26(5), 433-441.
- Tiri, B., Sensi, E., Marsiliani, V., Cantarini, M., Priante, G., Vernelli, C., ... & Cappanera, S. (2020). Antimicrobial stewardship program, COVID-19, and infection control: spread of carbapenem-resistant Klebsiella pneumoniae colonization in ICU COVID-19 patients. What did not work?. Journal of clinical medicine, 9(9), 2744.
- Chakraborty, A. K., Saha, S., Poria, K., Samanta, T., Gautam, S., & Mukhopadhyay, J. (2020). An Abundant New Saponin-Polybromophenol Antibiotic (CU1) from Cassia Fistula Bark Targeting RNA Polymerase. bioRxiv.
- Chakraborty, A. K. (2018). Heterogeneous phyto-antibiotics and other future therapeutics against multi-drug resistant bacteria. International Journal of Bio-Science and Bio- Technology, 10(8).
- Chakraborty, A. K. (2018). Nucleic-Acids Based Nanocarriers, in “Nanocarriers for Drug Delivery”(Eds.) Mahapatra et al. chapter-5. Elsevier Press, Amsterdam. ISBN, 978012814, 155-172.
- Lapointe, C. P., Grosely, R., Johnson, A. G., Wang, J., Fernández, I. S., & Puglisi, J. D. (2021). Dynamic competition between SARS-CoV-2 NSP1 and mRNA on the human ribosome inhibits translation initiation. Proceedings of the National Academy of Sciences, 118(6), e2017715118.
- Zhang, K., Miorin, L., Makio, T., Dehghan, I., Gao, S., Xie, Y., ... & Fontoura, B. M. (2021). Nsp1 protein of SARS- CoV-2 disrupts the mRNA export machinery to inhibit host gene expression. Science Advances, 7(6), eabe7386.



