Research Article - (2023) Volume 1, Issue 1
Virological Diagnosis of SARS-CoV-2 in a Tunisian Orthopedic Institute
2Faculty of Pharmacy, Department of Biology, University of Monastir, Tunisia
3Faculty of Medicine, LR99ES09 Laboratory of Antibiotic Resistance, University of Tunis El Manar, Tunisia
Received Date: Apr 29, 2023 / Accepted Date: May 10, 2023 / Published Date: Jun 05, 2023
Copyright: ©Â©2023 Hajer Kilani, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Kilani, H., Kaoual, S., Nakaa, H., BenLamine, Y., Chatti, L., et al. (2023). Virological Diagnosis of SARS-CoV-2 in a Tunisian Orthopedic Institute. Transl Med OA, 1(1), 01-08.
Abstract
Background: The objective of this study was to investigate the epidemiological profile of the virological diagnosis of COVID 19 at Mohamed Kassab Institute of Orthopedics.
Methods: The virological diagnosis was performed by RT-PCR from november 2020 to september 2021 on 33505 nasopha- ryngeal swabs which different reagent kits were used: Allplex® 2019 nCoV Assay, WANTAI® SARS-CoV-2 RT-PCR, WOND- FO® 2019- nCoV Real-Time RT-PCR Assay, GENESIG® Real time PCR coronavirus (COVID-19), GENESIG® COVID-19 2G . The duration from onset to laboratory test in COVID-19 suspected cases and contact individuals ranged from 0 to 14 days with a median of 3 days. The present study provides some genetic information on the lineages of SARS-CoV-2 that cir- culated in Tunisia over 6 months from april to september 2021. Lineages were assigned for 13 samples using whole-genome sequencing, partial S gene sequencing.
Results: A total of 33505 PCR tests were performed from five governorates including Manouba which represented 88.2% (29560). The positive samples were 35% (11695) and 57% (19223) were negative. The sex ratio of confirmed cases was 0.9 (5540/6155). The age group 20-65 years was the most represented with 65.9%. For sequencing her we describe the third wave was marked by the predominance of the Alpha VOC, and the fourth wave was characterized by the predominance of the Delta VOC. This study adds new genomic data to the global context of COVID-19, particularly from the North African region, and highlights the importance of the timely molecular characterization of circulating strains.
Conclusion: This study adds new genomic data to the global context of COVID-19, particularly from the North African re- gion, and highlights the importance of the timely molecular characterization of circulating strains.
Keywords
Diagnostics, Sars-Cov-2, Tunisia, Variant Of Concern, Nasopharayngeal Swabs, Reagent Kits.
Introduction
The COVID-19 pandemic first emerged in December 2019 in China, and then rapidly spread to other countries in the world in- cluding Tunisia. In the late December 2019, a novel virus called Severe Acute Respiratory Syndrome Coronavirus 2 (SARS- CoV-2), also known as 2019 novel coronavirus (2019-nCoV), was reported with an unidentified source [1]. The genomic se- quence of this newly emerged virus is highly similar to that of severe acute respiratory syndrome coronavirus (SARS-CoV) with a 79.6% sequence identity [2] (figure 1). This causes symp- toms such as cough and fever, severe pneumonia, and death. The WHO reported that more than 280 million cases of COVID-19, including approximately 5,4 million deaths, have occurred as of december 2021
Figure 1: Schematic presentation of structure and genome organization of SARS-CoV-2 based on reference sequence (EPI_ ISL_412026) [2].
(A) The virion is covered by the spike (S) proteins as well as the membrane (M) and envelope (E) proteins are placed among the S proteins in the virus envelope. The genomic RNA is surrounded by phosphorylated nucleocapsid (N) proteins inside phospholipid bilayers.
(B) The SARS-CoV-2 genome (29903 nucleotides) comprises of the 5′ UTR, ORF1a/b encoding 16 nsps for replication, four genes that encode structural proteins including S, E, M, and N proteins, six accessory genes that encode six accessory proteins such as ORF3a, ORF6, ORF7a, ORF7b, ORF8, and ORF10, as well as the 3′ UTR. The location of the seventeen high-frequency mutations and co-mutations reported in the literature are shown on the genome by vertical red lines and circles with similar color, respectively. Abbreviations: SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; 5′ UTR, 5′ untranslated region; OFR, open reading frame; nsp, non-structural protein. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
The first whole-genome sequence of SARS-CoV-2 was pub- lished on 5 January 2020, and since then, the analysis of viral sequences worldwide has been continuous, with more than 2.5 million complete genomes currently available in public databas- es, such as the GISAID platform [2-5]. Molecular analysis has shown significant genetic variability of the SARS-CoV-2 virus due to the accumulation of mutations over time. Most of these changes have little to no impact, but some mutations have an im- pact on viral properties and lead to an increase in virus transmis- sibility, more severe infection, a potential reduction in vaccine or immune effectiveness and/or escape from molecular diagnosis.
Variants that have acquired at least one of these characteristics are named variants of concern (VOCs) and require special mon- itoring. In addition, other variantsare classified as variants of interest (VOIs), variants under monitoring (VUMs) or variants under investigation (VUIs) [6-8]. By July 2021, four major vari- ants of concern (VOCs) had been described that led to increased surveillance efforts worldwide [9]. The Alpha variant, B.1.1.7 lineage (20I/501Y.V1, VOC-202012/01), also known as the UK variant, has an unusually high number of mutations and is more transmissible than the wild-type virus [10]. The Beta variant, B.1.351 lineage (501.V2, 20H/501Y.V2, VOC-202012/02), first detected and reported in South Africa in early October 2020, shares several mutations with B.1.1.7 and reduces vaccine effec- tiveness to some extent [11,12].
The Gamma variant, P.1 lineage (20J/501Y.V3, VOC- 202101/02), emerged in December 2020 in Brazil; it has 10 mu- tations in the spike protein that may affect its ability to be recog- nized by antibodies [13-15]. The Delta variant (B.1.617.2, AY.1 and AY.2 VOC-21APR-02) was first described in India and then widely spread all over the world [11,12]. This variant has eight mutations in the spike protein and is characterized by increased transmissibility in comparison with the Alpha variant [11,12]. In November 2021, a new VOC designated the Omicron variant (B.1.1.529) was first described in Botswana and in South Africa. This variant has rapidly spread all over the world and is present- ly the most frequently detected worldwide [16,17]. Therefore, the molecular monitoring of circulating strains is crucial for the timely identification of the emergence of novel SARS-CoV-2 variants. After a small decrease in disease incidence in February 2021, the country experienced a third wave of COVID-19 with the introduction of the Alpha variant in March 2021 and then fourth and fifth waves after the introduction of Delta and Omi- cron in May and December 2021, respectively.
Here, the objective of this work was to investigate the epide- miological profile of the virological diagnosis of COVID 19 at the Mohamed Kassab Institute of Orthopedics. This is a retro- spective study from November 2020 to September 2021 and includes suspected COVID-19 cases. Furthermore we provide genetic some information on the lineages of SARS-CoV-2 that circulated in Tunisia during a period of 6 months (April 2020– September 2021) covering three and the four waves.
Materials and Methods
Clinical Specimens
This study is based on nasopharyngeal samples tested in the Laboratory of medical Biology at Mohamed Kassab Institute of Orthopedics from November 2020 to September 2021. Nasopha- ryngeal swab samples were collected from suspected COVID-19 cases in fives governorates in Tunisia; Tunis, Manouba, Bizerte, Sfax and Kbeli.
Samples were mixed in 2 mL of viral transport media (VTM), consisting of Hanks’ balanced salt, 0.4% fetal bovine serum, HEPES, antibiotic and antifungal agents. Samples were trans- ported at 2–8 °C to the laboratory for processing within a few hours.
The detection of SARS-CoV-2 by conventional real-time RT- PCR using a different reagent kits: Allplex® 2019 nCoV Assay, WANTAI® SARS-CoV-2 RT-PCR, WONDFO® 2019- nCoV Real-Time RT-PCR Assay, GENESIG® Real time PCR corona- virus (COVID-19), GENESIG® COVID-19 2G [18].
Viral RNA Extraction
MagLEAD 12gC automated extraction platform (Precision Sys- tem Science, Chiba, Japan) was used to extract SARS-CoV-2 RNAs from 200 μL of nasopharyngeal and throat swabs. Ex- traction was performed according to the manufacturer’s instruc- tions. Viral RNA was eluted with 100 μL buffer and used for RT-PCR assay [19]. AlphaPrep TM VIRAL DNA/RNA Extraction Kit (Model: VDR-B096V) was used for extraction of nucleic acids from var- ious cell types. The extracted nucleic acids are applicable PCR, real-time PCR, and enzymatic reaction etc.
Preparation of Plate
Remove the sealing cover and add 200 µl of sample and 20 µl of proteinase K to the 1st or 7th well in the plate. Insert the plate correctly in the instrument (GenMagBio RNA extractor reagent kit with reliable quality)
Extraction of Nucleic Acids
After inserting the plate, close the door of the instrument and enter the correct program. When operation is finished, remove the plate from the instrument and then transfer 80 µl of extracted nucleic acids 6Th or 12th to the 1.5 ml tube.
SARSCoV2 RNA detection using real time RTPCR and Vari- ant Detection by Partial Sequencing of the S Gene The SARS- COV-2 RNA sequences are amplified by the presence of specific primers. These were designed to amplify in vitro a specific re- gion of nucleic acid: the structural genes. Depending on the type of used kit [20].
RT-PCR is the gold standard in the diagnosis of COVID-19. However, this test does not detect SARS-CoV 2 mutations, hence the genomic sequencing was maked by the laboratory of Clinical Virology, WHO Reference Laboratory for Poliomyelitis and Measles in the Eastern Mediterranean Region, Institut Pas- teur de Tunis.
Amplification by standard PCR and partial sequencing using Sanger technology was used for 13 samples collected from April to september 2021, as described previously [21]. The 648-nucle- otide-long S gene sequence encodes for the 477 to 693 amino acid residue region of the S protein. It includes key positions and allows the detection of the most important mutations character- izing most VOCs, VOIs and VUMs.
Results
A total of 33505 PCR tests were performed, from five gover- norates including Manouba which represented 88.2% (29560) (Figure 2)
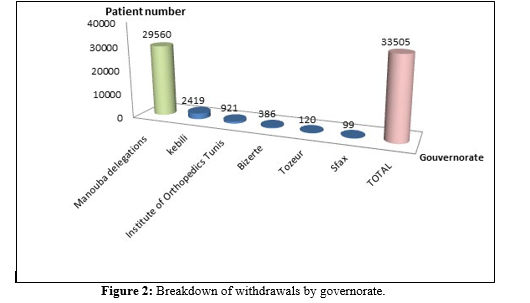
The positive samples were 35% (11695) and 57% (19223) were negative.
The sex ratio of confirmed cases was 0.9 (5540/6155).
The age group 20-65 years was the most represented with 65.9% (Table 1).
|
Age Range |
Number |
% |
|
7-19 ans |
625 |
5,34 |
|
20-55 |
6804 |
58,18 |
|
55-65 |
902 |
7,72 |
|
65-89 |
617 |
5,27 |
|
not mentioned |
2747 |
23,48 |
|
Total |
11695 |
100 |
Table 1: Distribution of confirmed cases of COVID-19 by age.
A first peak between January and February 2021 showed a positivity rate of 22.4% and a second peak between June and July with 41.2% positivity (Figure 3).
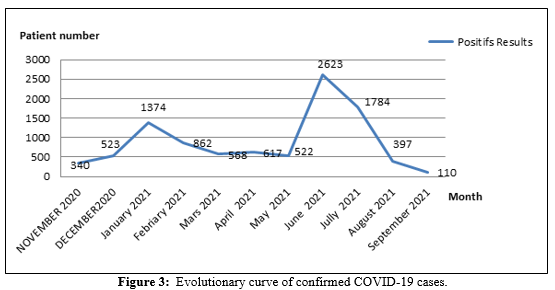
Nine out of ten of the COVID-19 positive samples sent from April were British B.1.1.7 (UK) variants of concern (Alpha) which have an accumulation of four substitution mutations at the nitrogen base levels 501-570-614- 681 and a variant A.27 (VOI) which presents four mutations by substitution at the levels of the nitrogenous bases 507-653-655 compared to the S gene of the Wuhan parent strain.\ The other three positive samples from September were B.1.617.2-like of concern (Delta) which have an accumulation of three substitution mutations; The effect of mutations in SAR- CoV-2 genomes highlighted similar profiles with D614G spike (S) variants which as the most change (Table 2).
|
Month |
Sample number |
Sequences |
Pango Lineage |
Mutations |
|
April |
1 |
S130 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
2 |
S132 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
3 |
S147 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
4 |
S151 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
5 |
S152 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
6 |
S154 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
7 |
S156 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
8 |
S161 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
9 |
S660 |
B.1.1.7 (UK) |
N501Y-A570D- D614G-P681H |
|
|
10 |
S159 |
A.27 (VOI) |
N501Y-A653V-H655Y |
|
|
September |
11 |
S-1516 |
Delta (B.1.617.2-like) |
L452R-T478K-D614G |
|
12 |
S-1519 |
Delta (B.1.617.2-like) |
L452R-T478K-D614G |
|
|
13 |
S-1524 |
Delta (B.1.617.2-like) |
L452R-T478K-D614G |
Table 2: Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) lineage distribution in Tunisia in april and Sep- tember months 2021.
This is confirming with the emergence of variants of concern (VOCs) and variants of interest (VOIs) in Tunisia between March 2020 and July 2021 (figure 4). In the background, is the number of positive cases detected in Tunisia according to the statistics published in the World Health Organization (WHO) website [22].
Figure 4: Emergence of variants of concern (VOCs) and variants of interest (VOIs) in Tunisia between March 2020 and July 2021. In the background, is the number of positive cases detected in Tunisia according to the statistics published in the World Health Or- ganization (WHO) website [22].
Discussion
Molecular tests are the standard laboratory diagnosis to confirm SARS-CoV-2 infection; RT-PCR assays for SARS-CoV-2 RNA detection in clinical specimens are widely used in COVID-19 diagnostic laboratories. In this retrospective observational study, we describe the SARS-CoV-2 that circulated in fives gover- norates in Tunisia for 11 months after its first introduction to the country in March 2020. First, Tunisia, a small country with an area of 163,610 km2 and a population of approximately 12 million, has a strategic geographic location, which makes it a junction point between the Arab world, Africa and Europe. Fur- thermore, it is known for its history of economic and cultural transactions, particularly with European and neighboring coun- tries. In addition, Tunisia was experiencing an economic and po- litical crisis that prevented total lockdown for long periods, and the introduction of the anti-SARS-CoV-2 vaccine to the popula- tion was relatively late.
The succession of the different waves observed in Tunisia is similar to the global picture of COVID-19 infection (https:// covid19.who.int/ (accessed on 30 August 2021)). In addition, the same picture of the circulation of several viral lineages has been reported in several countries around the world, such as the Czech Republic, Cyprus, the UK, Russia, South Africa and countries from the Middle East and North Africa (MENA) region [23-28].
The exchange between countries has played a crucial role in the importation of new lineages and their rapid spread across countries. This phenomenon is characteristic of airborne virus- es. Countries that have succeeded in stopping their transmission are those that have applied drastic measures, such as China and South Korea, or other countries that were able to contain the virus during the first phases of the pandemic with full contain- ment, border closures and the minimization of any contact, even between non-infected people [23-28].
In our study we observed the epidemic had first peak between January and February 2021 and a second peak between June and July with 2627 and 4825 confirmed cases respectively, which is close to the epidemiological situation in Tunisia. Such as we showed a positivity rate of 35% and more than half of the con- firmed cases were in the age group of 20-65 years. Studies done in Wuhan showed that the majority of Covid-19 patients are adults; their average age was 55.5 years. Elderly subjects ac- counted for only 10.1% of patients. Children are less frequently affected and less severely, but some authors have described the involvement of infants under one month of age [29].
The sex ratio was 0.9. Different results were found by Ketfi A. et al in Algeria with a sex ratio of 1.4 and by Nikpouraghdam M. et al in Iran with a sex ratio of 1.9 [30-31]. RT-PCR is the gold standard in the diagnosis of COVID-19. However, this test does not detect SARS-CoV 2 mutations, hence the genomic sequenc- ing was maked by the laboratory of Clinical Virology, WHO Reference Laboratory for Poliomyelitis and Measles in the East- ern Mediterranean Region, Institut Pasteur de Tunis.
In our study we researched the partial S gene sequencing for 13 specimens with Ct values less than 30 that were collected throughout April and September 2021. Nine out of ten of the COVID-19 positive samples sent from April were British B.1.1.7 (UK) variants of concern (Alpha) which have an accu- mulation of four substitution mutations at the nitrogen base lev- els 501-570-614- 681 and a variant A.27 (VOI) which presents four mutations by substitution at the levels of the nitrogenous bases 507-653-655 compared to the S gene of the Wuhan par- ent strain.The other three positive samples from September were B.1.617.2-like of concern (Delta) which have an accumulation of three substitution mutations; The effect of mutations in SAR- CoV-2 genomes highlighted similar profiles with D614G spike (S) variants which as the most change. Moreover, other lineages circulated for long periods, such as B.1.160 and B.1.177, which took an important place in the lineage landscape circulating in Tunisia. These lineages circulated from September 2020 until mid-2021 without any impact on the overall epidemiological sit- uation. B.1.160, known as 20A/EU2, is one of the main variants first reported in Europe [32,33]. The B.1.177 lineage, mostly de- tected in Europe, was first detected in early 2020 and is currently classified into more than 80 sub-lineages [32]. Further molecular characterization of a higher number of viruses will be of great interest to better characterize these two lineages.
In May 2021, the Delta variant, characterized by L452R and P681H amino acid substitutions in the spike protein, was de- tected in the country and rapidly displaced the Alpha variant, becoming the dominant variant in June–July 2021. This VOC, first detected in India in early 2021, became the most frequent- ly detected variant in many countries [5,34,35]. Indeed, it was demonstrated that the Delta variant emerged faster than the Al- pha variant and dominated the variant landscape worldwide. In the present study, the emergence of the Delta variant defined the fourth wave of SARS-CoV-2 infection in the country and partic- ipated in the resurgence of SARS-CoV-2 cases.
The circulation of the Delta variant coincided with high trans- missibility and with a large number of severe disease cases. In fact, infection with the Delta variant is characterized by the gen- eration of an average of 6 times more viral RNA copies per milli- liter than Alpha infections [12]. The Alpha VOC, first detected in the United Kingdom in late 2020, is defined by an N501Y amino acid substitution in the spike protein that increases its transmis- sibility. The Alpha VOC had become the dominant global vari- ant by early 2021 [5,28,30,31]. In Tunisia, the detection of the Delta variant decreased from August to December 2021 (data not shown). The disease incidence increased again with the in- troduction of Omicron, which was first detected in early Decem- ber and caused a new wave with a much higher transmission rate.
In order to limit the progression of the SARS-COV-2 virus in the population, it is essential to reinforce the application of the “Test-Trace-Isolate” strategy: Ø each person with symptoms suggestive of COVID-19 or having the slightest doubt must car- ry out a virus test as soon as possible. ØThe times for screening and isolating cases and their contact must be reduced for a great- er effectiveness of this strategy. ØPending the results, people must isolate themselves and contact must be reduced to a strict minimum.ØThis behavior must be coupled with adherence to all barrier gestures, in particular the wearing of mask, participation in contact identification measures, compliance with isolation measures for infected persons, persons likely to be infected or contacts classified at risk.
Preventive measures must be applied to minimize public health risks such as; Continue braking strategy, Respect health pro- tocols, Health monitoring should be associated with the vari- ousbarrier and physical distancing measures and Base prince: balance between the probability of occurrence of cases and ap- plication of general measures. This study describes the Tunisian experience in the molecular surveillance of SARSCoV2. The generated genomic data contribute to the enrichment of the glob- ally published data on SARS-CoV-2 circulation, particularly in North Africa.
Declaration of Competing Interest
The authors declare that they have no known competing finan- cial interests or personal relationships that could have appeared to influence the work reported in this paper. Les auteurs déclarent qu’ils n’ont pas d’intérêts financiers con- currents connus ou de relations personnelles qui auraient pu sembler influencer les travaux rapportés dans le présent docu- ment.
Funding: This study was funded by the authors.
Author Contributions: Original draft preparation writing, ed- iting Hajer Kilani. Review Salma.Kaoual methodology Hajer. Kilani, Hadhemi Nakaa, validation visualization supervision Sophia BouhalilaBesbes All authors read and approved the final version of the manuscript for submission.
References
- Zhu, N., Zhang, D., Wang, W., Li, X., Yang, B., Song, J., ... & Tan, W. (2020). A novel coronavirus from patients with pneumonia in China, 2019. New England journal of medi- cine.
- Rahimi, A., Mirzazadeh, A., & Tavakolpour, S. (2021). Ge- netics and genomics of SARS-CoV-2: A review of the liter- ature with the special focus on genetic diversity and SARS- CoV-2 genome detection. Genomics, 113(1), 1221-1232.
- Zhu, N., Zhang, D., Wang, W., Li, X., Yang, B., Song, J., ... & Tan, W. (2020). A novel coronavirus from patients with pneumonia in China, 2019. New England journal of medi- cine.
- WHO Coronavirus (COVID-19) Dashboard (2023).
- Elbe, S., & Bucklandâ?ÂÂMerrett, G. (2017). Data, disease and diplomacy: GISAID's innovative contribution to global health. Global challenges, 1(1), 33-46.
- World Health Organization (WHO). Tracking SARS-CoV-2Variants (2022).
- European Centre for Disease Prevention and Control(eCDC). SARS-CoV-2 Variants of Concern (2022).
- GOV-UK. Investigation of SARS-CoV-2 Variants of Con-cern: Technical Briefings (2020).
- Challen, R., Brooks-Pollock, E., Read, J. M., Dyson, L., Tsaneva-Atanasova, K., & Danon, L. (2021). Risk of mor- tality in patients infected with SARS-CoV-2 variant of con-cern 202012/1: matched cohort study. bmj, 372.
- Aksamentov, I., Roemer, C., Hodcroft, E. B., & Neher, R. A. (2021). Nextclade: clade assignment, mutation calling and quality control for viral genomes. Journal of open source software, 6(67), 3773.
- Tegally, H., Wilkinson, E., Althaus, C. L., Giovanetti, M., San, J. E., Giandhari, J., ... & de Oliveira, T. (2021). Rap- id replacement of the Beta variant by the Delta variant in South Africa. MedRxiv, 2021-09.
- Earnest, R., Uddin, R., Matluk, N., Renzette, N., Turbett, S. E., Siddle, K. J., ... & Grubaugh, N. D. (2022). Comparative transmissibility of SARS-CoV-2 variants delta and alpha in New England, USA. Cell Reports Medicine, 3(4), 100583.
- Faria, N. R., Mellan, T. A., Whittaker, C., Claro, I. M., Can-dido, D. D. S., Mishra, S., ... & Sabino, E. C. (2021). Ge- nomics and epidemiology of the P. 1 SARS-CoV-2 lineage in Manaus, Brazil. Science, 372(6544), 815-821.
- Chakraborty, S. (2022). E484K and N501Y SARS-CoV 2 spike mutants Increase ACE2 recognition but reduce affini- ty for neutralizing antibody. International immunopharma- cology, 102, 108424.
- Liu, Z., VanBlargan, L. A., Bloyet, L. M., Rothlauf, P. W.,Chen, R. E., Stumpf, S., ... & Whelan, S. P. (2021). Iden- tification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell host & microbe, 29(3), 477-488.
- World Health Organization: Home/News/ Classification of Omicron (B.1.1.529): SARS-CoV-2 Variant of Concern (2022).
- Saxena, S. K., Kumar, S., Ansari, S., Paweska, J. T., Mau- rya, V. K., Tripathi, A. K., & Abdel-Moneim, A. S. (2022). Characterization of the novel SARS-CoV-2 Omicron (B. 1.1. 529) variant of concern and its global perspective. Jour- nal of medical virology, 94(4), 1738-1744.
- Fund, G. (2020). List of SARS-CoV-2 Diagnostic test kits and equipments eligible for procurement according to Board Decision on Additional Support for Country Responses to COVID-19 (GF/B42/EDP11). Version 25.
- Chaimayo, C., Kaewnaphan, B., Tanlieng, N., Athipa- nyasilp, N., Sirijatuphat, R., Chayakulkeeree, M., ... & Hor- thongkham, N. (2020). Rapid SARS-CoV-2 antigen detec- tion assay in comparison with real-time RT-PCR assay for laboratory diagnosis of COVID-19 in Thailand. Virology journal, 17, 1-7.
- Chouikha, A., Fares, W., Laamari, A., Haddad-Boubaker, S., Belaiba, Z., Ghedira, K., ... & Triki, H. (2022). Molecu- lar epidemiology of SARS-CoV-2 in Tunisia (North Africa) through several successive waves of COVID-19. Viruses, 14(3), 624.
- Fares, W., Ghedira, K., Gdoura, M., Chouikha, A., Hadd- ad-Boubaker, S., Khedhiri, M., ... & Triki, H. (2021). Se- quencing using a two-step strategy reveals high genetic diversity in the S gene of SARS-CoV-2 after a high-trans- mission period in Tunis, Tunisia. Microbiology Spectrum, 9(3), e00639-21.
- World Health Organization (2022). WHO Health Emergen- cy Dashboard.
- Klempt, P., Brzon, O., Kašný, M., Kvapilová, K., Hubá-ek,P., Briksi, A., ... & Kvapil, P. (2021). Distribution of SARS-CoV-2 Lineages in the Czech republic, Analysis of Data from the First Year of the Pandemic. Microorganisms, 9(8), 1671.
- Richter, J., Fanis, P., Tryfonos, C., Koptides, D., Krashias, G., Bashiardes, S., ... & Christodoulou, C. (2021). Molec- ular epidemiology of SARS-CoV-2 in Cyprus. PLoS One, 16(7), e0248792.
- Du Plessis, L., McCrone, J. T., Zarebski, A. E., Hill, V., Ruis, C., Gutierrez, B., ... & Pybus, O. G. (2021). Establish- ment and lineage dynamics of the SARS-CoV-2 epidemic in the UK. Science, 371(6530), 708-712.
- Matsvay, A., Klink, G. V., Safina, K. R., Nabieva, E.,Garushyants, S. K., Biba, D., ... & Skvortsova, V. I. (2021). Genomic epidemiology of SARS-CoV-2 in Russia reveals recurring cross-border transmission throughout 2020. medRxiv, 2021-03.
- Engelbrecht, S., Delaney, K., Kleinhans, B., Wilkinson, E., Tegally, H., Stander, T., ... & de Oliveira, T. (2021). Mul- tiple Early Introductions of SARS-CoV-2 to Cape Town, South Africa. Viruses, 13(3), 526.
- Sallam, M., & Mahafzah, A. (2021). Molecular analysis of SARS-CoV-2 genetic lineages in jordan: tracking the intro- duction and spread of COVID-19 UK variant of concern at a country level. Pathogens, 10(3), 302.
- Amir, I. J., Lebar, Z., & Mahmoud, M. (2020). Covid-19: virologie, épidémiologie et diagnostic biologique. Option/ Bio, 31(619), 15.
- Ketfi, A., Chabati, O., Chemali, S., Mahjoub, M., Ghar- naout, M., Touahri, R., ... & Saad, H. B. (2020). Profil cl- inique, biologique et radiologique des patients Algériens hospitalisés pour COVID-19: données préliminaires. The Pan African Medical Journal, 35(Suppl 2).
- Nikpouraghdam, M., Farahani, A. J., Alishiri, G., Hey- dari, S., Ebrahimnia, M., Samadinia, H., ... & Bagheri, M. (2020). Epidemiological characteristics of coronavirus dis- ease 2019 (COVID-19) patients in IRAN: A single center study. Journal of Clinical Virology, 127, 104378.
- SARS-CoV-2 Lineages (2022).
- Hodcroft, E. B., Zuber, M., Nadeau, S., Vaughan, T. G.,Crawford, K. H., Althaus, C. L., ... & Neher, R. A. (2021). Spread of a SARS-CoV-2 variant through Europe in the summer of 2020. Nature, 595(7869), 707-712.
- Center of Diseases Control and Prevention (CDC). Delta Variant: What We Know About the Science. 2021.
- Challen, R., Dyson, L., Overton, C. E., Guzman-Rincon, L. M., Hill, E. M., Stage, H. B., ... & Danon, L. (2021). Early epidemiological signatures of novel SARS-CoV-2 variants: establishment of B. 1.617. 2 in England. MedRxiv, 2021-06.



