Research Article - (2025) Volume 4, Issue 1
Unified Diagnostic Intelligence: RESTful Integration of Validated Clinical Data and Machine Learning for Heart Health Prediction
Received Date: Apr 26, 2025 / Accepted Date: May 30, 2025 / Published Date: Jun 05, 2025
Copyright: ©©2025 Dr. Bhawna Singla, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Bansal, N., Singla, B. (2025). Unified Diagnostic Intelligence: RESTful Integration of Validated Clinical Data and Machine Learning for Heart Health Prediction. Curr Trends Mass Comm, 4(1), 01-15.
Abstract
Heart disease remains one of the leading causes of death globally, with early prediction being crucial for timely intervention and improved patient outcomes. However, existing predictive systems are often constrained by fragmented, non-standardized clinical data across diagnostic centers. This paper presents a RESTful framework for heart disease prediction that leverages validated clinical data and state-of-the-art machine learning (ML) models. The system integrates data collection via a user-friendly website, validation using Pydantic models, and a FastAPI-based RESTful API for scalable, asynchronous data ingestion. Data undergoes rigorous validation at both frontend and backend levels before being preprocessed and transformed for machine learning.
The ML pipeline includes imputation, scaling, and encoding of numeric and categorical features, followed by model selection using stratified 5-fold cross-validation on RandomForest and XGBoost classifiers. The best-performing model is trained on the entire dataset, and both the preprocessing pipeline and model are saved using joblib for reproducibility and inference. While the framework currently operates on synthetically generated data, it demonstrates the enormous potential of predictive modeling in healthcare. With institutional cooperation in sharing real, anonymized clinical data, this approach can significantly enhance the generalizability and accuracy of heart disease prediction, thereby transforming diagnostic decision support and ultimately benefiting public health outcomes.
Keywords
Heart Disease Prediction, RESTful API, Validated Clinical Data, Fast API, Pydantic, Random Forest, Healthcare Analytics.
Introduction
Heart disease remains one of the most significant public health challenges worldwide, contributing to high rates of morbidity and mortality. Early prediction of heart disease risk is crucial as it enables timely preventive interventions, personalized treatment plans, and more efficient allocation of healthcare resources. The effectiveness of predictive models in this domain, however, depends heavily on the availability of large, diverse, and high-quality datasets.
To date, much of the research and development in heart disease prediction using machine learning has relied predominantly on synthetic or limited dummy datasets and a constrained set of parameters. While these preliminary models have demonstrated the potential of machine learning, they fall short of capturing the complex variability and nuances present in real-world clinical data.
The power of modern machine learning techniques has grown far beyond initial expectations, capable of uncovering subtle patterns and complex interactions within data that were previously inaccessible. Therefore, if actual, comprehensive diagnostic data from diverse institutions is made available for predictive modeling, it could be a transformative boon for society. Accurate and timely predictions based on real clinical data would enable earlier interventions, reduce healthcare burdens, and ultimately save lives.
This paper advocates for a systematic approach to data sharing among diagnostic centers and healthcare institutions. By adopting standardized clinical data schemas—encompassing patient demographics, vital signs, laboratory test results, and imaging references—multiple institutions can collaboratively assemble comprehensive datasets. Such integration enhances model training by capturing population heterogeneity, thereby improving the generalizability and clinical utility of predictive algorithms in heart disease detection.
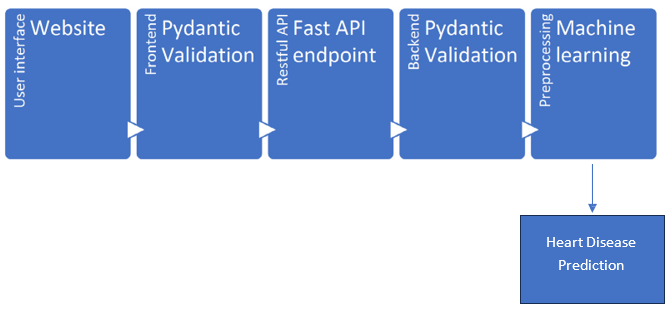
Figure 1: Flow Diagram from Accessing Data from Website to Heart Disease Prediction
Data Collection Via Web Platform
The first step in our framework is the collection of diagnostic data through a user-friendly web interface accessible to healthcare providers and diagnostic centers. This platform enables uploading of patient data records, including clinical history, lab results, and imaging references, into a centralized repository.
The web platform supports multiple input formats and enforces initial client-side validations to reduce data entry errors. This facilitates easy adoption across diverse institutions and encourages timely sharing of crucial diagnostic information.
Data Validation Using Pydantic Models: Ensuring Integrity and Clinical Relevance
Upon receiving data through the web interface, a crucial step before any meaningful analysis or predictive modeling is thorough data validation. In this framework, backend validation is performed using Pydantic models—a powerful Python library designed to enforce data integrity by validating input [1,2] according to explicitly defined schemas. Pydantic’s role is to ensure that incoming diagnostic data strictly adheres to the expected format, structure, and domain-specific rules, which is essential for building reliable and accurate machine learning models for heart disease prediction.
The Importance of Validation in Healthcare Data
Healthcare data is inherently complex, heterogeneous, and often prone to inconsistencies or errors. Inaccurate or malformed data can significantly degrade model performance, lead to incorrect clinical insights, and ultimately jeopardize patient safety. For example, a recorded age of 200 years or a negative cholesterol value is clearly erroneous and must be identified and rejected before analysis. Additionally, categorical data such as smoking status or chest pain type must fall within medically recognized categories to maintain clinical interpretability.
Therefore, validation is not merely a technical formality but a vital safeguard to ensure data quality. This process protects downstream processes—from preprocessing and feature extraction to model training and inference—from the pitfalls of corrupted or invalid input, thereby preserving the scientific rigor and trustworthiness of the entire pipeline.
Pydantic: Enforcing Strict Data Schemas
Pydantic models serve as the backbone of backend validation in this framework. These models define a strict schema specifying each data field’s expected data type, allowable value ranges, and domain constraints. For instance:
• Numeric Fields: Fields such as age, resting blood pressure, cholesterol levels, and heart rate are constrained within medically plausible ranges. Age is typically validated to be between 0 and 120 years; resting blood pressure values are restricted to ranges observed in clinical practice, e.g., 50 to 250 mmHg; cholesterol values are limited to physiologically possible numbers, preventing nonsensical or impossible values from corrupting the dataset.
• Categorical Fields: Fields representing qualitative information, such as smoking status (e.g., “yes,” “no,” “occasionally”), chest pain types (e.g., “no pain,” “mild,” “severe”), or diabetes status, are validated against predefined enumerations or sets of allowable values. This ensures that the input data aligns with clinical terminology and classifications, facilitating meaningful analysis and interpretability.
• Textual and Optional Fields: Patient names, contact information, and imaging references undergo format validation—such as phone number patterns or URL formats— ensuring syntactic correctness and usability for further processing.
The Pydantic model’s declarative nature also facilitates clear documentation, automated error messaging, and maintainable code, which is essential for evolving healthcare applications requiring adaptability to new clinical standards or expanded data elements.
Two-Tier Validation: Client-Side and Server-Side
Validation is implemented as a two-tiered approach, combining client-side and server-side mechanisms to maximize data quality and usability:
• Client-Side Validation: Basic input validation occurs in the user’s browser or device before data submission. This layer checks for common errors such as missing required fields, incorrect input formats (e.g., text in numeric fields), or values outside general bounds. Client-side validation provides immediate feedback to users, improving data entry accuracy and user experience.
• Server-Side Validation with Pydantic: Once data reaches the backend, Pydantic models perform comprehensive validation. Unlike client-side checks, server-side validation is authoritative and secure, ensuring that no malformed data bypasses safeguards even if client-side checks are disabled or circumvented. This tier handles complex inter-field dependencies and domain-specific rules that cannot be reliably enforced on the client side.
This layered validation approach significantly reduces the risk of invalid or inconsistent records entering the dataset. It balances usability—through instant feedback during data entry—with robust data integrity guarantees before storage or analysis.
Domain-Specific and Contextual Validation
Beyond basic type and range checks, the Pydantic validation framework incorporates domain-specific logic and contextual rules tailored to clinical requirements:
• Conditional Validation: For example, emergency contact information is mandated for patients above a certain age threshold (e.g., age > 60), reflecting real-world clinical practice where older patients are at higher risk and require additional contact points for urgent care. Such conditional rules ensure that data completeness aligns with patient risk profiles, improving dataset reliability and downstream clinical utility.
• Custom Validators: The Pydantic models include custom validation methods that enforce complex logic beyond basic constraints. For instance, validating that ECG or imaging references conform to acceptable URL or filename patterns ensures that only legitimate scan data is accepted. Similarly, ensuring that biomarker values such as Troponin I and CRP fall within clinically valid ranges prevents outlier values from skewing analysis.
• Cross-Field Consistency Checks: The model can also validate interdependencies between fields—for example, verifying that age and emergency contact number coexist logically or that reported symptoms align with expected diagnostic categories. This holistic validation prevents contradictory or illogical records.
Benefits of Strict Validation in Machine Learning Pipelines
The rigor enforced by Pydantic validation has direct and profound benefits for subsequent machine learning workflows:
• Improved Data Quality: Clean, consistent data reduces noise and enhances signal detection during model training. This leads to more reliable model convergence, improved accuracy, and better generalization to unseen data.
• Reduced Preprocessing Overhead: By filtering out invalid data early, the preprocessing pipeline becomes simpler and more efficient. There is less need for extensive data cleaning, imputation, or error handling downstream.
• Enhanced Model Interpretability and Trust: Models built on validated, clinically coherent data produce predictions that clinicians can trust and interpret with confidence, facilitating adoption in healthcare settings.
• Facilitated Regulatory Compliance: Healthcare data handling is subject to strict regulatory frameworks (e.g., HIPAA, GDPR). Enforcing strict validation policies supports compliance by ensuring data completeness, correctness, and auditability.
Challenges and Considerations
While Pydantic provides powerful validation capabilities, implementing such a system in real-world healthcare environments involves addressing several challenges:
• Data Variability Across Institutions: Different diagnostic sites may use varying conventions, units, or terminologies. Harmonizing this heterogeneity into a unified schema requires thoughtful model design and continuous refinement.
• Handling Missing or Incomplete Data: Clinical data often contain gaps or unreported fields. Validation must balance strictness with flexibility, allowing reasonable defaults or optional fields without compromising data quality.
• Scalability and Performance: Validating large volumes of data in real time requires efficient model implementations and backend infrastructure to avoid bottlenecks.
• User Training and Interface Design: Providing clear error messages and guidance helps end-users correct invalid inputs and improves overall data quality.
In summary, backend data validation using Pydantic models is a cornerstone of building trustworthy and effective heart disease prediction systems. By defining explicit schemas that encapsulate medically informed constraints and domain knowledge, Pydantic ensures that only high-quality, clinically meaningful data enters the modeling pipeline. This strict validation, combined with client-side checks, creates a robust two-tier defense against data errors and inconsistencies.
Ultimately, rigorous validation fosters confidence in predictive analytics, supports regulatory compliance, and paves the way for machine learning models that can reliably assist clinicians in early heart disease detection—potentially saving lives and improving healthcare outcomes at scale.
FastAPI-Driven Architecture for Scalable and Secure Data Management
Validated data is then managed via a RESTful API built using FastAPI, a modern, high-performance web framework designed for building APIs with Python. FastAPI is particularly well-suited for healthcare applications due to its ease of use, native support for asynchronous request handling, and robust data validation mechanisms through integration with Pydantic. These capabilities make it an ideal choice for designing scalable, secure, and interoperable platforms for managing sensitive diagnostic data in real time.
FastAPI serves as the central interface for communication between various components of the system, including front-end applications, databases, machine learning models, and external stakeholders. Once diagnostic data is collected and validated using strict schema definitions (as described in prior sections), FastAPI facilitates its structured ingestion into the backend system. The framework's declarative syntax ensures that every incoming data record complies with the predefined schema, thereby upholding data integrity and minimizing the risk of erroneous or incomplete records entering the processing pipeline.
Asynchronous Processing for High Throughput
One of the core advantages of FastAPI is its support for asynchronous processing using Python’s async and await capabilities. In the context of healthcare diagnostics, where multiple clinics, hospitals, and laboratories may concurrently submit data, asynchronous processing ensures that the API can handle thousands of concurrent requests without degrading performance. For instance, while one request is awaiting a response from a database or file storage system, FastAPI can continue to process other requests. This non-blocking behavior dramatically improves the system's throughput and responsiveness, making it viable for real-time applications such as emergency response systems or high-volume diagnostic centers.
Automated Interactive API Documentation
FastAPI automatically generates interactive API documentation using Swagger UI and ReDoc. These features are more than just conveniences—they are instrumental in enabling fast and accurate integration for diverse users. Developers from partner institutions or third-party system integrators can explore the API endpoints, understand the required input formats, and test different requests in real-time through the web interface. This significantly reduces onboarding time and ensures that integrations are consistent and reliable, particularly when extending the platform to new data providers or deploying it across multiple healthcare organizations.
API-Centric Architecture for Seamless Data Ingestion
The design philosophy behind using FastAPI emphasizes modularity and interoperability. This API-centric architecture enables seamless ingestion of diagnostic data from multiple sources, including hospital management systems, laboratory information systems, wearable devices, and mobile health apps. By exposing RESTful endpoints for data submission, retrieval, and modification, the system provides a standardized interface for all interactions. This unification is particularly important in healthcare, where data formats and collection protocols often vary widely between institutions.
The API endpoints are designed to accommodate both single-record submissions and bulk uploads, enabling healthcare providers to upload data manually via web forms or programmatically via batch jobs. For instance, a diagnostic center can upload an entire day’s worth of patient data in a single request, while a mobile app may submit real-time vital sign data for an individual patient at frequent intervals. FastAPI ensures that each submission—regardless of volume or frequency—is validated, timestamped, and logged for traceability.
Support for Secure Data Exchange
Given the sensitivity of patient data, FastAPI is equipped to enforce strong security measures. Authentication and authorization protocols (such as OAuth2 and JWT) are supported out of the box, allowing for secure access control. Role-based access can be implemented to ensure that users—whether doctors, nurses, researchers, or system administrators—can only view or modify data that they are authorized to handle. In addition, all API interactions are secured using HTTPS, and logging mechanisms are put in place to detect and audit any unauthorized or suspicious activity.
Modularity and Scalability of Services
Another key benefit of using FastAPI is the ability to design a modular architecture. Each major function—data validation, database storage, model prediction, and analytics—can be encapsulated in its own microservice. This modularity enables independent development, testing, deployment, and scaling of each component. For example, the machine learning service that predicts heart disease risk based on incoming data can be scaled independently of the user interface or database layers. As traffic increases, more resources can be allocated to the most heavily used services without affecting the performance of others.
Furthermore, the modular design facilitates easier maintenance and future upgrades. New services—such as integration with wearable health monitors, live ECG streaming, or AI-powered decision support—can be added as independent modules and connected to the central API, without disrupting the core functionality.
Real-Time Querying and Updating of Records
FastAPI also supports dynamic querying and updating of patient records. Healthcare providers can retrieve patient history, update lab values, append new imaging results, or adjust diagnosis inputs through intuitive API calls. This capability is critical in clinical settings where patient status can change rapidly, and care teams require the most up-to-date information to make informed decisions.
To optimize these operations, indexing and caching strategies are implemented at the database level, enabling fast retrieval times even when handling millions of records. Combined with asynchronous processing, this ensures that updates and queries are reflected almost instantaneously across the system.
Integration with ML Inference and Feedback Loops
One of the most impactful uses of FastAPI in this architecture is its ability to integrate with machine learning models for inference. Once a patient’s data is received and validated, it is routed through the preprocessing pipeline and passed to a trained ML model for prediction. The result—such as a risk score or a diagnostic category—is then returned via the API and can be displayed on dashboards or incorporated into electronic health records (EHRs). Additionally, FastAPI can capture clinician feedback on the prediction, forming the basis for a feedback loop that enhances future model accuracy. If a doctor overrides a model’s suggestion based on expert judgment, that data point can be flagged for retraining, enabling continuous learning and model refinement.
In summary, FastAPI serves as the backbone of a scalable, secure, and efficient healthcare data platform. Its RESTful API architecture enables the seamless ingestion, validation, and management of diagnostic data from multiple sources. Through features like asynchronous processing, modular deployment, and real-time data interaction, FastAPI provides the robust infrastructure needed to support predictive healthcare applications. By facilitating integration with machine learning systems and offering end-to-end security and compliance features, it empowers diagnostic centers and healthcare institutions to unlock the true potential of their data—paving the way for accurate, timely, and impactful heart disease prediction at scale.
Feature Extraction and Preprocessing
Before machine learning can be applied, diagnostic data is transformed into a structured format suitable for modeling. Feature extraction involves selecting clinically relevant attributes such as age, gender, blood pressure, cholesterol, ECG results, and biomarkers [3-12].
Data preprocessing addresses missing values through imputation strategies, normalizes numerical features, and encodes categorical variables. This ensures that the machine learning models receive consistent and clean input, improving training stability and prediction accuracy.
Machine Learning Algorithms for Prediction
The processed data feeds into machine learning models designed to classify heart disease risk and predict chest pain categories [13-31]. Algorithms such as Random Forest and XGBoost are chosen for their ability to handle heterogeneous data, capture complex nonlinear relationships, and provide robust performance.
Models are trained and evaluated using cross-validation techniques to assess generalizability and prevent overfitting. Performance metrics guide model selection and tuning. Once the best-performing model is identified, it is serialized alongside the preprocessing pipeline for efficient deployment.
Figure 2: Machine learning Prediction using Decision Forest
Societal Benefits and Future Directions
The integration and sharing of diagnostic data across institutions represent a transformative opportunity for modern healthcare. By enabling the development of robust, generalized predictive models, such collaborative frameworks can elevate diagnostic accuracy, support proactive healthcare delivery, and ultimately save lives. When institutions pool anonymized clinical data— spanning diverse geographies, demographics, and healthcare settings—they create a foundation for building machine learning models that are both representative and reliable. These models can be embedded into Clinical Decision Support Systems (CDSS) to assist healthcare professionals in making timely, data-driven decisions.
Figure 3: Integration of Results with LLM
One of the most immediate societal benefits of this paradigm is the early detection of heart disease risk, which remains a leading cause of death globally. Predictive models, trained on high-quality data from a broad population base, can identify at-risk individuals before clinical symptoms become evident. Early intervention not only improves prognosis but also reduces the long-term burden on healthcare systems by minimizing emergency cases, hospitalizations, and invasive procedures. For patients, this translates into improved health outcomes, longer life expectancy, and enhanced quality of life.
Economically, predictive diagnostics can lead to significant cost savings for healthcare providers and governments. Preventive care is invariably less expensive than managing advanced stages of chronic illness. With accurate risk prediction tools in place, healthcare systems can shift from a reactive model to a proactive model—allocating resources more efficiently, targeting high-risk populations, and optimizing care pathways. This shift also empowers physicians to focus more on personalized treatment plans, reducing unnecessary tests and improving patient satisfaction.
Looking ahead, the widespread adoption of secure data governance models will be essential to sustain and scale these benefits. While the collaborative use of healthcare data offers enormous potential, it must be balanced with rigorous standards for privacy, consent, and data security. Emerging technologies such as federated learning provide a promising solution to this challenge. In federated learning, data never leaves the institution where it was generated; instead, models are trained locally, and only the learning parameters are shared. This approach enables institutions to contribute to collective model training without exposing sensitive patient information, thus preserving data privacy and regulatory compliance.
Advancements in Few-shot and Zero-shot learning: LLMS can be made more understanding with minimal input(Few shot learning) or even no task specific training examples (zero shot learning). In addition, as healthcare data continues to evolve—with the incorporation of genomics, wearables, and real-time monitoring— models must be continuously updated to remain relevant. This calls for systems that support ongoing model retraining and validation using the latest data inputs. Automated pipelines that incorporate continuous integration, validation, and deployment of new model versions will ensure that predictions remain accurate over time and reflect the most current clinical knowledge and population trends. In conclusion, the structured sharing and utilization of diagnostic data, backed by secure and ethical frameworks, will not only improve the predictive power of healthcare systems but also lead to more equitable and effective care delivery. The future of medicine lies in data-informed decisions—and collaborative innovation today sets the stage for a healthier, smarter society tomorrow.
Author Contributions
Conceptualization, N.B. and B.S.; methodology, B.S.; validation, N.B. All authors have read and agreed to the published version of the manuscript.
Funding
This work received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The study is done on dummy data availability from diagnostics websites. But strongly suggest that diagnostics centers, Institutions can share their data.
Acknowledgments
The authors want to acknowledge their Institute administration, colleagues and family who motivated and supported them all time to write this article within time management.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Narayanan, P. K. (2024). Getting Started with Data Validation using Pydantic and Pandera. In Data Engineering for Machine Learning Pipelines: From Python Libraries to ML Pipelines and Cloud Platforms (pp. 163-196). Berkeley, CA: Apress.
- Dao, N., Hassan, S. M., Waxman, A. B., Washko, G. R., &Rahaghi, F. N. (2025). Validation Pipeline for Unstructured Data Extraction UsingArtificial Intelligence. American Journal of Respiratory and Critical Care Medicine, 211(Abstracts), A4303-A4303.
- Rajpurkar, P., et al. “Cardiologist-Level Arrhythmia Detection with Convolutional Neural Networks.” Nature Medicine, vol. 25, no. 1, 2019, pp. 65–69.
- Johnson, A. E., Pollard, T. J., Shen, L., Lehman, L. W. H.,Feng, M., Ghassemi, M., ... & Mark, R. G. (2016). MIMIC-III,a freely accessible critical care database. Scientific data, 3(1), 1-9.
- Lundberg, S. M., & Lee, S. I. (2017). A unified approach to interpreting model predictions. Advances in neural information processing systems.
- Lundberg, S. M., Nair, B., Vavilala, M. S., Horibe, M., Eisses,M. J., Adams, T., ... & Lee, S. I. (2018). Explainable machine-learning predictions for the prevention of hypoxaemia during surgery. Nature biomedical engineering.
- Deo, R. C. (2015). Machine learning inmedicine. Circulation, 132(20), 1920-1930.
- Krittanawong, C., et al. (2019 ). “Machine Learning and Deep Learning in Cardiovascular Disease: A Review.” Computers in Biology and Medicine, vol. 111.
- Obermeyer, Z., & Emanuel, E. J. (2016). Predicting the future—big data, machine learning, and clinical medicine. New England Journal of Medicine, 375(13).
- Miotto, R., Li, L., Kidd, B. A., & Dudley, J. T. (2016). Deep patient: an unsupervised representation to predict the future of patients from the electronic health records. Scientific reports.
- Beam, A. L., & Kohane, I. S. (2018). Big data and machine learning in health care. Jama, 319(13), 1317-1318.
- Chen, T., & Guestrin, C. (2016, August). Xgboost: A scalable tree boosting system. In Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining (pp. 785-794).
- Breiman, L. (2001). Random forests. Machine learning, 45,5-32.
- Friedman, J. H. (2001). Greedy function approximation: a gradient boosting machine. Annals of statistics.
- Lundberg, S. M., Erion, G., Chen, H., DeGrave, A., Prutkin,J. M., Nair, B., ... & Lee, S. I. (2020). From local explanations to global understanding with explainable AI for trees. Nature machine intelligence.
- Choi, E., Biswal, S., Malin, B., Duke, J., Stewart, W. F., & Sun,J. (2017, November). Generating multi-label discrete patient records using generative adversarial networks. In Machine learning for healthcare conference (pp. 286-305). PMLR.
- Zhang, Z., et al. (2019). “Predictive Analytics with Electronic Health Records.” Journal of Healthcare Informatics Research.
- Chen, I. Y., et al. (2020).“Ethical Machine Learning in Health Care.” Annual Review of Biomedical Data Science.
- Kaul, V., et al. (2019).“Using Machine Learning to Understand Clinical Risk and Enhance Patient Care.” American Journal of Medicine.
- Topol, E. J. (2019). High-performance medicine: the convergence of human and artificial intelligence. Nature medicine, 25(1), 44-56.
- Belle, A., Thiagarajan, R., Soroushmehr, S. R., Navidi, F., Beard, D. A., & Najarian, K. (2015). Big data analytics in healthcare. BioMed research international, 2015(1), 370194.
- Rajkomar, A., Oren, E., Chen, K., Dai, A. M., Hajaj, N., Hardt, M., ... & Dean, J. (2018). Scalable and accurate deep learning with electronic health records. NPJ digital medicine.
- Krittanawong, C., et al. “Machine Learning for Predicting Cardiovascular Events.” Nature Reviews Cardiology, 2020.
- Fawcett, T. (2006). An introduction to ROC analysis. Pattern recognition letters, 27(8), 861-874.
- Jiang, F., Jiang, Y., Zhi, H., Dong, Y., Li, H., Ma, S., ... & Wang, Y. (2017). Artificial intelligence in healthcare: past, present and future. Stroke and vascular neurology.
- Miotto, R., et al. (2017). “Deep Learning for Healthcare: Review, Opportunities and Challenges.” Briefings in Bioinformatics.
- Agarwal, R., et al. (2019).“The Role of Machine Learning in Precision Medicine.” Annual Review of Biomedical Engineering.
- Bansal, Neha, and Bhawna, Singla. (2025). Data-Driven decision making for car dealerships using web scraping techniques. Gradiva, 11(1), 52.
- Bansal, Neha, and Bhawna, Singla. (2025). Education 4.0 using Generative AI and metaverse. Liberte Journal Wos, 13(1).
- Bansal, Neha , and Bhawna, Singla. (2024). Heart Health Detector GPT Based on GPT-4o Model. Automation, Control and Intelligent Systems, 12(4), 114-124.
- Neha Bansal, Bhawna Singla “Healthcare Facilities Enhancement Using Artificial Intelligence Chatbots” proceedings published in Annual International Congress on Computer Science, Oxford, United Kingdom, April 17-18, 2025.




