Research Article - (2022) Volume 6, Issue 2
Structure/Epitope-Based Immunoinformatics Analysis of Structural Proteins of 2019 Novel Coronavirus
2Laboratory of Chemical Biology and State Key Laboratory of Rare Earth Resource Utilization, Changchun Institute of Applied Chemistry, Chinese Academy of Sciences, Changchun, Jilin 130022, China
3State Key Laboratory of Respiratory Disease, Guangzhou Medical College, Guangzhou 510006, China
Received Date: Sep 08, 2022 / Accepted Date: Sep 13, 2022 / Published Date: Sep 20, 2022
Copyright: ©Copyright: Ã?©2022 Liteng Yang. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Li., Y., Yang., L., Sun., X., Zhong., N. (2022). Structure/Epitope-Based Immunoinformatics Analysis of Structural Proteins of 2019 Novel Coronavirus. Stem Cell Res Int, 6(2),83-99.
Abstract
The newly identified 2019 novel coronavirus (2019-nCoV) has caused more than ten million laboratory-confirmed human infections, including 20,000 deaths, posing a serious threat to human health. Currently, however, there is no specific antiviral treatment or vaccine. To identify immunodominant peptides for designing global peptide vaccine for combating the infections caused by 2019-nCoV, the structure and immunogenicity of 2019-nCoV structural protein were analyzed by bioinformatics tools. 33 B-cell epitopes and 39 T-cell epitopes were determined in four structural proteins via different immunoinformatic tools in which include spike protein (22 B-cell epitopes, 25 T-cell epitopes ), nucleocapsid protein (7 B-cell epitopes, 6 T-cell epitopes), membrane protein (2 B-cell epitopes, 7 T-cell epitopes), and envelope protein (2 B-cell epitopes, 1T-cell epitopes), respectively. The proportion of epitope residues in primary sequence was used to determine the antigenicity and immunogenicity of proteins. The envelope protein has the largest antigenicity in which residue coverage of B-cell epitopes is 24%. The membrane protein possesses the largest immunogenicity in which residue coverage of T-cell epitopes is 55.86%. The reason that immune storm was caused by 2019-nCoV maybe that the membrane and envelope protein expressed plentifully in cell infected. Further, studies involving experimental validation of these predicted epitopes is warranted to ensure the potential of B-cells and T-cells stimulation for their effective use as vaccine candidates. These findings provide the basis for starting further studies on the pathogenesis, and optimizing the design of diagnostic, antiviral and vaccination strategies for this emerging infection.
Keywords
2019-nCoV; Structural protein; B-cell epitope; T-cell epitope; Prediction; Bioinformatics
Introduction
A novel coronavirus (2019-nCoV) associated with human to hu- man transmission and severe human infection has been recently reported from the city of Wuhan in Hubei province in China [1]. The ongoing outbreak of a novel coronavirus (2019-nCoV) caus- es great global concerns. Based on the advice of the International Health Regulations Emergency Committee and the fact that to date 24 other countries also reported cases, the WHO Director-Gener- al declared that the outbreak of 2019-nCoV constitutes a Public Health Emergency of International Concern on 30 January 2020 [2]. A total of 1,320 confirmed and 1,965 suspect cases were re- ported up to 25 January 2020; of the confirmed cases 237 were severely ill and 41 had died [3]. Currently, however, there is no specific antiviral treatment or vaccine [4].
Vaccines play a crucial role in providing protection against a particular disease to host organism, therefore it provides help in saving millions of lives annually across the globe [5]. Vaccine development processes has depended entirely upon experimental techniques, being generally very laborious and time-consuming [6]. There has been much encroachment in the area of computa- tional biology that aids many types of research and helps in dimin- ishing the expected time consumption [7]. Further combination of various bioinformatics prediction methods could significantly increase the prediction accuracy [8]. Thus bioinformatics studies could provide reliable guidance in selecting specific immunogen- ic epitopes, which will be significant for vaccine design, epitope mapping and antibody studies.
Coronavirus is a class of enveloped RNA virus with a 27–3l kb long single-stranded positive-sense genome [9]. The region down- stream of ORF1 contains at least 10 small ORFs, encoding the spike protein (S protein), small envelope protein (E protein), mem- brane protein (M protein), nucleocapsid protein (N protein) and the assumed nonstructural proteins [10]. The genome is packed inside a helical capsid formed by the nucleocapsid protein (N) and further surrounded by an envelope [11]. Associated with the viral envelope are at least three structural proteins: The membrane pro- tein (M) and the envelope protein (E) are involved in virus assem- bly, whereas the spike protein (S) mediates virus entry into hosT- cells, which are essential for viral assembly [4]. In the present study, we analyzed the CoV structural protein's sequence and 3D structures by using bioinformatics methods. B-cell epitopes and T-cell epitopes of structural protein were determined ultimately via different bioinformatic tools. Our study will be beneficial for the diagnosis and treatme nt of 2019-nCoV infection and the effective epitope-based vaccine design.
Materials and Methods
Sequence retrieval and physiochemical analysis
The structural protein sequence were obtained from the GenBank Nucleotide with the accession number of MN975262. The homol- ogous amino acid sequences were selected by BLAST in NCBI (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The multiple alignment of homologous amino acid sequences was performed using Clust- al X 2.1 program [12]. Physicochemical analysis of the protein sequences was predicted by ProtParam (http://web.expasy.org/ protparam/), including molecular weight, amino acid composition, negatively charged residues (Asp + Glu), positively charged resi- dues (Arg + Lys), theoretical isoelectric point pI, aliphatic index, grand average of hydropathicity (GRAVY) and instability index of proteins [13].
Tertiary structure prediction and validation
Homology modeling was used for constructing the tertiary struc- ture of proteins. The homologous templates suitable for proteins were selected by PSI-BLAST server and SWISS-MODEL server [14]. The best template was retrieved from the results of previous methods, which were based on the high score, lower e-value, and maximum sequence identity. Tertiary structure was constructed by MODELLER v9.20 [15]. The Ramachandran plot for all the mod- els was generated, showing the majority of the protein residues in the favored regions, allowed regions and disallowed regions [16].
In silico prediction of B-cell epitopes
Dozens of B-cell antigen epitopes have been discovered follow- ing various algorithms, offering us multiple reliable means of pre- dicting the antigen's hydrophilic regions, accessibility, flexibility and antigenic solubility [17]. We adopted the B-cell epitope pre- diction method based on machine learning and comprehensively used DNAStar, BepiPred 1.0 server, and COBEP to predict the common epitopes of protein. In the DNAStar, hydrophilicity, flex- ibility, accessibility, and antigenicity of the amino acid sequence were chosen as parameters for B-cell Epitopes [18]. The amino acid sequence is provided for the BepiPred 1.0 server. We used 0.35 as the threshold. Scores beyond 0.35 account for potential epitote, and a higher score indicates a higer probability of the ex- isting epitope [19]. COBEP are developed based on support vector machine, combining tripeptide similarity and propensity score pre- dicted B-cell epitope [20]. The consensus epitopes were accurate through combining the results of the three tools as the predicting results of B-cell epitopes. B-cell epitopes were selected further for the ultimate results of B-cell epitopes through excluding the amino acid of protein structure internal. B-cell epitopes predicted by computational tools were mapped on linear sequence and on the three dimensional model of structural protein to confirm their position.
In silico prediction of T-cell epitopes
T-cell epitopes are principally predicted on the basis of identifying the binding of amino acid fragments to the MHC complexes that can activate T-cells. The binding strength of each peptide to the given MHC is estimated by NetMHCII 2.2 at a set threshold level. Given the MHC alleles have tens of thousands of kinds, in order to ensure the representative and reliability prediction results, we selected the most common MHC in the population to predict their peptide binding activity with protein, including HLA-DR, HLA- DQ, and HLA-DP. HLA-DR 101, HLA-DR 301, HLA-DR 401 and HLADR 501 were used to predict HLA-DR-based T-cell epitope. HLA-DQA10102-DQB10502,HLA-DQA10201-DQB10301, HLA-DQA10501-DQB10302, and HLA-DQA10601-DQB10402 were used to predict HLA-DQ based T-cell epitope prediction. HLA-DPA10103-DPB10601,HLA-DPA10201-DPB10101, HLA- DPA10201-DPB10501, and HLA-DPA10301-DPB10402 were used to predict HLA-DP based T-cell epitope prediction. If the predicting results of all four alleles were non epitope, then the consensus result was 0% T-cell epitopesï¼?if the results were only one or no non-epitope, the consensus result was 75 or 100% T-cell epitopes, respectively [21]. Ultimately, the results that consensus epitope result was 75 or 100% were determined as the ultimate epitope results [22]. As a result, the ultimate consensus T-cell epi- tope results were obtained by combining the results of the HLA- DR alleles epitopes, HLA-DQ alleles epitopes, and HLA-DP al- leles epitopes.
Results and Discussion
Physiochemical analysis of structural protein of 2019-nCoV
The complete genome sequence of 2019-nCoV was available at GenBank accession (No.MN975262). The 2019-nCoV includes four structural proteins that are required to drive cytoplasmic viral assembly: S protein, M protein, N protein and E protein. The char- acterization of physicochemical attributes of antigenic proteins was a major step discerning the information about the biological activity of viral protein sequences [23]. Physicochemical analy- sis of the structural proteins was performed by ProtParam such as instability index, extinction coefficient, GRAVY, aliphatic index, theoretical pI of the protein sequences of 2019-nCoV (Table 1).
Table 1: Physicochemical characterization of structural proteins of 2019-nCoV.
|
Proteins |
ORF |
Number of amino acids |
Molecular weight |
theoretical pI |
Instability index |
GRAVY |
GenBank accession |
|
S protein |
ORF2 (21563..25384) |
1273 |
141.2 kDa |
6.24 |
33.01 |
-0.079 |
QHN73810.1 |
|
E protein |
ORF4 (26245..26472) |
75 |
8.4 kDa |
8.57 |
38.68 |
1.128 |
QHN73812.1 |
|
M protein |
ORF5 (26523..27191) |
222 |
25.2 kDa |
9.51 |
39.14 |
0.446 |
QHN73813.1 |
|
N protein |
ORF9a (28274..29533) |
419 |
45.6 kDa |
10.07 |
55.09 |
-0.971 |
QHN73817.1 |
The results show that The S protein is a large protein of 1273 ami- no acids, with a molecular weight of 141.2 kDa and a theoretical pI of 6.24. The M protein and N protein had 222 and 419 amino acids, with a molecular weight of 25.2 and 45.6 kDa and a theo- retical Pi of 9.5 and 10.1. The CoV E protein is a short protein of 75 amino acids, ranging from 8.4 kDa in size. Instability index of four structural protein is 30-60, meaning that all structural proteins of 2019-nCoV are stable. The Aliphatic index and Grand average of hydropathicity (GRAVY) are 50-120 and -1 to 2, respectively. The GRAVY of S protein and N protein is negative, meaning they exhibited hydrophilic character. The M protein and E protein of 2019-nCoV exhibited hydrophilic character. The physicochemical character may provide a selective advantage in the infected host [24].
Structure and antigenicity of structural proteins S protein
The spike forms large protrusions from the virus surface, giving coronaviruses the appearance of having crowns (hence their name; corona in Latin means crown). In addition to mediating virus entry, the spike is a critical determinant of viral host range and tissue tropism and a major inducer of host immune responses [25]. The coronavirus S protein is a multifunctional molecular machine that mediates coronavirus entry into hosT-cells [25]. Homology mod- eling can construct a target structure on the basis of suitable tem- plates extracted from homologous sequences [26]. Since S protein structure is not available in the PDB molecular database, we used S protein of SARS-CoV as template to predict the 3D structure of 2019-nCoV S protein through the SWISS-MODEL server and MODELLER software. The results revealed that the S protein of 2019-nCoV is a clove-shaped trimer with three S1 heads and a trimeric S2 stalk (Figure 1A). Monomer of S protein comprised of S1 and S2 subunits (Figure 1B). The S1 subunit contains a signal peptide, followed by N-terminal domain (NTD) and C-terminal domain (S1-CTD) (Figure 1C).
Figure 1: Structural features of S protein of 2019-nCoV. (A) Protein structure of trimeric S protein of 2019-nCoV. Three monomers are shown (blue, red, and yellow). (B) Protein structure of a monoFmer S protein of 2019-nCoV. (C)The important functional elements of the S protein of 2019-nCoV. The S1 N-terminal domain (NTD), the S1 C-terminal domain (CTD), and the S2 are colored in the same way.
S1-CTD contains a receptor-binding motif, which presents a gently concave outer surface to bind ACE2 (Figure 2A). The S2 subunit contains conserved fusion peptide, heptad repeat, transmembrane domain, and cytoplasmic domain. During virus entry, S1 binds to a receptor on the hosT-cell surface for viral attachment, and S2 fuses the host and viral membranes, allowing viral genomes to enter hosT-cells. Previous studies suggest that protein antigenic- ity is generally determined by its specified epitopes instead of the full length sequence [27]. To identify the antigen epitopes, bioin- formatics methods are used to predict their sequences. Predicted epitopes are further synthesized in vitro and validated with exper- iments. DNAStar, BepiPred 1.0 server, and COBEP were used to identify the potential antigen epitope regions of protein (Table 2).
Table 2: The predictions results of B-cell epitopes of S protein.
|
Tools |
Location of the prediction results |
|
DNAStar |
20-30,40-45,71-86,95-101,108-117,147-152,178-191,251-258,279-286,315-325,355-361,381-387,408-430,439-449,471- 485,526-540,552-559,569-581,601-607,657-660,675-687,773-781,789-798,806-817,935-942,1080-1088, 1112-1124. |
|
BepiPred |
21-31,71-81,110-113,250-258,280-287,382-387,407-428,439-447,461-484,496-506, 526-532,551-554,567-580,601-607,636- 643,656-659,675-687,700-707,772-780,789-797,805-816,936-941,1054-1059,1069-1090,1114-1122,1137-1150,1157-1171. |
|
CBTOPE |
21-32,66-77,134-147,164-167,174-181,203-215,256-274,280-286,304-325,345-356, 376-389,408-415,423-455,472-524,550- 557,598-621,636-643,659-674,725-737,753-768,776-788,804-824,903-941,961-976,1022-1049,1075-1088,1098-1119,113- 7-1146, 1154-1178. |
The final B-cell epitopes of S protein included 22 sequences through combining the results of the three tools. The S1 and S2 of spike protein has 16 and 6 B-cell epitopes, respectively. These peptides were also shown in the primary and tertiary structure of the S1 and S2 of spike protein (Figure 2).
Figure 2: B-cell epitopes distribution of S protein of 2019-nCoV. (A) B-cell epitopes distribute in the primary sequence of S protein. Sequences of the B-cell epitopes were are highlighted by green background and named B1-B22. The sequences of receptor-binding motif are highlighted by blue font. (B) B-cell epitopes on tertiary structure of S1 domain. (C) B-cell epitopes on tertiary structure of S2 domain.
Table 3. The predictions results of T-cell epitopes of S protein.
|
Tools |
Location of the prediction results |
|
DRB1∗01:01 |
2–10,4–12,5–13,7–15,40–48,46–54,53–61,55–63,58–66,62–70,65–73,98–106,104–112,115–123,117–125,157–165,168– |
|
|
176,169–177,186–194,200–208,220–228,223–231,233–241,235–243,238–246,239–247,241–249,242–250,258–266,259– |
|
|
267,267–275,268–276,269–277,275–283,300–308,306–314,312–320,318–326,342–350,347–355,365–373,377–385,379– |
|
|
387,394–402,451–459,453–461,489–497,502–510,505–513,510–518,515–523,517–525,531–539,538–546,543–551,582– |
|
|
590,590–598,627–635,642–650,643–651,648–656,686–694,689–697,693–701,695–703,696–704,711–719,718–726,724– |
|
|
734,752–760,759–767,764–772,797–805,802–810,818–826,821–829,826–834,833–841,855–863,858–866,869–867,870– |
|
|
878,874–882,875–883,886–884,888–896,891–899,898–906,927–935,959–967,962–970,963–971,970–978,993–1001,994– |
|
|
1002,996–1004,1001–1009,1005–1013,1007–1015,1010–1018,1032–1040,1041–1049,1042–1050,1047–1055,1048– |
|
|
1056,1060–1068,1062–1070,1075–1083,1103–1111,1129–1137,1169–1177,1215–1223,1216–1224,1217–1225,1218– |
|
|
1226,1220–1228,1241–1249,1262–1270,1265–1273. |
|
DRB3∗01:01 |
28–36,36–42,37–45,42–50,43–51,47–55,50–58,57–65,58–66,78–86,85–93,118–126,119–127,127–135,132–140,133– |
|
|
141,134–142,135–143,144–152,145–153,152–160,160–168,168–176,169–177,170–178,174–182,175–183,186–194,192– |
|
|
200,193–202,195–203,211–219,212–220,225–233,230–238,240–248,269–277,277–285,284–292,291–298,306–314,312– |
|
|
320,318–326,340–348,357–365,361–369,367–375,369–377,392–400,394–402,395–403,399–407,401–409,402–410,417– |
|
|
425,421–429,445–453,456–464,463–471,464–472,489–497,495–503,497–505,505–513,507–515,551–559,564–572,565– |
|
|
573,568–576,584–592,624–632,627–635,635–643,643–651,658–666,659–667,660–668,699–707,706–714,718–726,734– |
|
|
742,741–749,780–788,781,789–797,800–808,805–813,816–824,817–825,821–829,825–833,826–834,833–841,836– |
|
|
844,863–871,864–872,898–906,903–911,904–912,915–913,926–934,976–984,990–998,991–999,1067–1075,1081– |
|
|
1089,1094–1102,1095–1103,1102–1110,1114–1122,1115–1123,1148–1156,1208–1216,1220–1228,1256–1264. |
|
DRB4∗01:01 |
7–15,8–16,16–24,47–55,97–105,98–106,115–123,116–124,117–125,127–135,168–176,190–198,208–216,209–217,210– |
|
|
218,223–231,230–238,233–241,234–242,235–243,236–244,237–245,238–246,317–325,507–515,508–516,509–517,511– |
|
|
519,557–565,558–566,579–587,607–615,608–616,684–692,685–693,690–698,691–699,750–758,751–759,781–789,814– |
|
|
822,815–823,847–855,856–864,857–865,861–869,862–870,863–871,890–898,891–899,894–902,895–903,896–904,897– |
|
|
905,908–916,919–917,920–928,921–929,959–967,960–968,972–980,973–981,993–1001,995–1003,996–1004,999– |
|
|
1007,1004–1012,1005–1013,1006–1014,1007–1015,1010–1018,1011–1019,1047–1055,1048–1056,1057–1065,1129– |
|
|
1137,1175–1183,1176–1184,1195–,1218–1226,1225–1233,1226–1234,1263–1271,1264–1272. |
|
DRB5∗01:01 |
4–12,36–44,42–50,55–63,58–66,65–73,70–78,98–106,133–141,139–147,140–148,144–152,150–158,169–177,175– |
|
|
183,179–187,186–194,187–195,194–202,199–207,200–208,238–246,258–266,265–273,267–275,269–277,306–314,311– |
|
|
319,312–320,318–326,320–328,338–346,347–355,349–357,350–358,370–378,379–387,400–408,449–457,450–458,453– |
|
|
461,454–462,458–466,489–497,508–516,515–523,527–535,543–551,550–558,559–567,626–634,674–682,695–703,718– |
|
|
726,725–733,758–766,778–786,782–790,787–795,817–825,826–834,827–835,833–841,847–855,886–894,888–896,892– |
|
|
900,894–902,897–905,898–906,925–933,927–935,939–947,956–964,970–978,993–1001,994–1002,1007–1015,1011– |
|
|
1019,1020–1028,1032–1040,1046–1054,1065–1073,1075–1083,1094–1102,1095–1103,1103–1111,1141–1149,1208– |
|
|
1216,1217–1225,1264–1272. |
|
HLA-DQA10102- |
1–9,32–40,46–54,61–69,63–71,64–72,104–112,132–140,133–141,140–148,141–149,170–178,171–179,172–180,173– |
|
DQB10502 |
181,189–197,192–200,216–224,258–266,261–269,262–270,263–271,265–273,273–281,275–283,338–346,347–355,357– |
|
|
365,392–400,420–428,421–429,447–455,485–493,486–494,515–523,516–524,562–570,563–571,564–572,565–573,606– |
|
|
614,607–615,620–628,622–630,656–664,766–774,767–775,800–808,813–821,814–822,823–831,829–837,833–841,840– |
|
|
848,865–873,869–877,888–896,898–906,923–931,1011–1019,1044–1052,1046–1054,1047–1055,1061–1069,1098– |
|
|
1106,1099–1107,1106–1114,1132–1140,1144–1152,1145–1153,1192–1200,1193–1201,1202–1210,1214–1222,1216– |
|
|
1224,1217–1225,1220–1228. |
|
HLA-DQA10201- |
9–17,10–18,15–23,17–25,18–26,19–27,31–39,42–50,59–67,112–120,119–127,120–128,252–260,253–261,254–262,255– |
|
DQB10301 |
263,256–264,257–265,258–266,259–267,260–268,280–288,281–289,282–290,283–291,284–294,301–309,340–348,341– |
|
|
349,342–350,344–352,345–353,367–375,368–376,369–377,370–378,371–379,391–399,411–419,427–435,429–437,467– |
|
|
475,471–479,472–480,474–482,492–500,517–525,543–551,545–553,546–554,547–555,569–577,587–595,589–597,590– |
|
|
598,591–599,592–600,596–604,597–605,598–606,601–609,602–610,603–611,604–612,618–626,634–642,635–643,636– |
|
|
644,637–645,643–651,644–652,645–653,649–657,664–672,665–673,666–674,672–680,680–688,684–692,685–693,686– |
|
|
694,687–695,688–696,690–698,692–700,693–701,694–702,695–703,696–704,698–706,705–713,707–715,713–721,716– |
|
|
724,717–725,718–726,719–727,720–728,721–729,722–730,728–736,729–737,730–738,735–743,743–751,754–762,763– |
|
|
771,764–772,765–773,766–774,826–834,827–835,844–852,855–863,868–876,870–878,871–879,872–880,873–881,874– |
|
|
882,875–883,876–884,877–885,881–889,882–900,883–891,884–892,885–893,886–894,887–895,888–896,889–897,927– |
|
|
935,934–942,935–943,936–944,937–945,938–946,939–947,940–948,941–949,965–973,966–974,967–975,968–976,969– |
|
|
977,970–978,996–1004,997–1005,1002–1010,1008–1016,1012–1020,1017–1025,1018–1026,1019–1027,1020–1028,1021– |
|
|
1029,1022–1030,1026–1034,1027–1035,1062–1070,1072–1080,1074–1082,1075–1083,1076–1084,1096–1104,1114– |
|
|
1122,1115–1123,1116–1124,1120–1128,1121–1129,1122–1130,1169–1177,1170–1178,1218–1226,1220–1228,1227– |
|
|
1235,1230–1238,1235–1243,1236–1244,1237–1245,1247–1255. |
|
HLA-DQA10501- |
7–15,10–18,42–50,43–51,58–66,59–67,63–71,64–72,88–96,104–112,114–122,115–123,157–165,158–166,169–177,170– |
|
DQB10302 |
178,172–180,217–225,218–226,219–227,236–244,237–245,255–263,256–264,257–265,258–266,259–268,260–268,263– |
|
|
271,264–272,283–291,301–309,328–336,340–348,344–352,345–353,346–354,367–375,368–376,379–377,390–398,391– |
|
|
399,429–437,472–480,488–496,506–514,508–516,513–521,515–523,516–524,517–525,518–526,519–527,546–554,547– |
|
|
555,590–598,591–599,592–600,593–601,604–612,606–614,618–626,619–627,620–628,635–643,636–644,645–653,646– |
|
|
654,663–671,664–672,665–673,666–674,667–675,668–676,684–692,686–694,687–695,688–696,689–697,690–698,693– |
|
|
701,694–702,695–703,706–704,707–715,716–724,717–725,718–726,719–727,720–728,730–738,735–743,754–762,755– |
|
|
763,756–764,765–773,766–774,767–775,799–807,800–808,821–829,822–830,840–848,853–861,854–862,855–863,860– |
|
|
868,868–876,870–878,871–879,872–880,873–881,874–882,875–883,878–,885–,886–894,887–895,888–896,889–897,890– |
|
|
898,891–899,892–900,895–903,896–904,904–912,936–944,937–945,938–946,968–976,969–977,970–978,971–979,972– |
|
|
980,1001–1010,1009–1017,1010–1018,1017–1025,1018–1026,1019–1027,1024–1032,1045–1053,1046–1054,1048– |
|
|
1056,1049–1057,1055–1063,1061–1069,1062–1070,1063–1071,1064–1072,1074–1082,1075–1083,1076–1084,1110– |
|
|
1118,1131–1139,1169–1177,1170–1178,1217–1225,1218–1226,1221–1229,1235–1243,1236–1244. |
|
HLA-DQA10601- |
4–12,6–14,13–21,15–23,16–24,26–34,27–35,28–36,29–37,30–38,36–44,41–49,43–51,44–52,56–64,57–65,58–66,59–67,60– |
|
DQB10402 |
68,61–69,62–70,63–71,64–72,65–73,90–98,91–99,94–102,116–124,124–132,150–158,155–163,199–207,200–208,201– |
|
|
209,202–210,206–214,229–237,238–246,239–247,240–248,255–263,256–264,263–271,264–272,265–273,266–274,268– |
|
|
276,270–278,301–309,304–312,311–319,313–321,316–324,320–328,341–349,342–350,345–353,347–355,348–356,349– |
|
|
357,350–358,351–359,360–368,365–373,369–377,372–380,373–381,374–382,375–383,377–385,395–403,400–408,430– |
|
|
438,449–457,450–458,451–459,468–476,486–494,488–496,490–498,503–511,505–513,506–514,514–522,617–625,620– |
|
|
628,628–636,629–637,630–638,631–639,633–641,638–646,640–648,641–649,669–677,974–682,680–688,689–697,690– |
|
|
698,691–699,692–700,693–701,694–702,705–703,713–721,714–722,715–723,716–724,717–725,718–726,719–727,722– |
|
|
730,724–732,725–733,726–734,727–735,728–736,757–765,759–767,760–768,787–795,821–829,868–876,871–879,873– |
|
|
881,875–883,880–888,881–889,883–891,884–892,886–894,893–901,894–902,895–903,896–904,898–906,927–935,937– |
|
|
945,995–1003,1002–1010,1004–1012,1005–1013,1006–1014,1007–1015,1008–1016,1009–1017,1011–1019,1022– |
|
|
1030,1047–1055,1048–1056,1059–1067,1061–1069,1062–1070,1063–1071,1064–1072,1073–1081,1074–1082,1075– |
|
|
1083,1093–1101,1094–1102,1098–1106,1099–1107,1100–1108,1101–1109,1102–1110,1154–1162,1206–1214,1207– |
|
|
1215,1208–1216,1209–1217,1210–1218,1212–1220,1214–1222,1215–1223. |
|
HLA-DPA10103- |
1–9,2–10,3–11,4–12,5–13,31–39,32–40,42–50,50–58,51–59,52–60,53–61,54–62,58–66,59–67,78–86,100–108,104– |
|
DPB10601 |
112,127–135,128–136,130–138,132–140,133–141,134–142,135–143,139–147,140–148,167–175,168–176,170–178,188– |
|
|
196,189–197,233–241,237–245,238–246,264–272,265–273,273–281,288–296,387–495,390–398,391–399,395–403,429– |
|
|
437,448–456,449–457,450–458,451–459,456–464,489–497,492–500,507–515,508–516,509–517,510–518,511–519,512– |
|
|
520,513–521,514–522,538–546,557–567,581–589,749–857,754–762,800–808,815–823,816–824,817–825,818–826,855– |
|
|
863,1046–1054,1047–1055,1060–1068,1061–1069,1062–1070,1109–1117,1132–1140,1208–1210,1209–1217,1210– |
|
|
1218,1211–1219,1212–1220,1213–1221,1214–1222,1215–1223,1216–1224,1220–1228,1224–1232,1225–1233,1226– |
|
|
1234,1227–1235,1228–1236,1229–1237,1231–1239,1232–1240,1233–1241. |
|
HLA-DPA10201- |
2–10,4–12,40–48,41–49,42–50,48–56,51–59,52–60,53–61,54–62,55–63,58–66,59–67,78–86,102–110,110–118,112– |
|
DPB10101 |
120,133–141,138–146,139–147,140–148,152–160,167–175,168–176,169–177,170–178,186–194,188–196,189–197,193– |
|
|
201,200–208,233–241,235–243,237–245,238–246,262–270,269–277,302–310,312–320,340–348,341–349,342–350,369– |
|
|
377,392–400,449–457,450–458,453–461,505–513,508–516,509–517,510–518,511–519,512–520,556–564,557–565,558– |
|
|
566,559–567,577–585,579–587,581–589,719–727,725–733,797–805,814–822,815–823,816–824,817–825,818–826,820– |
|
|
828,821–829,822–830,833–841,853–861,854–862,870–878,893–901,915–923,962–970,996–1004,1000–1008,1059– |
|
|
1067,1062–1070,1107–1115,1148–1156,1195–1203,1208–1216,1209–1217,1213–1221,1214–1222,1217–1225,1220– |
|
|
1228,1264–1272,1265–1273. |
|
HLA-DPA10201- |
4–12,36–44,41–49,42–50,50–58,51–59,52–60,53–61,58–66,76–84,89–97,90–98,127–135,139–147,140–148,168–176,186– |
|
DPB10501 |
194,187–195,188–196,189–197,194–202,195–203,199–207,200–208,235–243,236–244,237–245,238–246,269–277,302– |
|
|
310,342–350,449–457,450–458,451–459,453–461,456–464,508–516,509–517,511–519,512–520,527–535,528–536,550– |
|
|
558,551–559,555–563,556–564,557–565,558–566,559–567,794–802,814–822,815–823,816–824,817–825,818–826,870– |
|
|
878,898–906,915–923,1012–1020,1149–1157,1203–1211,1214–1222,1215–1223,1264–1272. |
|
HLA-DPA10301- |
4–12,5–13,41–49,42–50,43–51,48–56,51–59,52–60,54–62,55–63,58–66,59–67,62–70,112–120,127–135,133–141,135– |
|
DPB10402 |
143,138–146,139–147,140–148,152–160,168–176,169–177,170–178,189–197,200–208,205–213,218–226,233–241,234– |
|
|
242,235–243,236–244,237–245,238–246,241–249,262–270,263–271,264–272,312–320,313–321,342–350,344–352,392– |
|
|
400,449–457,450–458,451–459,486–494,508–516,509–517,510–518,511–519,512–520,557–565,577–585,579–587,580– |
|
|
588,643–651,691–699,719–727,721–729,752–760,753–761,759–767,797–805,814–822,815–823,816–824,817–825,818– |
|
|
826,820–828,821–829,826–834,833–841,853–861,858–866,862–870,863–871,864–872,869–877,870–878,871–879,904– |
|
|
912,915–913,973–981,976–984,993–1001,996–1004,1005–1013,1006–1014,1007–1015,1057–1065,1058–1066,1060– |
|
|
1068,1062–1070,1214–1222,1264–1272. |
E protein
The E protein is the smallest of the major structural proteins in CoV which was identified as the structural component of the vi- rus. The E protein massively expressed virus envelope protein and plays an important role in virus membrane packaging [28]. The tertiary structure structure reveals that E protein of 2019-nCoV is a pentamer and contains multiple short α-helix (Figure 3B). Am- phipathic α-helix oligomerizes to form an ion-conductive pore in membranes [29]. BepiPred 1.0 server and COBEP were used to identify the potential antigen epitope regions of protein (Table 4).
Table 4: The predictions results of B-cell epitopes and T-cell epitopes of E protein.
|
Type |
Tools |
Location of the prediction results |
|
B-cell epitope prediction |
DNAStar |
3-10,52-70. |
|
CBTOPE |
3-8,52-63. |
|
|
BepiPred |
2-8,47-68. |
|
|
T-cell epitope prediction (HLA-DR) |
DRB1 01:01 |
10–18,11–19,26–34,31–39,36–44,44–52,49–57,50–58,57–65,62– 70. |
|
DRB3∗01:01 |
4–12,12–20,57–65. |
|
|
DRB4∗01:01 |
10–18,30–38,31–39. |
|
|
DRB5∗01:01 |
4–12,30–38,31–39,36–44,49–57,55–63,56–64,61–69. |
|
|
T-cell epitope prediction (HLA-DQ) |
HLA-DQA10102-DQB10502 |
1–9,17–25,20–28. |
|
HLA-DQA10201-DQB10301 |
6–14,7–15,8–16,14–22,24–32,27–35,28–36,29–37,30–38,32– 40,33–41,46–54. |
|
|
HLA-DQA10501-DQB10302 |
1–9,8–16,32–40,35–43,36–44. |
|
|
|
HLA-DQA10601-DQB10402 |
2–8,10–18,18–26,21–29,24–32,25–33,26–34,27–35,29–37,30– 38,32–40,33–41,40–48,42–50,45–53,52–60,54–62,55–63,56–64, 57–65. |
|
T-cell epitope prediction (HLA-DP) |
HLA-DPA10103-DPB10601 |
1–9,12–20,13–21,15–23,16–24,17–25,18–26,19–27,20–28,21– 29,22–30,23–31,24–32,25–33,26–34,27–35,28–36,29–37,31– 39,36–44,37–45,46–54,51–59. |
|
HLA-DPA10201-DPB10101 |
4–12,26–34,30–38,44–52,45–53. |
|
|
HLA-DPA10201-DPB10501 |
11–19,14–22,30–38,31–39,53–61,55–63,56–64,57–65. |
|
|
HLA-DPA10301-DPB10402 |
10–18,11–19,12–20,26–34,31–39,32–40,33–41,39–47,44–52,55– 63,57–65. |
Two B-cell epitopes in each monomer of E protein was identified through combining the two results of bioinformatic tools. The epi- topes were also shown in the primary and tertiary structure of the E protein (Figure 3). Two epitopes are located at the head and tail of the monomer of E protein and are regularly arranged at the lower and outer sides of the pentamer (Figure 3C and 3D). The residues of B-cell epitopes were summarized and extracted (Table 7).
Figure 3: B-cell epitopes distribution and structural features of E protein. (A) Sequences analysis of E protein from 2019-nCoV and SARS-CoV. Sequences of the B-cell epitopes were are highlighted by red background and named B1-B2. Residues marked “ * ” under the sequences, while these only conserved in the four first species were marked “ . or : ” under the sequences. (B) Protein structure of E protein. Five monomers are shown (purple, blue, red, green and yellow). (C) (D) B-cell epitopes on tertiary structure of E protein.
M protein
The M protein is the central organiser of CoV assembly and the most abundant structural protein , interacting with all other ma- jor coronaviral structural proteins [30]. [31]. Homotypic interac- tions between the M proteins are the major driving force behind virion envelope formation and defines the shape of the viral enve- lope [32]. The M protein composed of a long α-helices and three β-sheets (Figure 4). BepiPred 1.0 server and COBEP were used to identify the potential antigen epitope regions of protein (Table 5).
Figure 5: B-cell epitopes distribution of M protein of 2019-nCoV. (A) Sequences analysis of M protein from 2019-nCoV and SARS- CoV. Sequences of the B-cell epitopes were are highlighted by red background and named B1-B2. Residues marked “ * ” under the sequences, while these only conserved in the four first species were marked “ . or : ” under the sequences. (B) Protein structure of M protein. (C) B-cell epitopes on tertiary structure of M protein.
Table 6: The predictions results of B-cell epitopes and T-cell epitopes of N protein.
|
Type |
Tools |
Location of the prediction results |
|
B-cell epitope prediction |
DNAStar |
5-12,19-48,59-70,78-84,90-106,114-120,122-130,139-152,162-- |
|
|
|
215,227-264,274-300,323-329,334-349,357-389,400-405,414-418. |
|
|
CBTOPE |
1-8,23-30,43-46,54-56,70-73,79-86,100-111,145-147,164-178, 230- |
|
|
|
244, 266-300,310-327,355-383,404-410. |
|
|
BepiPred |
1-51,58-85,93-104,115-127,137-154,164-216,232-269,273-287, 291- |
|
|
|
301,306-310,323-331,338-347,361-387. |
|
T-cell epitope prediction (HLA-DR) |
DRB1∗01:01 |
52–60,53–61,110–118,111–119,130–138,132–140,157–165,159– |
|
|
|
167,165–173,215–223,219–227,222–230,227–235,314–322,330– |
|
|
|
338,331–339,349–357,351–359,360–368,392–400. |
|
|
DRB3∗01:01 |
86–94,100–108,125–133,213–221,221–229,222–230,266–274, |
|
|
|
306–314,307–315,345–353,351–359,355–363,374–382. |
|
|
DRB4∗01:01 |
130–138,153–161,154–162,155–163,156–164,157–165,217–225,218– |
|
|
|
226,219–227,220–228,221–229,222–230,303–311,348–352,349– |
|
|
|
357,350–358,351–359,392–400. |
|
|
DRB5∗01:01 |
6–14,28–36,32–40,52–60,53–61,85–93,86–94,87–95,94–102, |
|
|
|
110–118,157–165,171–179,177–185,181–189,183–191,187–195,249– |
|
|
|
257,253–261,254–262,258–266,268–276,291–299,292–300,307– |
|
|
|
315,311–319,314–322,330–338,331–339,349–357,353–361,379– |
|
|
|
387,392–400. |
|
T-cell epitope prediction (HLA-DQ) |
HLA-DQA10102- |
56–64,83–91,107–115,108–116,152–160,153–161,217–225,218– |
|
|
DQB10502 |
226,219–227,223–231,282–290,286–294,303–311,333–341,351– |
|
|
|
359,352–360,353–361,393–401,394–402,395–403. |
|
|
HLA-DQA10201- |
15–23,47–55,48–56,49–57,50–58,112–120,113–121,131–139, |
|
|
DQB10301 |
132–140,133–141,134–142,152–160,174–182,176–184,180–188,182– |
|
|
|
190,197–205,198–207,199–207.200–208,201–209,202–210,203– |
|
|
|
211,210–218,211–219,212–220,244–252,245–253,246–254,247– |
|
|
|
255,263–271,303–311,304–312,305–313,307–315,308–316,309– |
|
|
|
317,310–318,323–331,324–332,325–333,330–338,331–339,332– |
|
|
|
340,389–397,390–398,391–399,406–414,408–416,409–417,410– |
|
|
|
418,411–419. |
|
|
HLA-DQA10501- |
49–57,50–58,51–59,110–118,111–119,129–137,130–138,151– |
|
|
DQB10302 |
159,154–162,203–211,213–221,214–222,217–225,300–308,301– |
|
|
|
309,304–312,305–313,306–314,307–315,308–316,309–317,310– |
|
|
|
318,313–321,314–322,315–323,317–325,318–326,324–332,332– |
|
|
|
340,333–341,388–396,389–397,390–398,391–399,392–400,393– |
|
|
|
401,394–402,407–415,408–416. |
|
|
HLA-DQA10601- |
48–56,49–57,50–58,51–59,52–60,53–61,54–62,83–91,84–92,85– |
|
|
DQB10402 |
93,86–94,87–95,106–114,108–116,111–119,130–138,169–177, |
|
|
|
244–252,263–271,268–276,269–277,298–306,305–313,311–319,313– |
|
|
|
321,314–322,322–330,327–335,328–336,329–337,330–338,331– |
|
|
|
339,390–398. |
|
T-cell epitope prediction (HLA-DP) |
HLA-DPA10103- |
107–115,108–116,109–117,110–118,217–225,218–226,219–227. |
|
|
DPB10601 |
|
|
|
HLA-DPA10201- |
49–57,50–58,105–113,155–163,216–224,217–225,218–226,219– |
|
|
DPB10101 |
227,220–228,221–229,222–230,345–353,346–354,387–395. |
|
|
HLA-DPA10201- |
105–113,106–114,218–226,219–227,220–228,221–229,311–319, |
|
|
DPB10501 |
330–338,331–339,346–354,349–357,387–395. |
|
|
HLA-DPA10301- |
49–57,218–226,219–227,220–228,221–229,222–230,317–325, |
|
|
DPB10402 |
331–339,387–395. |
The final 7 potential B-cell epitopes were determined through in- tegrating the results from bioinformatic tools. The epitopes were also shown in the primary and tertiary structure of the N protein (Figure 5). The NTD and CTD has 3 and 4 B-cell epitopes, respec- tively (Figure 5C and 5D). The residues of B-cell epitopes were summarized and extracted (Table 7). In short, the structure and immunogenicity of 2019-nCoV struc- tural protein were analyzed by bioinformatics tools. Thirty-three B-cell epitopes of four structural proteins were determined in which include S protein (22), N protein (7), M protein (2), and E protein (2), respectively (Figure 6A). The B-cell epitopes residues of four structural proteins were summarized and extracted (Table 7).
Table 7: Predicted B-cell epitopes of 2019-nCoV structural proteins.
|
Protein |
Code |
Position |
Sequence |
|
S protein |
B1 |
21-30 |
RTQLPPAYTN |
|
B2 |
71-77 |
SGTNGTK |
|
|
B3 |
110-113 |
LDSK |
|
|
B4 |
251-258 |
PGDSSSGW |
|
|
B5 |
280-286 |
GTITD |
|
|
B6 |
315-325 |
TSNFRVQPTES |
|
|
B7 |
382-387 |
VSPTKL |
|
|
B8 |
408-415 |
RQIAPGQT |
|
|
B9 |
439-447 |
NNLDSKVGG |
|
|
B10 |
471-484 |
EIYQAGSTPCNGVE |
|
|
B11 |
526-532 |
GPKKSTN |
|
|
B12 |
551-557 |
VLTESNK |
|
|
B13 |
601-607 |
GTNTSNQ |
|
|
B14 |
636-643 |
YSTGSNVF |
|
|
B15 |
656-659 |
VNNS |
|
|
B16 |
675-687 |
QTQTNSPRRARSV |
|
|
B17 |
773-780 |
EQDKNTQE |
|
|
B18 |
789-797 |
YKTPPIKDF |
|
|
B19 |
805-816 |
ILPDPSKPSKRS |
|
|
B20 |
936-941 |
DSLSST |
|
|
B21 |
1080-1088 |
AICHDGKAH |
|
|
B22 |
1114-1122 |
IITTDNTFV |
|
|
N protein |
B1 |
43-46 |
QGLP |
|
B2 |
100-104 |
KMKDL |
|
|
B3 |
164-178 |
GTTLPKGFYAEGSRG |
|
|
B4 |
275-283 |
GRRGPEQTQ |
|
|
B5 |
296-300 |
TDYKH |
|
|
B6 |
323-329 |
EVTPSGT |
|
|
B7 |
361-365 |
KTFPP |
|
M protein |
B1 |
171-176 |
ATSRTL |
|
B2 |
185-194 |
QRVAGDSGFA |
|
|
E protein |
B1 |
3-8 |
SFVSEE |
|
B2 |
52-63 |
VKPSFYVYSRVK |
The proportion of epitope residues in primary sequence was cal- culated. The proportion of epitope residues in primary sequence was used to determine the antigenicity of proteins. The E protein has the largest antigenicity in which residue coverage of B-cell epitopes is 24% (Figure 6B). The E protein might be the potential targets to design effective 2019-nCoV vaccines and facilitate the development of rapid diagnostic methods in the future.
T-cell epitopes prediction of structural protein
T-cell immune response plays a very important role in persistent viral infection [35]. Cellular immune response impact disease pro- gression and is essential factor for the viral clearance from the pa- tients. T-cells immune response usually has been observed within 4–5 days after after a viral infection [36]. Antigenic peptides of the coronavirus protein can be recognised by T-cells on the sur- face of infected cells [37]. The structure of the MHC-I molecule HLAA*1101 in complex with such a peptide derived from the SARS-CoV protein has recently been determined [38]. T-cell epi- tope prediction is for identifying the shortest peptides of allergen that bind to the MHC complexes. T-cell epitopes are principally predicted on the basis of identifying the binding of amino acid fragments to the MHC complexes that can activate T-cells.
The binding strength of each peptide to the given MHC is esti- mated by NetMHCII 2.2 at a set threshold level. Given the MHC alleles have tens of thousands of kinds, in order to ensure the rep- resentative and reliability prediction results, we selected the most common MHC in the population to predict their peptide binding activity with protein, including HLA-DR, HLA-DQ, and HLA-DP. The predicted results of the four structural proteins are listed in Table 3-6. The number of MHC-binding peptides of the four struc- tural proteins predicted by different alleles is counted in Table 8.
Table 8: The number of MHC-binding peptides of different alleles prediction
|
Type |
Tools |
S protein |
N protein |
M protein |
E protein |
|
T-cell epitope prediction (HLA-DR) |
DRB1∗01:01 |
119 |
20 |
35 |
10 |
|
DRB3∗01:01 |
114 |
13 |
21 |
3 |
|
|
DRB4∗01:01 |
84 |
18 |
25 |
3 |
|
|
DRB5∗01:01 |
91 |
32 |
32 |
8 |
|
|
T-cell epitope prediction (HLA-DQ) |
HLA-DQA10102-DQB10502 |
77 |
20 |
17 |
3 |
|
HLA-DQA10201-DQB10301 |
182 |
52 |
29 |
12 |
|
|
HLA-DQA10501-DQB10302 |
162 |
39 |
15 |
5 |
|
|
HLA-DQA10601-DQB10402 |
174 |
33 |
57 |
20 |
|
|
T-cell epitope prediction (HLA-DP) |
HLA-DPA10103-DPB10601 |
96 |
7 |
52 |
23 |
|
HLA-DPA10201-DPB10101 |
93 |
14 |
34 |
5 |
|
|
HLA-DPA10201-DPB10501 |
64 |
12 |
36 |
8 |
|
|
HLA-DPA10301-DPB10402 |
96 |
9 |
34 |
11
|
The peptide will possess stronger immunogenicity in case it binds more alleles. The number of MHC-binding peptides of different allele coverage is counted in Table 9.
Table 9: The number of MHC-binding peptides of different allele coverage
|
Allele coverage |
S protein |
N protein |
M protein |
E protein |
||||||||
|
DR |
DQ |
DP |
DR |
DQ |
DP |
DR |
DQ |
DP |
DR |
DQ |
DP |
|
|
100% |
0 |
0 |
18 |
0 |
0 |
2 |
0 |
0 |
5 |
0 |
0 |
0 |
|
75% |
17 |
24 |
20 |
5 |
4 |
3 |
6 |
1 |
18 |
1 |
1 |
2 |
|
50% |
62 |
113 |
44 |
11 |
29 |
6 |
23 |
17 |
18 |
6 |
7 |
6 |
|
25% |
234 |
294 |
129 |
46 |
74 |
13 |
49 |
81 |
46 |
9 |
23 |
29 |
The final T-cell epitopes was determined through the allele cov- erage of more than 70%. The final T-cell epitopes included 39 sequences through combining the results of the three alleles, in which include spike protein (25), nucleocapsid protein (6), mem- brane protein (7), and envelope protein (1), respectively (Figure 6A). The residues of T-cell epitopes were summarized and extract- ed (Table 10).
Table 10: Predicted T-cell epitopes of 2019-nCoV structural protein.
|
Protein |
Code |
Position |
Sequence |
|
S protein |
T1 |
4-12 |
FLVLLPLVSS |
|
T2 |
41-76 |
KVFRSSVLHSTQDLFLPFFSNVTWFHAIHVSGTNGT |
|
|
T3 |
98-106 |
SNIIRGWIF |
|
|
T4 |
127-148 |
VIKVCEFQFCNDPFLGVYYHKN |
|
|
T5 |
168-178 |
FEYVSQPFLMD |
|
|
T6 |
186-197 |
FKNLREFVFKNI |
|
|
T7 |
233-246 |
INITRFQTLLALHR |
|
|
T8 |
255-277 |
SSGWTAGAAAYYVGYLQPRTFLL |
|
|
T9 |
301-326 |
CTLKSFTVEKGIYQTSNFRVQPTESI |
|
|
T10 |
342-353 |
FNATRFASVYAW |
|
|
T11 |
449-458 |
YNYLYRLFRK |
|
|
T12 |
489-497 |
YFPLQSYGF |
|
|
T13 |
508-520 |
YRVVVLSFELLHA |
|
|
T14 |
557-565 |
KKFLPFQQF |
|
|
T15 |
620-628 |
VPVAIHADQ |
|
|
T16 |
693-702 |
IAYTMSLGAE |
|
|
T17 |
716-727 |
TNFTISVTTEIL |
|
|
T18 |
766-774 |
ALTGIAVEQ |
|
|
T19 |
814-841 |
KRSFIEDLLFNKVTLADAGFIKQYGDCL |
|
|
T20 |
871-906 |
AQYTSALLAGTITSGWTFGAGAALQIPFAMQMAYRF |
|
|
T21 |
915-923 |
VLYENQKLI |
|
|
T22 |
993-1001 |
IDRLITGRL |
|
|
T23 |
1007-1015 |
YVTQQLIRA |
|
|
T24 |
1062-1083 |
FLHVTYVPAQEKNFTTAPAICH |
|
|
T25 |
1214-1222 |
WYIWLGFIA |
|
|
N protein |
T1 |
49-58 |
TASWFTALTQ |
|
T2 |
157-165 |
IVLQLPQGT |
|
|
T3 |
218-230 |
ALALLLLDRLNQL |
|
|
T4 |
305-313 |
AQFAPSA |
|
|
T5 |
351-359 |
ILLNKHIDA |
|
|
T6 |
387-400 |
KKQQTVTLLPAADL |
|
|
M protein |
T1 |
15-73 |
KLLEQWNLVIGFLFLTWICLLQFAYANRNRFLYIIKLIFLWLLWPVT- LACFVLAAVYRI |
|
T2 |
89-108 |
GLMWLSYFIASFRLFARTRS |
|
|
T3 |
117-125 |
NILLNVPLH |
|
|
T4 |
132-140 |
PLLESELVI |
|
|
T5 |
148-156 |
HLRIAGHHL |
|
|
T6 |
179-187 |
YKLGASQRV |
|
|
T7 |
193-201 |
FAAYSRYRI |
|
|
E protein |
T1 |
26-40 |
FLLVTLAILTALRLC |
The proportion of epitope residues in primary sequence was calculated (Figure 6B). The proportion of epitope residues in primary sequence was used to determine immunogenicity of proteins. The membrane protein has the largest immunogenicity in which residue coverage of T-cell epitopes is 55.86% (Figure 6B).
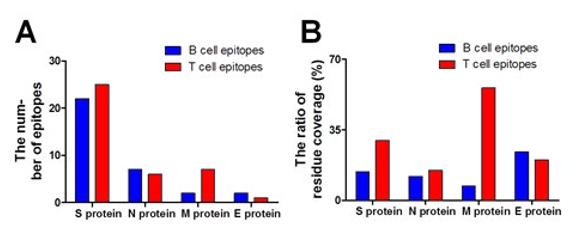
Figure 6: The number and residue coverage ratio of B- and T-cell epitopes of structural proteins. (A) The number and of B- and T-cell epitopes of structural proteins. (B) The residue coverage ratio of B- and T-cell epitopes of structural proteins.
The reason that immune storm was caused by 2019-nCoV maybe that the membrane and envelope protein expressed plentifully in cell infected. The knowledge of antigenic or epitopic sites of viral protein is important for the development of effective antiviral in- hibitors or vaccine. For effective vaccine, both T- and B-cells epi- topes identification is very important for prevention or clearance of infection [39].
These results indicate the importance of this region for the de- velopment of effective vaccine. Identified B-cell epitope can be utilized in the development of the effective 2019-nCoV vaccine because it is a vital step for development of epitope-based vaccines and diagnostic tools. Understanding the structural protein structure and function could possibly find therapeutic targets to prevent and control the coronaviruses related diseases. Therefore M protein and E protein was chosen as a potential antigenic target for the hu- moral immune responses, which might be significant for develop- ing better diagnostic and research reagents in the future. Thus our study suggests that E protein could possibly be a good candidate for B cell-line epitopes in preparing monoclonal antibodies, vac- cines and anti-viral inhibitors against CoV infection in the future.
Conclusion
In the present study, using bioinformatics methods, we analyzed the four structural proteins of 2019-nCoV and identified a potential T-cell epitope and B-cell epitope of the structural proteins, which might significantly improve our current 2019-nCoV vaccine de- velopment strategies. These predicted epitopes can be used as po- tential targets for the development of the effective vaccine against 2019-nCoV.
Author Contributions
Y.L. wrote the main manuscript and performed the experiments presented in Figure 1–5 and analyzed the data. M.M. performed the experiments presented in Table 1-3 and analyzed the data. Z.L., N.Z., X.S., and L.Y. designed the project, supervised the experi- ments and wrote the manuscript. All authors reviewed the manu- script.
Funding
This study was supported by grants from the National Natural Sci- ence Foundation of China (No. 81901684), Shenzhen Science and Technology Plan Project (No. JSGG20200225153031960)..
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to in- fluence the work reported in this paper.
References
- Hui, D. S., Azhar, E. I., Madani, T. A., Ntoumi, F., Kock, R., Dar, O., ... & Petersen, E. (2020). The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—The latest 2019 novel coronavirus outbreak in Wuhan, China. In- ternational journal of infectious diseases, 91, 264-266.
- Liu, J., Zheng, X., Tong, Q., Li, W., Wang, B., Sutter, K., ... & Yang, D. (2020). Overlapping and discrete aspects of the pathology and pathogenesis of the emerging human pathogen- ic coronaviruses SARS-CoV, MERS-CoV, and 2019-nCoV. Journal of medical virology, 92(5), 491-494.
- World Health Organization, & World Health Organization. (2020). Novel Coronavirus (2019-nCoV). Situation Report, 13(2).
- Tian, X., Li, C., Huang, A., Xia, S., Lu, S., Shi, Z., ... & Ying,T. (2020). Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerging microbes & infections, 9(1), 382-385.
- Rappuoli, R., Pizza, M., Del Giudice, G., & De Gregorio,E. (2014). Vaccines, new opportunities for a new society. Proceedings of the National Academy of Sciences, 111(34), 12288-12293.
- Weiss, G. A., Watanabe, C. K., Zhong, A., Goddard, A., & Sidhu, S. S. (2000). Rapid mapping of protein functional epi- topes by combinatorial alanine scanning. Proceedings of the National Academy of Sciences, 97(16), 8950-8954.
- Hua, C. K., Gacerez, A. T., Sentman, C. L., Ackerman, M. E., Choi, Y., & Bailey-Kellogg, C. (2017). Computationally-driv- en identification of antibody epitopes. Elife, 6, e29023.
- Li, Y., Wang, Y., Ran, P., Yang, P., & Liu, Z. (2019). IgE bind- ing activities and in silico epitope prediction of Der f 32 in Dermatophagoides farinae. Immunology Letters, 213, 46-54.
- Bartlam, M., Yang, H., & Rao, Z. (2005). Structural insights into SARS coronavirus proteins. Current opinion in structural biology, 15(6), 664-672.
- Van Regenmortel, M. H. V. (2000). Family flaviviridae. Virus Taxonomy: Seventh Report of the International Committee on Taxonomy of Viruses, 859-878.
- Chang, C. K., Hou, M. H., Chang, C. F., Hsiao, C. D., & Huang, T. H. (2014). The SARS coronavirus nucleocapsid protein–forms and functions. Antiviral research, 103, 39-50.
- Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F., & Higgins, D. G. (1997). The CLUSTAL_X windows inter- face: flexible strategies for multiple sequence alignment aid- ed by quality analysis tools. Nucleic acids research, 25(24), 4876-4882.
- Teng, F., Sun, J., Yu, L., Li, Q., & Cui, Y. (2018). Homology modeling and epitope prediction of Der f 33. Brazilian Journal of Medical and Biological Research, 51.
- Kiefer, F., Arnold, K., Künzli, M., Bordoli, L., & Schwede,T. (2009). The SWISS-MODEL Repository and associated resources. Nucleic acids research, 37(suppl_1), D387-D392.
- Eswar, N., Eramian, D., Webb, B., Shen, M. Y., & Sali, A. (2008). Protein structure modeling with MODELLER. In Structural proteomics (pp. 145-159). Humana Press.
- Ramachandran, S., Kota, P., Ding, F., & Dokholyan, N. V. (2011). Automated minimization of steric clashes in protein structures. Proteins: Structure, Function, and Bioinformatics, 79(1), 261-270.
- Van Regenmortel, M. H. V., & De Marcillac, G. D. (1988). An assessment of prediction methods for locating continuous epitopes in proteins. Immunology letters, 17(2), 95-107.
- Zheng, L. N., Lin, H., Pawar, R., Li, Z. X., & Li, M. H. (2011).Mapping IgE binding epitopes of major shrimp (Penaeus monodon) allergen with immunoinformatics tools. Food and Chemical Toxicology, 49(11), 2954-2960.
- Xie, Q., He, X., Yang, F., Liu, X., Li, Y., Liu, Y., ... & Zhao,W. (2017). Analysis of the genome sequence and prediction of B-cell epitopes of the envelope protein of Middle East re- spiratory syndrome-coronavirus. IEEE/ACM transactions on computational biology and bioinformatics, 15(4), 1344-1350.
- Wang, H. W., & Pai, T. W. (2014). Machine learning-based methods for prediction of linear B-cell epitopes. Immunoin- formatics, 217-236.
- Yang, H., Chen, H., Jin, M., Xie, H., He, S., & Wei, J. F.(2016). Molecular cloning, expression, IgE binding activi- ties and in silico epitope prediction of Per a 9 allergens of the American cockroach. International Journal of Molecular Medicine, 38(6), 1795-1805.
- Yang, X., & Yu, X. (2009). An introduction to epitope pre- diction methods and software. Reviews in medical virology, 19(2), 77-96.
- Gasteiger, E., Hoogland, C., Gattiker, A., Wilkins, M. R., Ap- pel, R. D., & Bairoch, A. (2005). Protein identification and analysis tools on the ExPASy server. The proteomics proto- cols handbook, 571-607.
- Ziebuhr, J. (2004). Molecular biology of severe acute respira- tory syndrome coronavirus. Current opinion in microbiology, 7(4), 412-419.
- Ou, X., Guan, H., Qin, B., Mu, Z., Wojdyla, J. A., Wang, M.,... & Cui, S. (2017). Crystal structure of the receptor binding domain of the spike glycoprotein of human betacoronavirus HKU1. Nature communications, 8(1), 1-10.
- Wong, A., Gehring, C., & Irving, H. R. (2015). Conserved functional motifs and homology modeling to predict hidden moonlighting functional sites. Frontiers in Bioengineering and Biotechnology, 3, 82.
- Hopp, T. P., & Woods, K. R. (1981). Prediction of protein an- tigenic determinants from amino acid sequences. Proceedings of the National Academy of Sciences, 78(6), 3824-3828.
- Raamsman, M. J., Locker, J. K., De Hooge, A., De Vries, A. A., Griffiths, G., Vennema, H., & Rottier, P. J. (2000). Charac- terization of the coronavirus mouse hepatitis virus strain A59 small membrane protein E. Journal of virology, 74(5), 2333- 2342.
- Verdiá-Báguena, C., Nieto-Torres, J. L., Alcaraz, A., DeDi- ego, M. L., Enjuanes, L., & Aguilella, V. M. (2013). Anal- ysis of SARS-CoV E protein ion channel activity by tuning the protein and lipid charge. Biochimica et Biophysica Acta (BBA)-Biomembranes, 1828(9), 2026-2031.
- Neuman, B. W., Kiss, G., Kunding, A. H., Bhella, D., Baksh,M. F., Connelly, S., ... & Buchmeier, M. J. (2011). A structural analysis of M protein in coronavirus assembly and morpholo- gy. Journal of structural biology, 174(1), 11-22.
- Masters, P. S. (2006). The molecular biology of coronavirus- es. Advances in virus research, 66, 193-292.
- De Haan, C. A., Vennema, H., & Rottier, P. J. (2000). Assem- bly of the coronavirus envelope: homotypic interactions be- tween the M proteins. Journal of virology, 74(11), 4967-4978.
- de Haan, C. A., & Rottier, P. J. (2005). Molecular interactions in the assembly of coronaviruses. Advances in virus research, 64, 165-230.
- Jayaram, H., Fan, H., Bowman, B. R., Ooi, A., Jayaram, J., Collisson, E. W., ... & Prasad, B. V. (2006). X-ray structures of the N-and C-terminal domains of a coronavirus nucleocapsid protein: implications for nucleocapsid formation. Journal of virology, 80(13), 6612-6620
- Claassen, M. A., Janssen, H. L., & Boonstra, A. (2013). Role of T cell immunity in hepatitis C virus infections. Current Opinion in Virology, 3(4), 461-467.
- Rehermann, B. (2009). Hepatitis C virus versus innate and adaptive immune responses: a tale of coevolution and coex- istence. The Journal of clinical investigation, 119(7), 1745- 1754.
- Boots, A. M., Kusters, J. G., Van Noort, J. M., Zwaagstra,K. A., Rijke, E., van der Zeijst, B. A., & Hensen, E. J. (1991). Lo- calization of a T-cell epitope within the nucleocapsid protein of avian coronavirus. Immunology, 74(1), 8.
- Blicher, T., Kastrup, J. S., Buus, S., & Gajhede, M. (2005). High-resolution structure of HLA-A* 1101 in complex with SARS nucleocapsid peptide. Acta Crystallographica Section D: Biological Crystallography, 61(8), 1031-1040.
- Fournillier, A., Wychowski, C., Boucreux, D., Baumert, T. F., Meunier, J. C., Jacobs, D., ... & Inchauspe, G. (2001). Induc- tion of hepatitis C virus E1 envelope protein-specific immune response can be enhanced by mutation of N-glycosylation sites. Journal of virology, 75(24), 12088-12097.


