Research Article - (2024) Volume 2, Issue 2
Screening of Rice Genotypes for Salinity Stress Tolerance
2Department of Rice, Tamil Nadu Agricultural University, Coimbatore, India
3Agricultural Officer, Vadipatti block, Madurai, India
4ICAR - Krishi Vigyan Kendra, Virudhunagar, Tamil Nadu, India
Received Date: Sep 02, 2024 / Accepted Date: Sep 20, 2024 / Published Date: Oct 07, 2024
Copyright: ©©2024 Chelvi Ramesh, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Surendar, K. K., Sritharan, N., Sathiyavani, E., Kumar, S. K., Venudevan, B., et al. (2024). Screening of Rice Genotypes for Salinity Stress Tolerance. Int J Bot Hor Res, 2(2), 01-14.
Abstract
Experiments were conducted at Unit of Crop Physiology, Department of Seed Science and Technology, Agricultural College and Research Institute, Madurai to study the effect of salinity on growth of various genotypes of rice. The varieties (ASD 16 MDU 6 and ADT 45) and landraces (Kuliyadichan, Kallurundaikar and Norungan) were taken for this study. Germination and seedling characters were measured at seventh day after inducing abiotic stress conditions such as, salinity by application of NaCl at 200mM concentration. Physiological and biochemical parameters were estimated at the twentieth day after inducing the salinity stress conditions. The number of seeds germinated, shoot and root length, vigor index, nitrate reductase, α-amylase content and the soluble protein content were studied under the laboratory conditions in comparison to the control. Analyses of variance, revealed significant differences for germination percentage, shoot and root length, vigor index, nitrate reductase, α-amylase and protein contents under salinity. It was evident from the data that increased levels of sodium chloride decreased the germination percent, shoot and root lengths and hence the vigor index nitrate reductase, α- amylase and protein contents linearly.
Keywords
Salinity Stress, Rice Varieties, Morphology, Physiological Characters
Introduction
Rice is the seed of the grass species Oryza sativa (Asian rice) or Oryzaglaberrima(African rice). As a cereal grain, rice, a monocot, is normally grown as an annual plant, although in tropical areas it can survive as a perennial and can produce a ratoon crop for up to 30 years. The rice plant can grow to 1 to 1.8 m tall, occasionally more depending on the variety and soil fertility. It has long, slender leaves 50 to 100 cm long and 2 to 2.5 cm broad. The small wind-pollinated flowers are produced in a branched arching to pendulous inflorescence 30 to 50 cm long. The edible seed is a grain, a fruit called caryopsis 5 to 12 mm long and 2 to 3mmthick. Rice cultivation is well-suited to countries and regions with low labor costs and high rainfall, as it is labor intensive to cultivate and requires ample water. However, rice can be grown practically anywhere, even on a steep hill or mountain area with the use of water- controlling terrace systems. The traditional method for cultivating rice is flooding the fields while or after setting the young seedlings. This simple method requires sound planning and servicing of the water damming and channeling, but reduces the growth of less robust weed and pest plants that have no submerged growth state, and deters vermin. While flooding is not mandatory for the cultivation of rice, all other methods of irrigation require higher effort in weed and pest control during growth periods and a different approach for fertilizing the soil. Water management : Uniform leveling of field and proper drainage are most essential for an effective water management in irrigated field. Efficient water-management facilitates, good tillering and better nutrient uptake and helps in reducing weed population. In India rice is grown in 43.86 million ha, the production level is 104.80 million tones and the productivity is about 2390 kg ha-1 (Agricultural Statistics at a glance, 2015). The rice production has registered an appreciable increase from 20.58 million tons in 1950-51to 104.86 million tons during 2014-15, which is nearly 5 times. The yield was 668 kg ha-1in 1950-51which has increased to 2390 kg ha-1 during 2014-15. Major share of rice production is in Kharif season. It is grown in almost all the states in the country however the major 5 states in rice production are West Bengal, UP, Andhra Pradesh, Punjab and Tamil Nadu.
It is one of the most important food crops and feeds more than 60 per cent population of India. Though the area in production of rice is high compared to that of china, the production and productivity of rice is low in India. And also the productivity of countries like Indonesia, Bangladesh and Vietnam is high to that of India. On the whole, Indian agriculture does not show high efficiency or productivity, though there is an improvement since independence. The reasons and constrains of the above said results may have connections like., population pressure, uneconomic holdings, uncertain monsoons and inadequate irrigation facilities, subsistence nature of farming, decline in soil fertility, lack of support services, poor organization of resources and lack of entrepreneurship and also poor crop plant population in case of broadcast sowing method resulting in uneven germination (upland and direct seeded lowland). Delay in monsoon onset often results in delayed and prolonged transplanting and sub-optimum plant population (mostly in rainfed low lands). Therefore, the productivity is low. In the high rainfed regions, the rain water is lost rapidly through deep percolation, because of the upland location and loose texture of the soil. In these soils the plant nutrient applied through fertilizers are lost rapidly and investment of fertilizer become risky. Further, low water retention capacity by the soil due to high permeability brings in moisture stress conditions quickly after cessation of rains. Such situation contributes low productivity. In the low rainfall regions, the crops suffer from iron and zinc deficiency in some soils. Among the above said problems and constrains, the increasing areas under salinity are the major problems leading to severe yield reduction. Abiotic stress, which includes salinity, heat and cold, critically threatens crop production and causes significant yield loss in large areas [1,2]. Among these, soil salinity is the second major environmental constraint, to crop production and is expected to increase due to global climate changes and as a consequence of many irrigation practices and depleting water resources respectively. Plant growth and developmental processes in terms of biochemical, physiological and morphological characteristics are inhibited by both water deficit and salt stresses [3-5].
Salinity is a common abiotic stress that severely limits crop growth and development, productivity and causes the continuous loss of arable land, which results in desertification in arid and semi-arid regions of the world [6]. It is estimated that more than 800 million hectares of land throughout the world are adversely affected by high salinity [7]. Ali et al opined that saline soils are characterized by excess of sodium ions with dominant anions of chloride and sulfate resulting in higher electrical conductivity (>4 dS m1) [8]. In general, salinity stress induces an initial osmotic stress and subsequent toxicity as a consequence of the accumulation of ions. However, damage can also ensue as a result of excessive reactive oxygen species (ROS) such as superoxide radicals (O2), hydrogen peroxide (H2O2) and hydroxyl radicals (OH) produced at a high rate commonly accumulated in plant tissues due to ion imbalance and hyper osmotic stresses. ROS accumulation leads to lipid oxidation and has a negative effect on cellular metabolism and physiology, thus adversely ruining the membrane integrity [9]. Salinity tolerance in glycophytic crops including rice is predominantly associated with the maintenance of ion homeostasis, particularly low Na+/K+ or high K+/Na+ ratios, through exclusion, compartmentation, and partitioning of Na+ [10]. In addition to ion homeostasis strategies, many plants have evolved mechanisms to regulate the synthesis and accumulation of compatible solutes such as proline and glycine betaine, which function as osmoprotectants and have a crucial role in plant adaptation to osmotic stress through stabilization of the tertiary structure of proteins [5]. Rice is the most important global food crop that feeds over half of the world population and still getting a major proportion of their energy requirement from rice and its derived products with the demand for food expected to increase by another 38% within 30 years [11]. However, rice productivity in many areas is affected by salinity stress, which originates from the accumulation of underground salt and is exacerbated by salt mining, deforestation and irrigation [12]. Rice is generally characterized as a salt sensitive crop but the extent of its sensitivity varies during different growth and developmental stages. It is tolerant to salinity stress during germination and active tillering, whereas it displays more sensitivity during early vegetative and reproductive stages [13].
Attempts to measure the tolerance capacity in a genotypes or hybrid with single parameter have limited value because of the multiplicity of the factors and their interactions contributing to salinity tolerance. Different researchers used different traits to appraise genetic variances in salinity tolerance. Hence, in the present study, rice genotypes and hybrids have been selected to assess their tolerant capacity under salinity and the various traits contributing to salinity tolerance during germination and seedling establishment stage of the crop with the following objectives viz., To investigate the germination, physiological and biochemical responses of rice genotypes to salinity stress at germination level and To screen the rice genotypes that are tolerant and perform good to the induced salinity stress conditions.
Materials and Methods
A laboratory study was conducted in the Unit of Crop Physiology, Agricultural College and Research Institute, Madurai with the objective to screen the rice genotypes for salinity stress tolerance at seed germination level of selected rice genotypes. A brief account of the materials used and methodologies followed in the present study are presented in this chapter. Three varieties and three landraces were choosen for this experiments with two treatments T1-Control (Water only used) and T2 – NaCl @200mM withfour replications.
Varietal Details
The experiment was conducted with six rice genotypes. It includes 3 landraces ( Kuliyadichan, Kallurundaikar, NORUNGAN) ,3 varieties ( ADT 45, MDU6, ASD16) .The seeds for the experiment were obtained from Department of plant breeding & genetics , Agricultural College and Research Institute, Madurai.
Methodology
The experiment was laid out under laboratory with 6 rice genotypes with 4 replications. NaCl stress treatment was imposed. The standardized protocols using NaCl stress (200mM equivalent to -1.26 Mpa water potential). Change solutions once in every two days for NaCl stress. Drain completely rinses three to four times with fresh solutions if possible so as to avoid increased stress level due to NaCl. Experiment was continued at lab for one month and seedling vigour, germination and other related parameters were observed against control.
Screening for Salinity Tolerance
• Screening for salinity tolerance need to be carried out hydropon-ically at early vegetative stage of the plants that is when the plant reaches 3 to 4 leaf stages
• For this floating styrofoam panel as shown in figure need to be prepared as per the size of the tray using thermocool sheet, mos-quito nets and Adhesive. The styroform panel should have 6* 4 holes
• Individual accession need to be grown as single line along with standard check lines and 6 replications
• The tray need to be filled with a nutrient solution comprising of both macro and micronutrients and should have pH of 5.5
• Preparation of stock solution Preparation of working solution
• During the experiment the solution need to be checked every day for maintaining the pH of the solution if the pH shifts to far then it is better to replace the solution Beforestartingtheexperimenttheseedsofeachgenotypesneedstobep-reheatedinhot air oven for 3 to 5 days at 50 degree centigrade to break seed dormancy( if any)
• The surface sterilized seeds need to be replaced in dishes with moistured filter papers and incubator at 30 degree centigrade for 48 hours to germinate
• These pre-germinated seeds should be placed in the individual holes of the styrofoam panel two seedlings per hole • Initially the Seedling should be kept in normal water for 2 to 3 days.
• After that it need it to be kept in nutrient solution
• After 5 days of growth in nutrients solution salinity stress need to be imposed in one set of the .Other set would be kept as search for control
• Salt stress need to be imposed as 6ds m-1 NaCl solution (approx¬imately 60 mM NaCl i. e. 3.0-gram NaCl per liter solution would be required level of EC) for initial 2 days. After 2 days it should be increases to 12 ds m-1 NaCl solution( approximately 120 mm NaCl i.e. 6.0-gram NaCl per liter solution)
• Visual scoring of genotype should be started as soon as the ap¬pearance of the saltspecific symptoms. Scoring should be contin¬ued until 60% of plants of most suspectable genotypes reaches the score of "9" .Genotype should be ranked based on the final scoring at the stage
Preparation of Stock Solution
|
Element (macronutrient) |
Reagent(ARgrade) |
Preparation(g/250mlsolution) |
|
N |
Ammoniumnitrate(NH4NO3) |
22.85 |
|
P |
Sodiumphosphate,monobasicmonohydrate (NaH2PO4.H2O) |
8.9 |
|
K |
Potassiumsulphate(K2SO4) |
17.85 |
|
Ca |
Calciumchloride,Dihydrate(CaCl2.2H2O) |
29.3 |
|
Mg |
Magnesium sulphate, 7-hydrate (MgSO4.7H2O) |
81 |
Micro nutrient dissolve each reagent separately & mix in 250 ml of distilled water, then add 50 mL of conc. H2SO4 and makeup volume to 7 lit.
|
Element(micro nutrient) |
Reagent(ARGRADE) |
Preparation (g/250ml solution) |
|
Mn |
ManganousChloride,4hydrate(MnCl3.4H2O) |
0.375 |
|
Mo |
Ammoniummolybdate,4-hydrate[(NH4)6 Mo7O24. 4H2O] |
0.0185 |
|
Zn |
Zincsulphate,7-hydrate(ZnSO4. 7H2O) |
0.00875 |
|
B |
Boric acid (H3BO3) |
0.2335 |
|
Cu |
Cupricsulphate,5-hydrate(CuSO4.5H2O) |
0.0075 |
|
Fe |
Ferricchloride6-hydrate(FeCl3.6H20) |
1.925 |
|
|
Citricacid, monohydrate (C6H8O7.H2O) |
2.975 |
|
Element |
Reagent |
ml of stock required solution/1Lof working nutrient solution |
Concentration of element in ppm |
|
Macro nutrient |
|
|
|
|
N |
NH4NO3 |
1.25 |
40 |
|
P |
NaH2PO4.H2O |
1.25 |
10 |
|
K |
K2SO4 |
1.25 |
40 |
|
Ca |
CaCl2.2H2O |
1.25 |
40 |
|
Mg |
MgSO4.7H2O |
1.25 |
40 |
|
Micro nutrient |
|
|
|
|
Mn |
MnCl34H2O |
1.25 |
0.50 |
|
Mo |
(NH4)6 Mo7O24.4H2O |
1.25 |
0.05 |
|
Zn |
ZnSO4.7H2O |
1.25 |
0.01 |
|
B |
H3BO3 |
1.25 |
0.20 |
|
Cu |
CuSO4.5H2O |
1.25 |
0.01 |
|
Fe |
FeCl3.6H2O |
1.25 |
2.00 |
Observations Recorded
The morphological, physiological and biochemical parameters were recorded at different date of stress imposition of rice seeds. Three plants in each replication were selected at random as sample seedlings and tagged to record observations in salinity stress. The methods followed in recording each of these parameters are described below.
Morphological and Growth Characters
Germination Percentage
Seed germination tests were carried out and performed with three replications of 5 seeds each holes. The mean values were expressed in percentage.
Root Length
The root length was measured and expressed as cm.
Shoot Length
The shoot length was measured and expressed as cm.
Vigour Index
The Vigour Index was computed as per the procedure suggested by Abdul-Baki and Anderson, (1973) by using the following formula. (Shoot length + Root length) x Germination percentage
Stress Tolerance Index (STI)
The Stress Tolerance Index was computed as per the procedure suggested by Dhopte and Livera, (1989) by using the following formula.
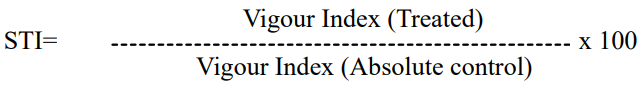
Physiological and Biochemical Parameters
Soluble Protein Content
Soluble protein content of the leaf was estimated by following the procedure described by Lowry et al. and expressed as mg g-1 of fresh weight [14].
Determination of Soluble Protein Content
Proteins were estimated using method.0.50 g of seed sample is macerated with 10ml of phosphate buffer (pH 7.0) [15]. The extract was centrifuged at 3000 rpm for 10 minutes and the supernatant was collected. One ml of the supernatant is transferred to a tube and 5ml of alkaline copper tartarate (ACT) reagent is added. The solution is kept as such for 30 minutes for color development. Then, 0.5ml of phenol reagent is added and the OD value of the sample for the intensity of blue color is read at 660nm in spectrophotometer. The protein content is expressed as mg g-1 of sample. Protein concentration was measured using bovine serum albumin as standard.
Alpha Amylase Content
Alpha amylase content of the germinated seedlings was estimated by following the procedure described by Miller, (1959) and expressed as enzyme unit g-1 fresh weight.
Determination of Alpha Amylase Content
One g of sprouting cereal (here rice) is milled in 5ml of prechilled 0.05M citrate buffer, pH 6.0; the resulting homogenate is centrifuged at 10,000rpm for 10 minutes. Enzyme activity is assayed in the supernatant as yield of crude enzyme. 0.1ml of this supernatant is pipette into a separate test tube and 0.9ml of 2%soluble starch is added and is incubated in shaking water bath at 50OC for 30 minutes. The reaction is stopped by adding DNSA reagent and boiled for at least 3 minutes for color development. Absorbance is read at 550nm against blank. Standard glucose curve is prepared from a series of glucose concentrations.
Nitrate Reductase Activity
Nitrate reductase activity in young leaves was estimated as per the method described by Nicholas et al. and the enzyme activity was expressed as μg NO2 g-1 h-1.
Determination of Nitrate Reductase Activity
A seedling of one g weight is taken in a test tube and 5ml of as-say medium is added.Thisiskeptinsidethevacuumdesiccator-for5minutes,undisturbed.2mlof the supernatant is pippeted out into a test tube and 1ml of 1M zinc acetate is added followed by 1ml of 70% ethanol. This is filtered using watman filter paper. To the filtrate 1ml of 1% sulphanilamide and 1ml of 0.02% NEDD is added. The OD Value is read at 540nm after the development of pink color using the spectrophotometer
Result and Discussion
Effect of Salinity on Morphological Characters
Germination Percentage (%)
The germination percentage was reduced by the salinity compared to control. Significant difference was noticed in all the treatments with respect to seed germination. Control recorded the highest germination percentage (92 to 100%) and salinity (40 to 80%) record¬ed least germination percentage (Table 1).
|
Varieties & Landraces |
Seed germination(%) |
Shoot length (cm) |
||
|
Control |
Salinity |
Control |
Salinity |
|
|
Kuliyadichan |
100.0 |
80.0 |
16.2 |
15.9 |
|
Kallurundaikar |
100.0 |
80.0 |
17.8 |
16.0 |
|
Norungan |
100.0 |
100.0 |
19.3 |
17.7 |
|
ASD16 |
98.0 |
60.0 |
12.8 |
13.3 |
|
MDU6 |
100.0 |
40.0 |
15.3 |
9.5 |
|
ADT45 |
92.0 |
80.0 |
13.5 |
7.9 |
|
Mean |
98.3 |
73.3 |
15.8 |
11.8 |
|
SEd |
1.91 |
1.71 |
0.20 |
0.12 |
|
CD(0.05) |
3.95 |
3.53 |
0.43 |
0.24 |
Table 1. Screening of rice genotypes on seed germination and shoot length for salinity stress tolerance
Among the rice genotypes, ADT45 recorded the minimum germination percentage (40%) followed by kuliyadichan (96%) under controlled conditions. The lowest rate of reduction in germination percentage in the genotype Kuliyadichan and Norungan under salinity stress imposed. Under salinity (NaCl 200 mM) conditions, the lowest rate of reduction in germination percentage recorded in the genotyes 3 land races Kuliyadichan, Kallurundaikar and Norungan. Highest reduction was observed in ASD16, ADT45 and MDU6 among the genotypes studied. Seed germination is very important stage for the victorious establishment of vigorous seedlings. This germination stage is very sensitive to salinity as compared to other stages. Salinity accumulates the toxic ion in plant cells causing a membrane damage and mineral imbalance. The most evident effect of salinity to the seed germination of rice was reducing the osmotic potential of soil which makes decline in water imbibition by seed to the creation of ionic toxicity which alters enzymes action involved in nucleic acid metabolism [16]. Other impacts of salt stress on seed germination include change in metabolism of protein [17]. In this present study, a significant increase was observed in germination percentage under controlled conditions. However, a considerable reduction could also be noticed in germination percentage due to the influence of sodium chloride treatments. The genotypes land races Kuliyadichan, Kallurundaikar and Norungan maintained its superiority with about 10 to 12 percent reduction, whereas, all the other genotypes showed about 25 to 30 percent reduction due to salinity.
Shoot and Root Length (Cm)
The results of shoot and root length showed a significant reduction under salinity stress compared to controlled condition. Among the rice genotypes, Norungan recorded the maximum shoot length (17.7cm) followed by Kallurundaikar (11.8cm) and Kuliyadichan (15.9cm) under controlled conditions. The lowest rate of reduction in shoot length due to the influence of sodium chloride in the genotypes of Norungan, Kuliyadichan and Kallurundaikar had lowest reduction in shoot length under salinity stress imposed and the highest reduction was observed in ASD16, MDU6 and ADT45 among the rice genotypes and hybrids studied. The root length was reduced by the influence of sodium chloride compared to control. Significant differences were noticed in all the treatments and genotypes with respect to root length. The control was recorded the maximum mean root length (17.9m) and sodium chloride 200mM recorded the minimum mean root length (7.4cm, 8.9cm and 4.0cm) (Table 2).
|
Varieties & Land races |
Root length (cm) |
Seedling vigour |
||
|
Control |
Salinity |
Control |
Salinity |
|
|
Kuliyadichan |
9.9 |
8.15 |
2610 |
1924 |
|
Kallurundaikar |
9.7 |
6.2 |
2750 |
1776 |
|
Norungan |
10.3 |
7.9 |
2960 |
2560 |
|
ASD16 |
9.3 |
5.8 |
2165.8 |
1146 |
|
MDU6 |
11.2 |
7.9 |
2650 |
1392 |
|
ADT45 |
9.6 |
6.3 |
2125.2 |
568 |
|
Mean |
10.0 |
7.04 |
2543.5 |
1561 |
|
SEd |
0.23 |
0.07 |
42.45 |
15.88 |
|
CD |
0.47 |
0.15 |
87.43 |
32.72 |
Table 2. Screening of rice genotypes on root length and seedling vigour for salinity stress tolerance
Among the rice genotypes, Kuliyadichan, Kallurundaikar, Norungan, ASD16, MDU6 and ADT45 recorded the maximum root length (15 to 17 cm) under salinity conditions and genotype kuliyadichan, kallurundaikar and MDU6 had maximum root length under salinity stress imposed. The rate of reduction in root length was the lowest in the genotypes Norungan, ASD16 andADT45 under the highest reduction was observed in ASD16 and ADT 45among the genotypes and hybrids studied.
The main functions of root are absorption of water and inorganic nutrients and anchoring of the plant body to the ground. Many researchers reported that with an increase in salinity there was a decrease in the development of the xylem and phloem. Many studies have shown that the fresh and dry weights of the shoot system are affected, either negatively or positively, by changes in salinity concentration, type of salt present, or type of plant species [18]. Root length was more suppressed than shoot by salinity at each specific salt concentration level. According to Rahman et al. who stated that, the gradual decrease in root length with the increase in salinity as observed might be due to more inhibitory effect of NaCl salt to root growth compared to that of shoot growth. In this present study, a lesser reduction was observed in the genotype of kuliyadichan and Norungan under salinity stress (NaCl) imposed showed about 11 to 15 percent and the highest reduction was observed in ASD16 and ADT45 with about 22 to 31 percent reduction due to the impact of salinity.
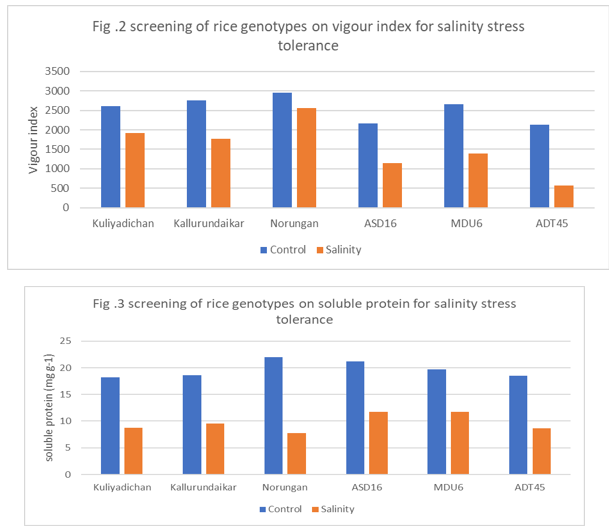
Vigour Index
Significant differences were noticed in all treatments and genotypes with respect to vigour index. The highest mean vigour index was recorded in control (2960) and lowest mean vigour index was recorded by salinity (568). Among the rice genotype, Norungan, Kuliyadichan and Kallurundaikar recorded the maximum vigour index (2560,1924 and 1776 ) under salinity stress (NaCl 200mM) imposed. The rate of reduction in vigour index was the lowest in the genotypes ASD16, MDU6, ADT45 and the highest reduction was observed in ADT45 among the genotypes and hybrids studied.
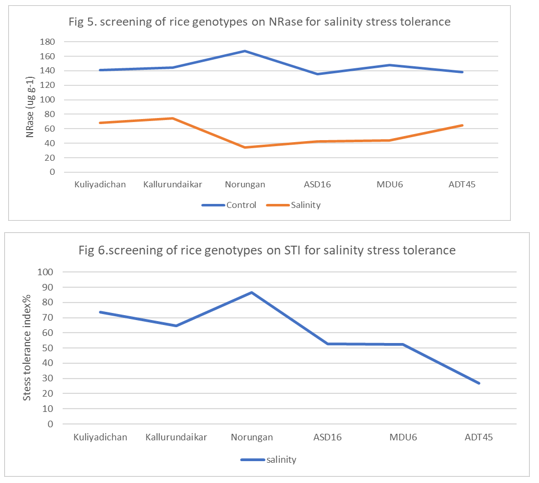
Stress Tolerance Index
The highest mean Stress Tolerance Index was recorded in Norungan (86.4)and lowest mean Stress Tolerance Index was recorded by salinityADT45(26.7) (Table 3). Among the rice genotypes, Norungan recorded the maximum Stress Tolerance Index (86.4%) followed by kuliyadichan (73.7) and kallurundaikar (64.5) had maximum Stress Tolerance Index under salinity stress (NaCl 200mM) imposed among the genotypes and hybrids studied.
Effect of Salinity Stress on Physiological and Biochemical Parameters
Soluble Protein Content (mg/g)
The data on soluble protein content was reduced by the influence of sodium chloride compared to control. The control was recorded the maximum mean soluble protein content (22.0mg g-1) and the sodium chloride 200 mM recorded the minimum mean soluble protein content (8.8mg g-1, 8.7mg g-1 and 7.8mg g-1).
|
Varieties & Land races |
Stress tolerance index (%) |
Soluble protein (mg g-1) |
|
|
Salinity |
Control |
Salinity |
|
|
Kuliyadichan |
73.7 |
18.2 |
8.8 |
|
Kallurundaikar |
64.5 |
18.6 |
9.6 |
|
Norungan |
86.4 |
22.0 |
7.8 |
|
ASD16 |
52.9 |
21.2 |
11.7 |
|
MDU6 |
52.5 |
19.7 |
11.7 |
|
ADT45 |
26.7 |
18.5 |
8.7 |
|
Mean |
59.45 |
19.7 |
9.71 |
|
SEd |
0.73 |
0.38 |
0.14 |
|
CD |
0.51 |
0.79 |
0.29 |
Table 3. Screening of rice genotypes on stress tolerance index and soluble protein content for salinity stress tolerance
Among the rice genotypes, Lalat recorded the genotypes Norungan, MDU6, ASD16 and kallurundaikar had maximum soluble protein content (11.7 mg g-1) under salinity stress imposed. The rate of reduction in soluble protein content was the highest reduction was observed in ADT45 and Norungan among the genotypes and hybrids studied. The soluble protein content of the rice seedlings, being a measure of RuBP carboxylase activity was considered as an index for photosynthetic efficiency. There were reports that RuBP-case enzyme forms nearly 80% of the soluble proteins in leaves of many plants [19]. Diethelm and Shibles opined that the RUBISCO content per unit leaf area was positively correlated with that of soluble protein content of the leaf [20]. In the present study, salinity caused a significant reduction in soluble protein content of seedlings of all the rice genotypes. Among the genotypes studied, MDU6, ASD16 and kallurundaikar maintained its superiority with about 8 to 12 percent reduction in soluble protein content due to the influence of sodium chloride. These findings were in accordance with the results of who observed a significant reduction in soluble protein content of rice plants grown under salinity stress conditions [21]. Besides these results, Martignone et al. observed that in soybean soluble protein content was the first nitrogenous compound affected under abiotic stress conditions, which at severity got denatured and lost the activity [22].
Nitrate Reductase Enzyme Activity(µgg-1d-1)
The result on nitrate reductase enzyme activity was reduced by the influence of sodium chloride compared to control. The control was recorded the maximum mean nitrate reductase enzyme activity (167.1µgg-1d-1) and the sodium chloride 200mM recorded the minimum mean nitrate reductase enzyme activity (141.1µg g-1 d-1, 138.1µg g-1 d-1and 135.7µg g-1 d-1) (Table 4).
Among the rice genotypes, Lalat, kallurundaikar and kuliyadichan had maximum nitrate reductase enzyme activity (74.6 to 68.1µg g-1 d-1) under salinity stress imposed. The highest reduction was observed in ASD16and Norungan among the genotypes and hybrids studied. Nitrate reductase is the important key enzyme of the nitrogen assimilation pathway in plant system. Its catalytic action in tissue is subjected to complex regulation in response to different abiotic factors. Salinity is one of the most important abiotic factors that restrictions growth and development of plants. It is well known that initial steps of nitrogen metabolismin plants (NO3−ions uptake, reduction of nitrates and assimilation of ammonium) aresensitive to salt stress [23]. The plants are treated with NaCl reduces the level of nitrate ions in the cytoplasms incite so inhibits nitrate uptake by plants [24]. In the present study, salinity caused a significant reduction in nitrate reductase enzyme activity of seedlings of all the rice genotypes. Among the genotypes studied, kallurundaikar (74.6), kuliyadichan (68.1), ADT45 (64.8) maintained its superiority with about 10 to 14 percent reduction in nitrate reductase enzyme activity due to the influence ofsodium chloride. These findings were in accordance with the results of Rao and Gnaham, who stated that, the nitrate uptake and nitrate reductase activity (NRA) decrease in plants under salinity stress [25].
Alpha Amylase Enzyme Activity (enzyme unitg-1 fresh weight)
The data on alpha amylase enzyme activity was reduced by the influence of sodium chloride compared to control. The control was recorded the maximum mean alpha amylase enzyme activity (15.1) and the sodium chloride 200 mM recorded the minimum mean alpha amylase enzyme activity (12.7, 12.5 and 12.3).
|
Varieties & Land races |
Nitrate reductase activity (µg g-1 d-1) |
Alpha amylase (enzyme unit g-1 fresh weight) |
||
|
Control |
Salinity |
Control |
Salinity |
|
|
Kuliyadichan |
141.1 |
68.1 |
12.7 |
6.2 |
|
Kallurundaikar |
144.7 |
74.6 |
13.0 |
6.7 |
|
Norungan |
167.1 |
34.2 |
15.1 |
3.1 |
|
ASD16 |
135.7 |
42.4 |
12.3 |
3.8 |
|
MDU6 |
148.2 |
43.8 |
13.4 |
3.9 |
|
ADT45 |
138.1 |
64.8 |
12.5 |
5.8 |
|
Mean |
145.8 |
54.65 |
13.16 |
4.9 |
|
SEd |
2.97 |
1.11 |
0.27 |
0.10 |
|
CD |
6.12 |
2.12 |
0.55 |
0.21 |
Table 4. Screening of rice genotypes on NRase and alpha amylase enzyme activity for water and salinity stress tolerance.
Among the rice genotypes, the genotypes kallurundaikar, kuliyadichan, ADT45 had maximum alpha amylase enzyme activity (6.7, 6.2 and 5.8) under salinity stress imposed. Higher accumulation of alpha-amylase during grain development, especially the filling stage, could result in rice grains with reduced quality, chalky, and with few stored starch grains. In rice, seed germination is dependent on the degradation of storage reserves in mature seeds, and the sugars from starch hydrolysis are the major source of energy for seedling emergence [26]. α-Amylase is the important enzyme implicated in starch mobilization; thus, α-amylase enzyme activity is an important factor in seed germination [27]. In this study, salinity caused a significant reduction in alpha-amylase enzyme activity of seedlings of all the rice genotypes. Among the genotypes studied, kallurundaikar, kuliyadichan and ADT45 maintained its superiority with about 12 to 17 percent reduction in alpha-amylase enzyme activity due to the influence of sodium chloride. These findings were in accordance with the results of who found that, positive relationships between bioactive GA content and α-amylase activity and between α-amylase activity and the rice seed germination rate and the seeds are growing under NaCl induced bioactive GA deficiency inhibits seed germination by decreasing α- amylase activity [28-92].
Acknowledgement
We thank Unit of Crop Physiology, Department of Seed Science and Technology, AC & RI, Madurai for product and laboratory support. We would like to thank Department of Plant Breeding and Genetics, AC & RI, Madurai for providing the rice seed materials.
References
- Pareek, A., Sopory, S. K., & Bohnert, H. J. (2009). Abiotic stress adaptation in plants. Dordrecht, The Netherlands: Springer.
- Mantri, N., Patade, V., Penna, S., Ford, R., & Pang, E. (2012). Abiotic stress responses in plants: present and future. Abiotic stress responses in plants: metabolism, productivity and sustainability, 1-19.
- Hasegawa, P. M., Bressan, R. A., Zhu, J. K., & Bohnert, H. J. (2000). Plant cellular and molecular responses to high salinity. Annual review of plant biology, 51(1), 463-499.
- Wang, Y., & Nii, N. (2000). Changes in chlorophyll, ribulose bisphosphate carboxylase-oxygenase, glycine betaine content, photosynthesis and transpiration in Amaranthus tricolor leaves during salt stress. The Journal of Horticultural Science and Biotechnology, 75(6), 623-627.
- Parida, A. K., & Das, A. B. (2005). Salt tolerance and salinity effects on plants: a review. Ecotoxicology and environmental safety, 60(3), 324-349.
- Pons, R., Cornejo, M. J., & Sanz, A. (2011). Differential salinity-induced variations in the activity of H+-pumps and Na+/H+ antiporters that are involved in cytoplasm ion homeostasis as a function of genotype and tolerance level in rice cell lines. Plant physiology and biochemistry, 49(12), 1399-1409.
- Munns, R., & Tester, M. (2008). Mechanisms of salinity tolerance. Annu. Rev. Plant Biol., 59(1), 651-681.
- Ali, S., Gautam, R. K., Mahajan, R., Krishnamurthy, S. L., Sharma, S. K., Singh, R. K., & Ismail, A. M. (2013). Stress indices and selectable traits in SALTOL QTL introgressed rice genotypes for reproductive stage tolerance to sodicity and salinity stresses. Field crops research, 154, 65-73.
- Munns, R., James, R. A., & Läuchli, A. (2006). Approaches to increasing the salt tolerance of wheat and other cereals. Journal of experimental botany, 57(5), 1025-1043.
- Blumwald, E. (2000). Sodium transport and salt tolerance in plants. Current opinion in cell biology, 12(4), 431-434.
- Joseph, B., Jini, D., & Sujatha, S. (2010). Biological and physiological perspectives of specificity in abiotic salt stress response from various rice plants. Asian J. Agric. Sci, 2(3), 99-105.
- Akbar, M. 1986. Breeding for salinity tolerance in rice. In: IRRI (Ed.), Salt-affected Soils of Pakistan, India and Thailand. International Rice Research Institute, Manila, Philippines,pp. 39–63.
- Zhu, G. Y., Kinet, J. M., & Lutts, S. (2001). Characterization of rice (Oryza sativa L.) F3 populations selected for salt resistance. I. Physiological behaviour during vegetative growth. Euphytica, 121, 251-263.
- Lowry, 0 H; Rosebrough, N J; Farr, A L and Randall, R J (1951). Proteins measurement with folin-phenol reagent., J. Bio. Che. 193: 269 - 275. Origin, dispersal, cultivation and variation of rice.
- Lowry, O.H., N.J. Rosebrough, A.L. Farr, R.J. Randall, J.Biol. Chem., 1951. 193, 265 – 275.
- Khan, M. A., & Weber, D. J. (Eds.). (2006). Ecophysiology of high salinity tolerant plants (Vol. 40). Springer Science & Business Media.
- Rasheed, R. (2009). Salinity and extreme temperature effects on sprouting buds of sugarcane (Saccharum oficinarum L.): some histological and biochemical studies (Doctoral dissertation, UNIVERSITY OF AGRICULTURE, FAISALABAD PAKISTAN).
- Taffouo, V. D., Wamba, O. F., Youmbi, E., Nono, G. V., & Akoa,(2010). Growth, yield, water status and ionic distribution response of three bambara groundnut (Vigna subterranea (L.) Verdc.) landraces grown under saline conditions.
- Joseph, M. C., Randall, D. D., & Nelson, C. J. (1981). Photosynthesis in polyploid tall fescue: II. Photosynthesis and ribulose-1, 5-bisphosphate carboxylase of polyploid tall fescue. Plant physiology, 68(4), 894-898.
- Diethelm, R., & Shibles, R. (1989). Relationship of enhanced sink demand with photosynthesis and amount and activity of ribulose 1, 5-bisphosphate carboxylase in soybean leaves. Journal of plant physiology, 134(1), 70-74.
- Chaves, M. M., Flexas, J., & Pinheiro, C. (2009). Photosynthesis under drought and salt stress: regulation mechanisms from whole plant to cell. Annals of botany, 103(4), 551-560.
- Martignone, R. A., Guiamet, J. J., & Nakayama, F. (1987). Nitrogen partitioning and leaf senescence in soybean as related to nitrogen supply. Field Crops Research, 17(1), 17-24.
- Flores, P., Botella, M. Á., Cerdá, A., & Martínez, V. (2004). Influence of nitrate level on nitrate assimilation in tomato (Lycopersicon esculentum) plants under saline stress. Canadian journal of botany, 82(2), 207-213.
- Klobus, G., Ward, M. R., & Huffaker, R. C. (1988). Characteristics of injury and recovery of net NO3− transport of barley seedlings from treatments of NaCl. Plant Physiology, 87(4), 878-882.
- Rao, R. K., & Gnanam, A. (1990). Inhibition of nitrate and nitrite reductase activities by salinity stress in Sorghum vulgare. Phytochemistry, 29(4), 1047-1049.
- Beck, E., & Ziegler, P. (1989). Biosynthesis and degradation of starch in higher plants.
- Karrer, E. E., Litts, J. C., & Rodriguez, R. L. (1991). Differential expression of α-amylase genes in germinating rice and barley seeds. Plant molecular biology, 16, 797-805.
- Appleford, N. E., Wilkinson, M. D., Ma, Q., Evans, D. J.,Stone, M. C., Pearce, S. P., ... & Lenton, J. R. (2007). Decreased shoot stature and grain α-amylase activity following ectopic expression of a gibberellin 2-oxidase gene in transgenic wheat. Journal of Experimental Botany, 58(12), 3213-3226.
- Prisco, J. T., & Vieira, G. H. F. (1976). Effects of NaCl salinity on nitrogenous compounds and proteases during germination of Vigna sinensis seeds. Physiologia Plantarum, 36(4), 317-320.
- Ratnakar, A., & Rai, A. (2013). Effect of sodium chloride salinity on seed germination and early seedling growth of Trigonella foenum-graecum L. Var. Peb. Octa Journal of Environmental Research, 1(4).
- Abdul-Baki, A. A., & Anderson, J. D. (1973). Vigor determination in soybean seed by multiple criteria 1. Crop science, 13(6), 630-633.
- Agastian, P., Kingsley, S. J., & Vivekanandan, M. (2000). Effect of salinity on photosynthesis and biochemical characteristics in mulberry genotypes. Photosynthetica, 38, 287-290.
- Aiyswaraya, S., Saraswathi, R., Ramchander, S., Vinoth, R., Uma, D., Sudhakar, D., & Robin, S. (2017). An insight into total soluble protein across rice (Oryza sativa L.) germplasm accessions. International Journal of Current Microbiology and Applied Sciences, 6(12), 2254-2269.
- Ali, S., Gautam, R. K., Mahajan, R., Krishnamurthy, S. L., Sharma, S. K., Singh, R. K., & Ismail, A. M. (2013). Stress indices and selectable traits in SALTOL QTL introgressed rice genotypes for reproductive stage tolerance to sodicity and salinity stresses. Field crops research, 154, 65-73.
- Andersen, M. N., Asch, F., Wu, Y., Jensen, C. R., Næsted, H., Mogensen, V. O., & Koch, K. E. (2002). Soluble invertase expression is an early target of drought stress during the critical, abortion-sensitive phase of young ovary development in maize. Plant physiology, 130(2), 591-604.
- Ashraf, M., & Fatima, H. (1995). Responses of salt-tolerant and salt-sensitive lines of safflower (Carthamus tinctorius L.) to salt stress.
- Ashraf, M. P. J. C., & Harris, P. J. (2004). Potential biochemical indicators of salinity tolerance in plants. Plant science, 166(1), 3-16.
- Badieh, M. M. S., Farshadfar, E., Haghparast, R., Rajabi, R., & Zarei, L. (2012). Evaluation of gene actions of some traits contributing in drought tolerance in bread wheat utilizing diallel analysis.
- Bänziger, M., Edmeades, G. O., Beck, D., & Bellon, M. (2000). Breeding for drought and nitrogen stress tolerance in maize: from theory to practice.,(CIMMYT, Mexico DF, Mexico). ISBN 970-648-46-368.
- Bartels, D., & Sunkar, R. (2005). Drought and salt tolerance in plants. Critical reviews in plant sciences, 24(1), 23-58.
- Bhargava, S., & Paranjpe, S. (2004). Genotypic variation in the photosynthetic competence of Sorghum bicolor seedlings subjected to polyethylene glycol-mediated drought stress. Journal of plant physiology, 161(1), 125-129.
- Bhatt, D., Negi, M., Sharma, P., Saxena, S. C., Dobriyal,K., & Arora, S. (2011). Responses to drought induced oxidative stress in five finger millet varieties differing in their geographical distribution. Physiology and Molecular Biology of Plants, 17, 347-353.
- Erica, B., Larsson, C. M., & Larsson, M. (1996). Responsesof nitrate assimilation and N translocation in tomato (Lycopersicon esculentum Mill) to reduced ambient air humidity. Journal of experimental botany, 47(7), 855-861.
- Chaum, S., Nhung, N.T.H., Kirdmanee, C. 2010. Effect of mannitol and salt induced isoosmotic stress on proline accumulation, photosynthetic abilities and growth characters of rice cultivars (Oryza sativa L. spp. indica). Pak. J. Bot., 42: 927–941.
- Chennafi, H., Aïdaoui, A., Bouzerzour, H., & Saci, A. (2006). Yield response of durum wheat (Triticum durum Desf.) cultivar waha to deficit irrigation under semi arid growth conditions.
- Clarke, J. M., DePauw, R. M., & Townley-Smith, T. F. (1992). Evaluation of methods for quantification of drought tolerance in wheat. Crop science, 32(3), 723-728.
- Cooper, M., Podlich, D. W., & Smith, O. S. (2005). Gene-to-phenotype models and complex trait genetics. Australian Journal of Agricultural Research, 56(9), 895-918.
- Dai, H. P., Zhang, P. P., Lu, C., Jia, G. L., Song, H., Ren, X.M., ... & Zhang, S. Q. (2011). Leaf senescence and reactive oxygen species metabolism of broomcorn millet ('Panicum miliaceum'L.) under drought condition. Australian Journal of Crop Science, 5(12), 1655-1660.
- Evans, T., Rosenthal, E. T., Youngblom, J., Distel, D., & Hunt,T. (1983). Cyclin: a protein specified by maternal mRNA in sea urchin eggs that is destroyed at each cleavage division. Cell, 33(2), 389-396.
- Flowers, T. J., & Yeo, A. R. (1981). Variability in the resistance of sodium chloride salinity within rice (Oryza sativa L.) varieties. New Phytologist, 88(2), 363-373.
- Garg, A. K., Kim, J. K., Owens, T. G., Ranwala, A. P., Choi, Y. D., Kochian, L. V., & Wu, R. J. (2002). Trehalose accumulation in rice plants confers high tolerance levels to different abiotic stresses. Proceedings of the National Academy of Sciences, 99(25), 15898-15903.
- Gharoobi, B., Ghorbani, M., & GHASEMI, N. M. (2012). Effects of different levels of osmotic potential on germination percentage and germination rate of barley, corn and canola.
- Gomes Filho, E., & Sodek, L. (1988). Effect of salinity on ribonuclease activity of Vigna unguiculata cotyledons during germination. Journal of plant physiology, 132(3), 307-311.
- Gregorio, G. B. (1997). Tagging salinity tolerance genes in rice using amplified fragment length polymorphism (AFLP).
- Handa, S., Handa, A. K., Hasegawa, P. M., & Bressan, R. A. (1986). Proline accumulation and the adaptation of cultured plant cells to water stress. Plant physiology, 80(4), 938-945.
- Hsiao, T. C. (1973). Plant responses to water deficit. Ann. Rev. Plant Physiol, 24, 519-570.
- Huber, S. C., McMichael, R. W., Bachmann, M., Huber, J. L., Shannon, J. C., Kang, K. K., & Paul, M. J. (1996). Regulation of leaf sucrose-phosphate synthase and nitrate reductase by reversible protein phosphorylation.
- Hussain, M., Malik, M. A., Farooq, M., Ashraf, M. Y., & Cheema, M. A. (2008). Improving drought tolerance by exogenous application of glycinebetaine and salicylic acid in sunflower. Journal of Agronomy and Crop Science, 194(3), 193-199.
- Jha, B. N., & Singh, R. A. (1997). Physiological responses of rice varieties to different levels of moisture stress.
- Jin, J., Zhang, H., Kong, L., Gao, G., & Luo, J. (2014).PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic acids research, 42(D1), D1182-D1187.
- Joseph, B., Jini, D., & Sujatha, S. (2010). Biological and physiological perspectives of specificity in abiotic salt stress response from various rice plants. Asian J. Agric. Sci, 2(3), 99-105.
- Hakim, M. A., Juraimi, A. S., Begum, M., Hanafi, M. M., Ismail, M. R., & Selamat, A. (2010). Effect of salt stress on germination and early seedling growth of rice (Oryza sativa L.). African journal of biotechnology, 9(13), 1911-1918.
- Kaiser, W. M., & Brendle-Behnisch, E. (1991). Rapid modulation of spinach leaf nitrate reductase activity by photosynthesis: I. Modulation in vivo by CO2 availability. Plant Physiology, 96(2), 363-367.
- Kaiser, W. M., & Forster, J. (1989). Low CO2 prevents nitratereduction in leaves. Plant Physiology, 91(3), 970-974.
- Kaiser, W. M., & Huber, S. C. (1994). Posttranslational regulation of nitrate reductase in higher plants. Plant Physiology, 106(3), 817.
- MK, K. (2002). Evaluation Of Sunflower (Helianthus Annuus L.) Genotypes At Gametophytic And Sporophytic Stages For Water Stress Tolerance And Development Of Tolerant Hybrids Through Selective Fertilization Technique (Doctoral dissertation, TAMIL NADU AGRICULTURAL UNIVERSITY COIMBATORE).
- Khaayatnezhad M., Zaeifizadeh, M., Gholamin, R.(2010). Investigation and selection index for drought stress Article in Australian Journal of Basic and Applied Sciences 4(10):4815-4822.
- Khush, G.S. (1997). Authors Khush GS1.Author information Plant Breeding, Genetics and Biochemistry Division, International Rice Research Institute, Manila, Philippines. Plant Mol Biol. Sep 35(1-2) 25-34.
- Kumar, U., Joshi, A. K., Kumari, M., Paliwal, R., Kumar, S., & Röder, M. S. (2010). Identification of QTLs for stay green trait in wheat (Triticum aestivum L.) in the ‘Chirya 3’בSonalika’population. Euphytica, 174, 437-445.
- Larsson, M., Larsson, C. M., Whitford, P. N., & Clarkson,D. T. (1989). Influence of osmotic stress on nitrate reductase activity in wheat (Triticum aestivum L.) and the role of abscisic acid. Journal of Experimental Botany, 40(11), 1265-1271.
- Lefevre, I., Gratia, E., & Lutts, S. (2001). Discrimination between the ionic and osmotic components of salt stress in relation to free polyamine level in rice (Oryza sativa). Plant science, 161(5), 943-952.
- Lutts, S., Kinet, J. M., J. BouharmontLaboratoire de Cytogenetique, Universite Catholique de Louvain, 5 (Bte 13) Place Croix du Sud, B-1348 Louvain-la-Neuve, Belgium.
- Manabendra, D., Baruah, K. K., & Deka, M. (1998). Moisture stress induced changes in seed germination and seedling growth of upland ‘Ahu’rice (Oryza sativa L.). Indian J. Ecol, 25, 133-137.
- Mori IK, Kinoshita T (1987). Salt tolerance of rice callus clones. Rice Genet. Newsletter 4: 112–113. Article information J Transl Med. 2009; 7: 75.
- Munns, R. (2002). Comparative physiology of salt and water stress. Plant, cell & environment, 25(2), 239-250.
- Nasim, M. U. H. A. M. M. A. D., Qureshi, R., Aziz, T. A.R. I. Q., Saqib, M., Nawaz, S. H. A. F. Q. A. T., Sahi, S. T.,& Pervaiz, S. (2008). Growth and ionic composition of salt stressed Eucalyptus camaldulensis and Eucalyptus teretcornis. Pakistan Journal of Botany, 40(2), 799-805.
- Nedjimi, B., & Daoud, Y. (2006). Effect of Na 2 SO 4 on the growth, water relations, proline, total soluble sugars and ion content of Atriplex halimus subsp. schweinfurthii through in vitro culture. In Anales de Biología (No. 28, pp. 35-43). Servicio de Publicaciones de la Universidad de Murcia.
- Nonami, H. (1998). Plant water relations and control of cell elongation at low water potentials. Journal of Plant Research, 111, 373-382.
- Pareek, A., Singla, S. L., & Grover, A. (1997). Salt responsive proteins/genes in crop plants.
- Passioura, J. B. (1983). Roots and drought resistance. In Developments in agricultural and managed forest ecology (Vol. 12, pp. 265-280).
- Prisco, J. T., Eneas Filho, J., & Gomes Filho, E. (1981). Effect of NaCl salinity on cotyledon starch mobilization during germination of Vigna unguiculata (L.) Walp seeds. Revista brasileira de Botânica, 4, 63-71.
- Rajakumar, R. (2011). A study on effect of salt stress in the seed germination and biochemical parameters of rice (Oryza sativa L.) under in vitro condition. Asian Journal of Plant Science & Research.
- Rauf, S., Sadaqat, H. A., & Khan, I. A. (2008). Effect of moisture regimes on combining ability variations of seedling traits in sunflower (Helianthus annuus L.). Canadian Journal of Plant Science, 88(2), 323-329.
- Sahi, C., Singh, A., Blumwald, E., & Grover, A. (2006). Beyond osmolytes and transporters: novel plant salt-stress tolerance-related genes from transcriptional profiling data. Physiologia Plantarum, 127(1), 1-9.
- Olivié, D. J. L., Peters, G. P., & Saint-Martin, D. (2012). Atmosphere response time scales estimated from AOGCM experiments. Journal of Climate, 25(22), 7956-7972.
- Sari-Gorla, M., Krajewski, P., Di Fonzo, N., Villa, M., & Frova, C. (1999). Genetic analysis of drought tolerance in maize by molecular markers. II. Plant height and flowering. Theoretical and Applied Genetics, 99, 289-295.
- Singh, N. K., Bracker, C. A., Hasegawa, P. M., Handa,K., Buckel, S., Hermodson, M. A., ... & Bressan, R. A. (1987). Characterization of osmotin: a thaumatin-like protein associated with osmotic adaptation in plant cells. Plant physiology, 85(2), 529-536.
- Sivaramakrishnan, S., Patell, V. Z., Flower, D. J., & Peacock,J. M. (1988). Proline accumulation and nitrate reductase activity in contrasting sorghum lines during mid-season drought stress. Physiologia Plantarum, 74(3), 418-426.
- Talouizite, A., & Champigny, M. L. (1988). Response of wheat seedlings to short-term drought stress with particular respect to nitrate utilization. Plant, Cell & Environment, 11(3), 149-155.
- Tavakkoli, E., Fatehi, F., Coventry, S., Rengasamy, P., & McDonald, G. K. (2011). Additive effects of Na+ and Cl–ions on barley growth under salinity stress. Journal of ExperimentalBotany, 62(6), 2189-2203.
- Hsiao, T. C., & Acevedo, E. (1975). Plant responses to water deficits, water-use efficiency, and drought resistance. Developments in Agricultural and Managed Forest Ecology, 1, 59-84.
- Wang, W. B., Kim, Y. H., Lee, H. S., Kim, K. Y., Deng, X.P., & Kwak, S. S. (2009). Analysis of antioxidant enzyme activity during germination of alfalfa under salt and drought stresses. Plant physiology and Biochemistry, 47(7), 570-577.



