Research Article - (2024) Volume 9, Issue 1
In-Silico Analysis of Heart and Neural Crest Derivatives (HAND) Subclass Basic Helix-Loop-Helix (bHLH) Transcription Factor in Cardiogenesis
Received Date: Oct 05, 2023 / Accepted Date: Dec 27, 2023 / Published Date: Jan 08, 2024
Copyright: ©Shouhartha Choudhury. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Choudhury, S. (2024). In-Silico Analysis of Heart and Neural Crest Derivatives (HAND) Subclass Basic Helix- Loop-Helix (bHLH) Transcription Factor in Cardiogenesis. Cardio Open, 9(1), 01-15.
Abstract
Background: The HANDs are muscle-specific bHLH TFs crucial for proper cardiac and extra-embryonic development. The eHAND and dHAND functioned in developing ventricular chambers (right and left ventricle), aortic arch arteries, cardiac neural crest, endocardium and epicardium. The down-regulated response of the eHAND and dHAND genes reflects permissiveness. A recent report suggested that cardiac hypertrophy intimate eHAND corresponds to cardiomyopathy and dHAND in the atrium. Those reports supported the cardiac muscles may re-initiate a fetal gene result and initiate physiological changes, which allow the heart to recompense.
Objective: In this study, the objective is an investigation of the HAND subclass bHLH transcription factors in mammals. I like to classify the bHLH TFs and discuss the genetic evidence of both eHAND and dHAND genes in cardiogenesis. So, perform bioinformatics and computational tools and techniques to the current knowledge of the HAND subclass bHLH transcription factor in the mammalian genome. This application may be valuable for future functional analysis of particular TFs in different organisms.
Results: The observation data demonstrated that the heart and neural crest derivative transcription factors are present in mammals. The two mammalian genomes' likelihood of Homo sapiens and Mus musculus perform for comparative analysis. Analysis data suggested the eHAND and dHAND genes and a total number of bHLH domains in Homo sapiens and Mus musculus. Also, the conserved domain, motifs, phylogeny, gene expression and chromosome location analysis demonstrated the heart and neural crest derivative factors associated with cardiogenesis.
Conclusion: Over the last decades, a wealth of new reports has been composed of unique genetic and phenotypic characteristics of cardiac morphogenesis. The mutational analysis of the eHAND and dHAND transcription factors enabled precise resolution of specialized function during the developing myocardium. Also, analysis data concluded the muscle-specific transcription factors eHAND and dHAND are associated with cardiac disease and development. In contrast, the tissue-specific bHLH and other TFs lead to the development of myogenesis and vasculogenesis.
Keywords
eHAND, dHAND, bHLH Transcription Factor, Cardiogenesis
List Of Abbreviations
TF’s: Transcription Factors
bHLH: Basic Helix-Loop-Helix
HAND1: Heart and Neural Crest Derivative 1
HAND2: Heart and Neural Crest Derivative 2
HANDs: Heart and Neural Crest Derivatives
cNCCs: Cardiac Neural Crest Cells
BLAST: Basic Local Alignment Search Tools
HMM: Hidden Markov Model
GO: Gene Ontology
MSA: Multiple Sequence Alignment
KEGG: Kyoto Encyclopedia of Genes and Genomes
UniProt: Universal Protein Resource
NCBI: National Center for Biotechnology Information
EMBL: European Molecular Biology Laboratory
DDBJ: DNA Data Bank of Japan
SMART: Simple Modular Architecture Research Tool
STRING: Search Tool for the Retrieval of Interacting Genes/ Proteins
NGM: Neighbor-Joining Methods
MEGA: Molecular Evolutionary Genetics Analysis
MEME: Multiple Expectation Maximizations for Motif Elicitation
Introduction
The HANDs (heart and neural crest derivatives) sub-class bHLH TFs of eHAND and dHAND genes initiate normal progression of cardiac and extra-embryonic development in mammals. Both eHAND and dHAND functions govern the growth of the cNCCs, left and right ventricle, aortic arch, endocardium and epicardium. HAND1 (eHAND) and HAND2 (dHAND) reveal sided expressions in ventricles (left and right), despite their genetic function subtly across species. The HAND factors express at the cardiac crescent stage also their expressions maintain cardiac development [1-6]. The downregulated response of eHAND and dHAND propose the developmental progress of the adult heart [7-9]. Recent data suggested the eHAND and dHAND expressions develop tetralogy of fallot and hypertrophic cardiomyopathy (HCM) [10]. Also, congenital heart defects suggested HAND1 and HAND2 functions in the cardiac tissues and their clinical anomalies. So, those defects occur between the distinct regions in the chamber-specific heart.
The molecular study proposed that cardiac outgrowth and inherent heart disease are normal and abnormal segments between HAND1 and HAND2. So, the HANDs mutants reveal hypoplasia in the right and left ventricles, aortic arch, and branchial arches. The dHAND mutant drives into dissecting molecules and controls the morphogenesis of the particular components in the heart. Inherited heart symptom is a deficiency of the segment between the ventricles (left and right) in the mature heart. So, the eHAND and dHAND regulate the normal growth of ventricles (left and right) and also malformed cardiogenesis. The molecular study suggested eHAND and dHAND expression in ventricles. Also, HAND2 expresses in the atrium, but the HAND1 response is not clear in the atrium. The expression of eHAND identified by Hon’ble Natarajan and D Srivastava et al. reported cardiomyopathy. The heart responds to chronic arterial blood pressure by reactivating genes linked with the fetal gene program result of changes in expression is hypertrophy of cardiomyocytes [11]. The eHAND reveals differentiation in trophoblast cell lineage and over-expresses in the cell. As hypertrophy increases in the cytoplasm with less cell division, it’s possible the reduction of eHAND and dHAND expression in ventricles. A recent study supported that the heart reinitiates a fetal gene sequence and instructs cells to increase their cytoplasmic mass, and then eHAND and dHAND expression may reduce hypertrophic cardiomyopathy. The molecular mechanisms that initiate the re-expression of the fetal gene sequence (innate cell-survival pathways) are critical to gain improved treatment of patients with heart disease. The downregulated expression of eHAND and dHAND may be early markers for the onset of cardiac disease [12,13].
Therefore, the muscle-specific bHLH TFs (eHAND and dHAND) express in a complementary fashion during the growth of the left and right ventricles. Both genes express in the aortic arch and neural crest-derived cardiac sections. The mutations of HAND1 and HAND2 impart superior organogenesis in mesoderm and ectoderm with neural crest. Also, HAND1 and HAND2 function in cardiac growth proposed a high degree of the evolutionarily conserved mechanism. The above study conducted a compressive genome-wide survey of the HANDs transcription factors (TFs) in two mammalian genomes using the In-Silico approach. So, both muscle-specific bHLH TFs studies are crucial for organism research and development.
Materials and Methods
Primary Sequence and Database
The primary sequence retrieves from different specialised databases (UniProt, KEGG, GenBank, EMBL, DDBJ and NCBI), then performs web-based application SMART for identification of the particular domain in the objective sequence. Pfam searches for retrieving protein family information. PROSITE performs for identification of domain, family, and functional residue and linked patterns and profiles. PROCHECK perform for the stereo- chemical property of the primary peptide sequences. Also, the draft genome sequence of particular organisms downloads from the genomic data in the different specialised databases (NCBI and Ensemble).
Standalone Tools and GO Annotation
HMMER executes through MSA (multiple sequence alignment) of the particular domain as a profile search. HMMER is a statistical algorithm making MSA of the particular domains as a profile search and system program using probabilistic models called the profile HMM. Standalone BLAST performs for homologs gene in different organisms. Also, the BLAST2GO performs for the sequence accuracy of the particular genes in the genome. BLAST2GO is a bioinformatics and computational tool for high-throughput gene annotation of sequences. The functional information retrieves via GO annotation, a standard library of functional attributes.
Domain, Motif, and Phylogeny
A multiple sequence alignment (MSA) method performs for the conserved domain in the organism. The MSA method calculates the best match of the homolog sequences and lines them up, and then the identities, similarities and differences can be seen. MSA of highest hits sequence analysis carried out with a web-based tool MultAlin for identification of conserved domain. The analysis of the molecular evolutionary relationship of the particular gene in different organisms. The MEGA7 for constructing the phylogenetic tree using Neighbor-Joining Methods. The MEME suite performs for retrieving sequence motifs. The MEME is a computational web-based tool for the analysis of sequence motifs.
Gene Expression, Chromosome Location, and Gene Network
The gene expression analysis utilised using the GENEVESTIGATOR tool is a high-performance search engine for gene expression in different biological contexts. GENEVESTIGATOR used to identify and validate targets. Chromosome location retrieves using a gene card is a database of the genes that provide genomic information of known also predicted human genes. This database is currently available for biomedical research such as genes, encoded proteins and associated diseases. The gene regulatory matrix or network is a molecular interaction in the cellular process to dominate the expression volume of mRNA or proteins. Some proteins act to activate genes are the TFs bind to the promoter area and initiate the response of different proteins known as regulatory cascades. STRING database retrieves for prediction of protein-protein interaction. STRING database includes various resources like experimental data and computational prediction of nucleic acids and proteins.
Results
Structural Analysis of the eHAND and dHAND
The primary sequence includes nucleotides and peptides. The eHAND compose of 648 nucleotides and 215 peptides among 53 peptides binding to the DNA, in dHNAD is composed of 654 nucleotides and 217 peptides with 52 peptide binding to the DNA, which is well known as a muscle-specific bHLH transcription factors (Table 1). The peptide structure demonstrated the bHLH domain characterized by two alpha-helix connected by a loop. The adaptability of the loop allows the dimerization of folding and packing against another helix. Both amphipathic alpha- helices are separate by a linker region of length (Figure 1).

Table 1: Primary Sequence (a) Nucleotides and (b) Peptides
Figure 1: Structure of the bHLH Domain
Genome-Wide Analysis of eHAND and dHAND
Genome-wide analysis by HMMER algorithm results obtains a total of 116 and 100 bHLH domains in Homo sapiens and Mus musculus, respectively (Table 2). Standalone BLAST2 results represent 45 and 46 homologs in Homo sapiens and Mus musculus, respectively (Table 2). The gene ontology (GO) annotation result demonstrated the sequence accuracy of eHAND and dHAND transcription factors (TFs) in Homo sapiens and Mus musculus (Table 3).
|
Organisms |
HMMER |
BLAST2 |
BLSAT2GO |
|
Homo sapiens |
116 |
45 |
1 |
|
Mus musculus |
100 |
46 |
2 |
|
Total |
216 |
91 |
3 |
Table 2: Summary of the bHLH Domain in Homo Sapiens and Mus Musculus
|
Gene |
Homo sapiens |
Mus musculus |
|
HAND1 |
1 |
2 |
|
HAND2 |
2 |
1 |
|
TCF15 |
1 |
1 |
|
TWIST1 |
3 |
1 |
|
NHLH1 |
1 |
1 |
|
NHLH2 |
3 |
3 |
|
TWIST2 |
3 |
2 |
|
TAL2 |
1 |
1 |
|
TCF21 |
2 |
2 |
|
MSC |
1 |
1 |
|
PTF1 |
1 |
1 |
|
BHLHA15 |
2 |
1 |
|
TAL1 |
2 |
3 |
|
TCF24 |
1 |
1 |
|
LYL1 |
1 |
1 |
|
ATOH7 |
1 |
1 |
|
TCF23 |
1 |
1 |
|
NEUROG1 |
1 |
1 |
|
NEUROG2 |
1 |
1 |
|
NEUROG3 |
1 |
2 |
|
ASCL1 |
1 |
1 |
|
ATOH1 |
1 |
1 |
|
ASCL5 |
2 |
1 |
|
ASCL4 |
1 |
1 |
|
FER3 |
1 |
1 |
|
NEUROD1 |
1 |
1 |
|
NEUROD2 |
1 |
1 |
|
ASCL3 |
1 |
1 |
|
NEUROD6 |
1 |
1 |
|
NEUROD4 |
1 |
1 |
|
BHLHE22 |
1 |
1 |
|
ASCL2 |
1 |
2 |
|
FIGLA |
1 |
1 |
|
BHLHE23 |
2 |
1 |
|
OLIG2 |
2 |
1 |
|
OLIG3 |
1 |
1 |
|
MESP1 |
1 |
1 |
|
ATOH8 |
1 |
1 |
|
MESP2 |
1 |
0 |
|
MYF6 |
1 |
1 |
|
ID3 |
2 |
1 |
|
MYF5 |
1 |
1 |
|
TCFAP4 |
1 |
1 |
|
OLIG1 |
1 |
1 |
|
MYOD1 |
1 |
1 |
|
MAX |
10 |
4 |
|
BHLHA9 |
1 |
1 |
|
MYC |
5 |
10 |
|
ID2 |
3 |
3 |
|
ARNTL1 |
7 |
4 |
|
ARNTL2 |
9 |
5 |
|
ID1 |
2 |
2 |
|
ID4 |
1 |
1 |
|
SREBP2 |
3 |
1 |
|
MSGN1 |
1 |
1 |
|
SREBF1 |
1 |
0 |
|
USF2 |
5 |
4 |
|
USF3 |
1 |
0 |
|
MYOG |
1 |
1 |
|
SIM1 |
2 |
1 |
|
Total |
112 |
91 |
Table 3: Summary of the GO annotation of Homo sapiens and Mus musculus
Analysis of the Domain, Motifs, and Phylogeny
The highest hits of eHAND and dHAND listed from both organisms for sequence alignment, a multiple sequence alignment (MSA) results demonstrated conserved bHLH domain. The high consensus (90%) indicates extended HAND subclass bHLH domain (Figure 2) and their specific motifs (Figure 3). The analysis of the phylogeny demonstrated the evolutionary link of the HAND1 and HAND2 between Homo sapiens and Mus musculus. Particular clades represent multi-functional bHLH domain-involved genes in both organisms (Figure 4).\
Figure 2: MAS (Conserve bHLH domain)
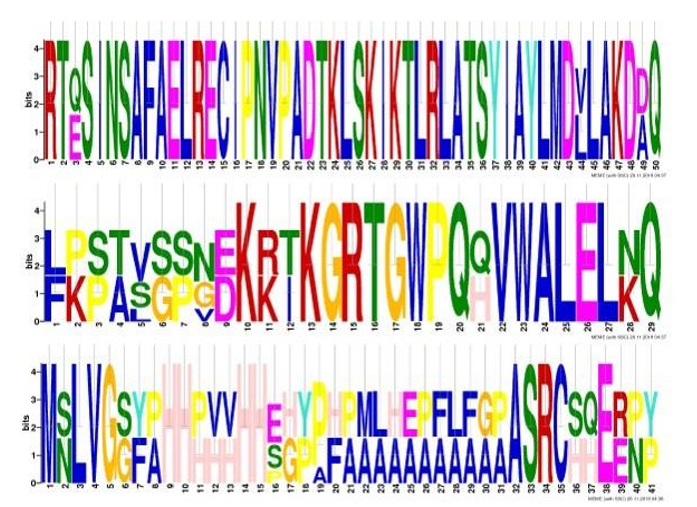
Figure 3: Specific Motifs (a, b, and c)
Figure 4: Phylogenetic Tree of the Different bHLH Domains
Gene Expression, Chromosome Location, and Gene Network Analysis of eHAND and dHAND
The eHAND expression is normal in tissue, the circulatory and cardiovascular system. Also, dHAND highly expresses in tissue, the circulatory and cardiovascular system, heart muscle (myocardium, cardiac muscle), heart ventricle, and heart left ventricle of the human (Figure 5). Chromosome location study confirmed that the eHAND located band 5q33.2 (Figure 6) and dHAND located band 4q34.1 (Figure 7) in human. The gene network study determines that the HAND1/HAND2 interact with other molecules such as Forkhead box protein P4 (FOXP4), Transcription factor GATA-4 (GATA4), 2',3'-Cyclic Nucleotide 3' Phosphodiesterase (CNP), Xylosyltransferase 1 (XYLT1), Adenylosuccinate Synthase 2 (ADSS), Home box protein Nkx-2.5 (NKX2-5), Epididymal Sperm Binding Protein 1 (ELSPBP1), Myocyte-specific enhancer factor 2C (MEF2C) and Nuclear factor of activated T-cells, cytoplasmic 1 (NFATC1).
Figure 5: Gene Expression of eHAND and dHAND
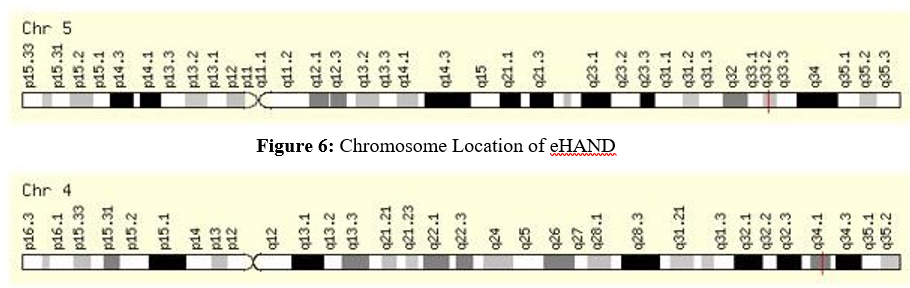

Figure 7: Chromosome Location of dHAND
Those molecular interactions govern the outcome of the HAND1/HAND2 gene in the cells (Figure 8). So, the principle of TFs data analysis suggested multiple hits of HANDs subclass bHLH TF and a total of 94 and 84 multiple bHLH domains in Homo sapiens and Mus musculus respectively (Table 4).
Figure 8: Gene Regulatory Network of eHAND and dHAND
|
Gene Id |
Protein |
|
ENSP00000231121.2 |
Heart and neural crest derivatives expressed 1 |
|
ENSP00000477638.1 |
HAND2 isoform 3 |
|
ENSP00000352565.4 |
dHand protein |
|
ENSP00000427084.1 |
HAND2 isoform 1 |
|
ENSP00000246080.3 |
transcription factor 15 |
|
ENSP00000346582.5 |
twist-related protein 1 |
|
ENSP00000302189.5 |
helix-loop-helix protein 1 |
|
ENSP00000322087.3 |
helix-loop-helix protein 2 |
|
ENSP00000358519.1 |
helix-loop-helix protein 2 |
|
ENSP00000405176.2 |
twist-related protein 2 |
|
ENSP00000482581.1 |
twist-related protein 2 |
|
ENSP00000242261.5 |
twist-related protein 1 |
|
ENSP00000334547.3 |
T-cell acute lymphocytic leukemia protein 2 |
|
ENSP00000237316.3 |
transcription factor 21 |
|
ENSP00000356857.4 |
transcription factor 21 |
|
ENSP00000321445.4 |
musculin |
|
ENSP00000365687.3 |
pancreas transcription factor 1 |
|
ENSP00000326391.2 |
class A basic helix-loop-helix protein 15 |
|
ENSP00000476312.1 |
class A basic helix-loop-helix protein 15 |
|
ENSP00000294339.3 |
T-cell acute lymphocytic leukemia protein 1 |
|
ENSP00000360951.1 |
T-cell acute lymphocytic leukemia protein 1 |
|
ENSP00000455444.1 |
transcription factor 24 |
|
ENSP00000264824.3 |
protein lyl-1 |
|
ENSP00000362777.3 |
protein atonal homolog 7 |
|
ENSP00000296096.5 |
transcription factor 23 |
|
ENSP00000317580.4 |
neurogenin-1 |
|
ENSP00000317333.3 |
neurogenin-2 |
|
ENSP00000242462.4 |
neurogenin-3 |
|
ENSP00000266744.3 |
achaete-scute homolog 1 |
|
ENSP00000302216.3 |
protein atonal homolog 1 |
|
ENSP00000469019.2 |
achaete-scute homolog 5 |
|
ENSP00000472681.1 |
achaete-scute homolog 5 |
|
ENSP00000345420.4 |
achaete-scute homolog 4 |
|
ENSP00000275461.3 |
fer3-like protein |
|
ENSP00000295108.3 |
neurogenic differentiation factor 1 |
|
ENSP00000306754.4 |
neurogenic differentiation factor 2 |
|
ENSP00000435770.1 |
achaete-scute homolog 3 |
|
ENSP00000297142.3 |
neurogenic differentiation factor 6 |
|
ENSP00000242994.3 |
neurogenic differentiation factor 4 |
|
ENSP00000318799.1 |
class E basic helix-loop-helix protein 22 |
|
ENSP00000332293.4 |
achaete-scute homolog 2 |
|
ENSP00000333097.6 |
factor in the germline alpha |
|
ENSP00000359371.2 |
class E basic helix-loop-helix protein 23 |
|
ENSP00000480998.1 |
class E basic helix-loop-helix protein 23 |
|
ENSP00000416986.1 |
twist-related protein 1 |
|
ENSP00000371794.3 |
oligodendrocyte transcription factor 2 |
|
ENSP00000356708.2 |
oligodendrocyte transcription factor 3 |
|
ENSP00000300057.4 |
mesoderm posterior protein 1 |
|
ENSP00000304676.3 |
protein atonal homolog 8 |
|
ENSP00000342392.3 |
mesoderm posterior protein 2 |
|
ENSP00000228641.3 |
myogenic factor 6 |
|
ENSP00000363689.5 |
DNA-binding protein inhibitor ID-3 |
|
ENSP00000489102.1 |
DNA-binding protein inhibitor ID-3 |
|
ENSP00000228644.3 |
myogenic factor 5 |
|
ENSP00000204517.6 |
Transcription factor AP-4 |
|
ENSP00000371785.1 |
oligodendrocyte transcription factor 1 |
|
ENSP00000250003.3 |
myoblast determination protein 1 |
|
ENSP00000452286.1 |
protein max isoform X4 |
|
ENSP00000375248.1 |
class A basic helix-loop-helix protein 9 |
|
ENSP00000378104.2 |
protein max isoform X3 |
|
ENSP00000451907.1 |
protein max isoform X1 |
|
ENSP00000452378.1 |
protein max isoform |
|
ENSP00000480127.1 |
protein max isoform X1 |
|
ENSP00000452197.1 |
protein max isoform X2 |
|
ENSP00000450818.1 |
protein max isoform X4 |
|
ENSP00000284165.6 |
protein max isoform X1 |
|
ENSP00000351175.4 |
protein max isoform X2 |
|
ENSP00000351490.4 |
protein max isoform X1 |
|
ENSP00000491476.1 |
Myc proto-oncogene protein |
|
ENSP00000234091.4 |
DNA-binding protein inhibitor ID-2 |
|
ENSP00000379585.1 |
DNA-binding protein inhibitor ID-2 |
|
ENSP00000385465.2 |
DNA-binding protein inhibitor ID-2 |
|
ENSP00000405062.1 |
helix-loop-helix protein 2 |
|
ENSP00000433571.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000436313.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000281043.3 |
N-myc proto-oncogene protein isoform X1 |
|
ENSP00000365280.3 |
DNA-binding protein inhibitor ID-1 |
|
ENSP00000365273.3 |
DNA-binding protein inhibitor ID-1 |
|
ENSP00000367207.3 |
Myc proto-oncogene protein |
|
ENSP00000367972.3 |
DNA-binding protein inhibitor ID-4 |
|
ENSP00000430235.1 |
myc proto-oncogene protein |
|
ENSP00000479618.2 |
myc proto-oncogene protein |
|
ENSP00000478887.1 |
myc proto-oncogene protein |
|
ENSP00000484441.1 |
sterol regulatory element-binding protein 2 |
|
ENSP00000395728.1 |
sterol regulatory element-binding protein 2 |
|
ENSP00000281047.3 |
mesogenin-1 |
|
ENSP00000354476.4 |
sterol regulatory element-binding protein 2 |
|
ENSP00000361943.3 |
hairy/enhancer-of-split related with YRPW motif-like protein |
|
ENSP00000379100.4 |
sterol regulatory element-binding protein 1 |
|
ENSP00000470081.1 |
upstream stimulatory factor 2 |
|
ENSP00000438545.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSP00000445836.1 |
ARNTL2 protein |
|
ENSP00000385581.3 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000455289.1 |
CCR4-NOT transcription complex subunit 1 |
|
ENSP00000457869.1 |
CCR4-NOT transcription complex subunit 1 |
|
ENSP00000385915.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000261178.5 |
ARNTL2 protein |
|
ENSP00000442438.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSP00000400185.2 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSP00000379238.2 |
ARNTL2 protein |
|
ENSP00000374357.4 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000384517.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000385897.3 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSP00000312247.5 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSP00000266503.5 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSP00000174618.4 |
max-binding protein MNT |
|
ENSP00000420752.1 |
basic helix-loop-helix domain-containing protein USF3 |
|
ENSP00000241651.4 |
myogenin |
|
ENSP00000368429.3 |
upstream stimulatory factor 2 |
|
ENSP00000340633.4 |
upstream stimulatory factor 2 |
|
ENSP00000262901.4 |
single-minded homolog 1 |
|
ENSP00000358210.3 |
single-minded homolog 1 |
|
ENSP00000222305.2 |
upstream stimulatory factor 2 |
|
ENSP00000471511.1 |
upstream stimulatory factor 2 |
Table 4A: Summary of the GO annotation of Homo-Sapiens
|
Gene Id |
Protein |
|
ENSMUSP00000046999.2 |
heart and neural crest derivatives expressed transcript 1 |
|
ENSMUSP00000124951.2 |
heart and neural crest derivatives expressed transcript 1 |
|
ENSMUSP00000044983.3 |
dHand protein |
|
ENSMUSP00000086511.5 |
transcription factor 15 |
|
ENSMUSP00000043668.7 |
basic helix-loop-helix transcription factor scleraxis |
|
ENSMUSP00000057489.3 |
helix-loop-helix protein 1 |
|
ENSMUSP00000064355.4 |
helix-loop-helix protein 2 |
|
ENSMUSP00000142746.1 |
helix-loop-helix protein 2 |
|
ENSMUSP00000143362.1 |
helix-loop-helix protein 2 |
|
ENSMUSP00000007949.3 |
twist-related protein 2 |
|
ENSMUSP00000139531.1 |
twist-related protein 2 |
|
ENSMUSP00000040089.5 |
twist-related protein 1 |
|
ENSMUSP00000030124.3 |
T-cell acute lymphocytic leukemia protein 2 |
|
ENSMUSP00000151767.1 |
transcription factor 21 |
|
ENSMUSP00000053178.7 |
transcription factor 21 |
|
ENSMUSP00000027062.5 |
musculin |
|
ENSMUSP00000028068.2 |
pancreas transcription factor 1 |
|
ENSMUSP00000055493.7 |
class A basic helix-loop-helix protein 15 |
|
ENSMUSP00000030489.2 |
T-cell acute lymphocytic leukemia protein 1 |
|
ENSMUSP00000124983.1 |
T-cell acute lymphocytic leukemia protein 1 |
|
ENSMUSP00000125202.1 |
T-cell acute lymphocytic leukemia protein 1 |
|
ENSMUSP00000138827.1 |
transcription factor 24 |
|
ENSMUSP00000006818.2 |
transcription factor 23 |
|
ENSMUSP00000046010.4 |
protein lyl-1 |
|
ENSMUSP00000039801.3 |
protein atonal homolog 7 |
|
ENSMUSP00000029587.7 |
neurogenin-2 |
|
ENSMUSP00000050484.4 |
neurogenin-1 |
|
ENSMUSP00000054054.1 |
neurogenin-3 |
|
ENSMUSP00000020243.7 |
achaete-scute homolog 1 |
|
ENSMUSP00000098903.4 |
protein atonal homolog 1 |
|
ENSMUSP00000151551.1 |
neurogenin-3 |
|
ENSMUSP00000058994.3 |
fer3-like protein |
|
ENSMUSP00000137746.1 |
achaete-scute homolog 5 |
|
ENSMUSP00000137650.1 |
achaete-scute homolog 4 |
|
ENSMUSP00000040364.4 |
neurogenic differentiation factor 1 |
|
ENSMUSP00000041373.6 |
neurogenic differentiation factor 2 |
|
ENSMUSP00000047016.8 |
neurogenic differentiation factor 6 |
|
ENSMUSP00000037702.1 |
achaete-scute homolog 3 |
|
ENSMUSP00000051379.3 |
neurogenic differentiation factor 4 |
|
ENSMUSP00000009392.4 |
achaete-scute homolog 2 |
|
ENSMUSP00000026120.6 |
class E basic helix-loop-helix protein 22 |
|
ENSMUSP00000113012.1 |
achaete-scute homolog 2 |
|
ENSMUSP00000104506.1 |
class E basic helix-loop-helix protein 23 |
|
ENSMUSP00000032070.3 |
factor in the germline alpha |
|
ENSMUSP00000036797.8 |
oligodendrocyte transcription factor 2 |
|
ENSMUSP00000057106.5 |
oligodendrocyte transcription factor 3 |
|
ENSMUSP00000036981.7 |
protein atonal homolog 8 |
|
ENSMUSP00000032760.5 |
mesoderm posterior protein 1 |
|
ENSMUSP00000103017.1 |
mesoderm posterior protein 2 |
|
ENSMUSP00000047529.3 |
myogenic factor 6 |
|
ENSMUSP00000008016.2 |
DNA-binding protein inhibitor ID-3 |
|
ENSMUSP00000000445.1 |
myogenic factor 5 |
|
ENSMUSP00000061408.5 |
oligodendrocyte transcription factor 1 |
|
ENSMUSP00000005862.7 |
transcription factor AP-4 |
|
ENSMUSP00000072330.1 |
myoblast determination protein 1 |
|
ENSMUSP00000054158.3 |
protein S-Myc |
|
ENSMUSP00000080778.5 |
protein max isoform X2 |
|
ENSMUSP00000106025.3 |
protein max isoform X1 |
|
ENSMUSP00000151816.1 |
SIM1 |
|
ENSMUSP00000050516.1 |
class A basic helix-loop-helix protein 9 |
|
ENSMUSP00000092019.4 |
DNA-binding protein inhibitor ID-1 |
|
ENSMUSP00000045993.7 |
N-myc proto-oncogene protein |
|
ENSMUSP00000114225.1 |
N-myc proto-oncogene protein |
|
ENSMUSP00000123821.1 |
myc proto-oncogene protein |
|
ENSMUSP00000123852.1 |
myc proto-oncogene protein |
|
ENSMUSP00000124758.1 |
myc proto-oncogene protein |
|
ENSMUSP00000141139.1 |
myc proto-oncogene protein |
|
ENSMUSP00000130285.1 |
myc proto-oncogene protein |
|
ENSMUSP00000140183.2 |
myc proto-oncogene protein |
|
ENSMUSP00000022971.7 |
myc proto-oncogene protein |
|
ENSMUSP00000021810.1 |
DNA-binding protein inhibitor ID-4 |
|
ENSMUSP00000105449.1 |
DNA-binding protein inhibitor ID-1 |
|
ENSMUSP00000121170.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSMUSP00000020974.6 |
DNA-binding protein inhibitor ID-2 |
|
ENSMUSP00000152052.1 |
DNA-binding protein inhibitor ID-2 |
|
ENSMUSP00000023100.6 |
sterol regulatory element-binding protein 2 |
|
ENSMUSP00000152069.1 |
DNA-binding protein inhibitor ID-2 |
|
ENSMUSP00000055001.1 |
mesogenin-1 |
|
ENSMUSP00000040576.4 |
hairy/enhancer-of-split related with YRPW motif |
|
ENSMUSP00000107265.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSMUSP00000107263.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSMUSP00000079373.4 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSMUSP00000107266.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 2 |
|
ENSMUSP00000147764.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSMUSP00000046235.7 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSMUSP00000147989.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSMUSP00000147823.1 |
aryl hydrocarbon receptor nuclear translocator-like protein 1 |
|
ENSMUSP00000000291.2 |
max-binding protein MNT |
|
ENSMUSP00000118435.1 |
max-binding protein MNT |
|
ENSMUSP00000027730.4 |
myogenin |
|
ENSMUSP00000132833.1 |
upstream stimulatory factor 2 |
|
ENSMUSP00000020071.3 |
single-minded homolog 1 |
|
ENSMUSP00000118475.1 |
NEDD4-binding protein 2-like 2 |
|
ENSMUSP00000132021.1 |
upstream stimulatory factor 2 |
|
ENSMUSP00000125520.2 |
upstream stimulatory factor 2 |
|
ENSMUSP00000132256.1 |
upstream stimulatory factor 2 |
Table 4B: Summary of the GO annotation of Mus Musculus
Discussion
The above study epitomized that the subclass bHLH transcription factors of eHAND and dHAND are present in two mammalian genomes. The neuron and muscle-specific transcription factors of eHAND and dHAND are consistent in the human heart and conserved across species. The observation demonstrated the HAND1 in the left and HAND2 in the right ventricles, and both side patterns correlated with the growth of ventricles (left/ right) during maturation. The preface of eHAND and dHAND expression in embryogenesis and cardiac hypertrophy suggested that the HANDs (heart and neural crest derivatives) transcription factors (TFs) are essential for maintaining cardiomyocyte phenotype [11]. The function of HANDs subclass bHLH TFs allowed the hypertrophic genetic facts. The downregulated response of eHAND reduces the prior onset of cardiac hypertrophy. However, subtle difference responses between eHAND and dHAND involve neural-humoral mechanisms that govern the growth of cardiac hypertrophy and the development of heart disease. Therefore, the notable finding of the above study is the differences between eHAND and dHAND in hypertrophy. The eHAND and dHAND express in the earliest onset of cardiogenesis in a non-restricted pattern. In humans, limited evidence suggested that embryological expression of adult samples demonstrated a non-restricted response of eHAND and dHAND in ventricles [11].
The observation of eHAND and dHAND in the ventricle raises questions: (a) why exhibit chamber-restrictive expression during development? (b) why do expression factors throughout a heart in an adult? Because the mitigation of eHAND and dHAND matches embryological heart expression. The chamber-restricted variation of vertebrates suggested that eHAND and dHAND have an overlapping function to allow survival. The overlapping response appears from both genes (eHAND and dHAND) co- express throughout the heart [1]. Also, the combined expression of eHAND and dHAND disrupts cardiac growth and suggests functional redundancy. The dHAND function in the developing heart and limbs documented in biology [14]. So additional functions and affirm of embryonic ventricles provided evidence of the distinct function of eHAND and dHAND in cells and suggested partially overlapping functions [1,3,7,9]. The possible differences in eHAND and dHAND reflect an aspect of their non- overlapping functions. But an overlapping function of eHAND and dHAND involves with dimerization factors [15]. Also, the eHAND and dHAND form heterodimers with class A and B proteins [16]. The functional overlaps involve with dimerization with a prevalent factor exhibiting dominant function. The result of the dominant response drives gene expression. Nevertheless, during trophoblast development, eHAND play a dominant role in the terminal differentiation of trophoblast cell via exit from the cell cycle. So, the downregulated expression of eHAND treats cardiomyopathies, a role of a negatively regulated hypertrophic gene program. If eHAND reveals a dominant role in terminal differentiation and activates the hypertrophic gene program, the simple term is cell growth with less cell division. Then eHAND and dHAND expression may reduce cardiac hypertrophy. The hypothesis is a constitutive expression of eHAND and dHAND in the ventricles through a promoter response to ensure hypertrophic signals. If a similar analysis performs in such experimental animals, the answer to this question might solve. The above study demonstrated eHAND and dHAND expression in adult hearts. Also, the down-regulated expression of the HANDs reveals chamber-specific hypertrophy in ventricles [2,4, 15]. Therefore, the molecular mechanisms initiating the re-expression of the fetal cardiac program may significantly improve treatments for heart disease.
Highlights
• The study associated with organism development.
• The neuron and muscle-specific bHLH TFs are responsible for growth.
• The eHAND and dHAND response may be an early marker for cardiac development.
• Also, the HANDs and other bHLH domains lead to the development of organogenesis.
• The method and analysis data may be valuable for biological databases and clinical research.
Conclusion
Over the last decades, a wealth of new reports has been composed of unique genetic and phenotypic characteristics of cardiac morphogenesis. The mutational analysis of the eHAND and dHAND transcription factors enabled precise resolution of specialized function during the developing myocardium. Also, analysis data concluded the muscle-specific transcription factors eHAND and dHAND are associated with cardiac disease and development. In contrast, the tissue-specific bHLH and other TFs lead to the development of myogenesis and vasculogenesis [17-25].
Declaration
Ethics Approval and Consent to Participate
The study contains an in-silico analysis of the mammalian genome for investigation and upgradation of particular genes in different organisms.
Consent for Publication
The work presented in this paper is original and carried out by the author addressed in the manuscript. The author declares that the document is not concerned elsewhere and is not under simultaneous consideration by other journals.
Availability of Data and Material
The data and samples are not deposited in the database but are available on reasonable request or demand.
Funding
The author availed financial assistance through a National Fellowship from the University Grant Commission (UGC) and the Ministry of Social Justice and Empowerment, Government of India.
Author Contribution
The above research paper contains the sole author. The author proposed the idea, experimented, analysed the data and prepared the manuscript.
Dedication
The author dedicated this article to his beloved friend Catherine Tresa Alexander
Acknowledgement
The author is grateful to Assam University, Silchar, Assam, India, for providing the requisite lab facilities for completing this research work.
References
- Srivastava, D., Thomas, T., Lin, Q., Kirby, M. L., Brown, D., & Olson, E. N. (1997). Regulation of cardiac mesodermal and neural crest development by the bHLH transcription factor, dHAND. Nature genetics, 16(2), 154-160.
- Firulli, A. B., McFadden, D. G., Lin, Q., Srivastava, D., & Olson, E. N. (1998). Heart and extra-embryonic mesodermal defects in mouse embryos lacking the bHLH transcription factor Hand1. Nature genetics, 18(3), 266-270.
- Riley, P., Anaon-Cartwight, L., & Cross, J. C. (1998). The Hand1 bHLH transcription factor is essential for placentation and cardiac morphogenesis. Nature genetics, 18(3), 271-275.
- Yelon, D., Ticho, B., Halpern, M. E., Ruvinsky, I., Ho,R. K., Silver, L. M., & Stainier, D. Y. (2000). The bHLH transcription factor hand2 plays parallel roles in zebrafish heart and pectoral fin development. Development, 127(12), 2573-2582.
- Thattaliyath, B. D., Livi, C. B., Steinhelper, M. E., Toney,G. M., & Firulli, A. B. (2002). HAND1 and HAND2 are expressed in the adult-rodent heart and are modulated during cardiac hypertrophy. Biochemical and biophysical research communications, 297(4), 870-875.
- Cserjesi, P., Brown, D., Lyons, G. E., & Olson, E. N. (1995). Expression of the novel basic helix-loop-helix gene eHAND in neural crest derivatives and extraembryonic membranes during mouse development. Developmental biology, 170(2), 664-678.
- Cross, J. C., Flannery, M. L., Blanar, M. A., Steingrimsson, E., Jenkins, N. A., Copeland, N. G., ... & Werb, Z. (1995). Hxt encodes a basic helix-loop-helix transcription factor that regulates trophoblast cell development. Development, 121(8), 2513-2523.
- McFadden, D. G., Charité, J., Richardson, J. A., Srivastava, D., Firulli, A. B., & Olson, E. N. (2000). A GATA-dependent right ventricular enhancer controls dHAND transcription in the developing heart. Development, 127(24), 5331-5341.
- Ritter, O., Haase, H., Schulte, H. D., Lange, P. E., & Morano, I. (1999). Remodeling of the hypertrophied human myocardium by cardiac bHLH transcription factors. Journal of cellular biochemistry, 74(4), 551-561.
- Natarajan, A., Yamagishi, H., Ahmad, F., Li, D., Roberts, R., Matsuoka, R., ... & Srivastava, D. (2001). Human eHAND, but not dHAND, is down-regulated in cardiomyopathies.Journal of molecular and cellular cardiology, 33(9), 1607- 1614.
- Scott, I. C., Anson-Cartwright, L., Riley, P., Reda, D., & Cross, J. C. (2000). The HAND1 basic helix-loop-helix transcription factor regulates trophoblast differentiation via multiple mechanisms. Molecular and cellular biology, 20(2), 530-541.
- Cross, J. C. (2000, April). Genetic insights into trophoblast differentiation and placental morphogenesis. In Seminars in cell & developmental biology (Vol. 11, No. 2, pp. 105-113). Academic Press.
- Murre, C., Bain, G., van Dijk, M. A., Engel, I., Furnari, B.A., Massari, M. E., ... & Stuiver, M. H. (1994). Structure and function of helix-loop-helix proteins. Biochimica et Biophysica Acta (BBA)-Gene Structure and Expression, 1218(2), 129-135.
- Thattaliyath, B. D., Firulli, B. A., & Firulli, A. B. (2002). The basic-helix-loop-helix transcription factor HAND2 directly regulates transcription of the atrial naturetic peptide gene. Journal of molecular and cellular cardiology, 34(10), 1335-1344.
- Firulli, B. A., Hadzic, D. B., McDaid, J. R., & Firulli, A.B. (2000). The Basic Helix-Loop-Helix Transcription factorsdHAND and eHAND Exhibit Dimerization Characteristics That Suggest Complex Regulation of Function. Journal of Biological Chemistry, 275(43), 33567- 33573.
- Mitchell, P. J., & Tjian, R. (1989). Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins. Science, 245(4916), 371-378.
- Massari, M. E., & Murre, C. (2000). Helix-loop-helix proteins: regulators of transcription in eucaryotic organisms. Molecular and cellular biology, 20(2), 429-440.
- Amoutzias, G. D., Robertson, D. L., Van de Peer, Y., & Oliver, S. G. (2008). Choose your partners: dimerization in eukaryotic transcription factors. Trends in biochemical sciences, 33(5), 220-229.
- Chaudhary, J., & Skinner, M. K. (1999). Basic helix-loop- helix proteins can act at the E-box within the serum response element of the c-fos promoter to influence hormone- induced promoter activation in Sertoli cells. Molecular endocrinology, 13(5), 774-786.
- Latchman, D. S. (1997). Transcription factors: an overview. The international journal of biochemistry & cell biology, 29(12), 1305-1312.
- Karin, M. (1990). Too many transcription factors: positive and negative interactions. The new biologist, 2(2), 126-131.
- Ptashne, M., & Gann, A. (1997). Transcriptional activation by recruitment. Nature, 386(6625), 569-577.
- Babu, M. M., Luscombe, N. M., Aravind, L., Gerstein, M., & Teichmann, S. A. (2004). Structure and evolution of transcriptional regulatory networks. Current opinion in structural biology, 14(3), 283-291.
- Brivanlou, A. H., & Darnell Jr, J. E. (2002). Signal transduction and the control of gene expression. Science, 295(5556), 813-818.
- McFadden, D. G., McAnally, J., Richardson, J. A., Charité, J., & Olson, E. N. (2002). Misexpression of dHAND induces ectopic digits in the developing limb bud in the absence of direct DNA binding. Development, 129(13), 3077-3088.