Research Article - (2022) Volume 3, Issue 3
Diversity and Distribution of Spa Types among Methicillin Resistant Staphylococcus Aureus Isolated From Humans and Livestock in Kabale District-South Western Uganda
2Department of Medical Laboratory Sciences, Faculty of Health Sciences, Muni University, P.O. Box 725, Arua, Uganda
3Department of Microbiology, University of Global Health Equity. P.O. Box 6955, Kigali, Uganda
4Department of Microbiology, Faculty of Medicine, Mbarara University of Science and Technology, P.O. Box 1410, Mbarara, Uganda
Received Date: Aug 15, 2022 / Accepted Date: Aug 22, 2022 / Published Date: Sep 09, 2022
Copyright: ©Copyright: ©2022 Andrew Baguma. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Baguma A, Musinguzi B, Orikiriza P, and Bazira J. (2022). Diversity and Distribution of Spa Types among Methicillin Resistant Staphylococcus Aureus Isolated From Humans and Livestock in Kabale District - South Western Uganda. J Vet Heal Sci, 3(3), 283-290.
Abstract
Background: S. aureus is a skin and mucosal bacterial commensal of both humans and animals which has evolved as an important pathogen implicated to cause various infections. High levels of antibiotic use have resulted into multi-drug resistance MRSA, especially among HA-MRSA, CA-and LA - MRSA. Awareness on coexistence and diversity of MRSA clones among humans and household Livestock particularly cattle and swine in our region is limited. We used spa typing method to determine spa diversity, distribution and coexistence in outpatients, household contacts and respective livestock (cattle and swine) in Kabale region, south western Uganda.
Methods: This was a cross sectional study by design consisting of outpatients, household contacts and livestock. Outpatients (n =100) colonized with MRSA were traced back to their respective homesteads where household members, domestic cattle, and, swine were tested for S. aureus and subsequently MRSA colonization. High-resolution DNA melting analysis was used to determine spa types among MRSA isolates. Overlap of MRSA isolates among humans and livestock was based on the presence of similar spa types.
Results: A total of 3371 S.aureus isolates were collected from outpatients (n =376), household contacts (n = 1531), Cattle (n = 1159) and Swine (n = 305), among which 482 had mecA gene where 27% (100/376) and 8% (123/1531) were outpatients and household contacts respectively while 11% (132/1159) and 42% (127/305) were cattle and swine respectively. Twenty different spa types were identified; t034, t4677, t108, t1451, t9377, t1081, t040, t701, t041, t002, t044, t037,t121, t127, t922, t032, t019, t018, t012 and t030, among which t034 (109/482), t4677 (53/482), t9377 (63/482) and t1081 (53/482) were most prevalent and distributed among human and livestock. All the MRSA isolates were multidrug resistant to antibiotics tested.
Conclusion: In Kabale region, there is high diversity of spa types among MRSA. Presence of similar spa types was found circulating among humans and their respective livestock which demonstrates a possible bidirectional transmission. Presence of MDR - MRSA highlights the need for effective prevention and control of MRSA among livestock and in the community using One Health approach.
Keywords
MRSA, spa types, diversity and distribution.
Background
S. aureus is a skin and mucosal bacterial commensal of both hu¬mans and animals which has evolved as an important pathogen implicated to cause various infections. Certainly, some strains have become methicillin resistant commonly known as methi¬cillin-resistant S. aureus (MRSA) upon acquisition of the Staph¬ylococcal Chromosomal Cassette mec (SCCmec) mobile genetic element [1]. SCCmec harbours mec A or mec C, both of which encode alternate penicillin- binding proteins, which mediate re¬sistance to almost all β-lactam antibiotics. The problem is that MRSA can resist several other classes of antibiotics, limiting the choice of treatment options [2]. High levels of antibiotic use in healthcare settings resulted into HA-MRSA among the isolates. The multi-drug resistance posed by MRSA, especially among the Healthcare associated MRSA (HA-MRSA) presents a serious public threat [3]. HA-MRSA infections are associated with higher mortality and prolonged lengths-of-stay, thus mak¬ing the control rather compelling.
Coexistence of MRSA isolates carrying SCCmec types IV or V (CA-MRSA) and SCCmec types I, II, or III (HA-MRSA) due to hospital-community interactions have been described before in Uganda [4]. Community and Livestock interactions in our region are common and therefore, presence of Livestock – associated (LA - MRSA) in human and vice versa needs to be investigated. However, Livestock associated MRSA (LA-MRSA) is known to cause clinical infections in humans [4]. MRSA transmission from animals to humans (zoonoses) and vice versa (zooanthro-ponosis) has been reported, and direct contact with livestock and other animals is associated with transmission and spread [1]. High prevalence of multi-drug resistant MRSA derived from livestock, particularly among the swine has been previously re¬ported in Kabale region and elsewhere [2-4]. The existence of MRSA carrying SCCmec types IV or V (CA- MRSA) and LA – MRSA clones in hospital settings is of serious concern. These may harbor genes that encode other non-β-lactam antimicrobial resistance genes especially those that led to aminoglycosides, macrolides, lincosamides and fluoroquinolones resistance or en¬hanced pathogenicity [5].
Knowledge on coexistence, diversity, and distribution of MRSA clones among humans and household Livestock particularly cat¬tle and swine in our region is limited due to under resourced lab¬oratories to provide meaningful data [6]. Several tools such as pulsed-field gel electrophoresis (PFGE), multi-locus sequence typing (MLST), staphylococcal cassette chromosome typing and DNA microarray hybridization may provide meaningful data to this phenomenon and constantly provides epidemiologi¬cal surveillance. However, these tools are expensive for routine use in our settings. Therefore, we chose spa typing method, a cheaper technique with high degree of typing ability, excellent reproducibility, providing interchangeable information and can distinguish relapse from re-infection among human and live¬stock. In our one health perspective, the aim of this study was to determine the MRSA carriage rate, spa diversity, distribution and coexistence in outpatients, household contacts and respec¬tive livestock (cattle and swine) in Kabale region, south western Uganda.
Materials and Methods
Study Subjects and Design
This was a cross sectional study, conducted between June 2016 and June 2018 and it included persons who were seeking medi¬cal care, diagnostics and treatment at Kabale Regional Referral Hospital (KRRH) outpatient unit without staying in the Hospital overnight. These personnel were labeled as outpatients. In addi¬tion, other study subjects included family (household) members and livestock (cattle and swine) of the outpatients who were test¬ed and found to have nasopharyngeal MRSA colonization. The inclusion criteria were: i) MRSA positive outpatient; ii) owner¬ship of cattle, swine or both in their respective homes.
Sample Size Determination
The minimum sample size required to accurately assess outpa¬tient MRSA carriage was estimated to be 384 using Kish and Lisle (1965) formula, basing on the 51% prevalence of MRSA community nasal colonization according to [7]. The sample size of 384 out patients attending KRRH were further translate into 1536 household contacts since the average number of people per homestead in Kabale region is estimated to be 04 according to National population census of 2014(https://www.ubos.org/). In addition, according to Ministry of Agriculture, Animal Indus-tries and Fisheries (https://www.agriculture.go.ug/) it is estimat¬ed that in Kabale region; each household has an average of 04 cattle, and 01 pig. Therefore, we estimated 1232 cattle and 308 pigs for inclusion in the study.
Sample Collection and Processing
The informed consent were obtained from all participants in-cluding Household heads who consented on behalf of animal subjects before commencement of the study. The collections of all nasopharyngeal swabs from human and animal subjects were not invasive with full respect of Human and animal rights following standard ethical guidelines. After consenting, we pro¬spectively collected nasopharyngeal swab from all outpatients attending care at Kabale Regional Referral Hospital (KRRH) using sterile swabs (Fisherbrand™). The samples were trans¬ported at 2 - 8°C in a cold box to the microbiology laboratory at KRRH within 6 hours for processing and isolation of MRSA. Outpatients whose nasopharyngeal swabs had MRSA isolated were followed up to their respective homesteads, where similar samples were collected among family members (Households) and their respective cattle, swine or both. Cattle and swine nasal swabs were collected from the upper nasal cavity using 6 – inch sterile cotton swabs after restrain. In brief, the by wiping off the snout with sterile gauze and inserted sterile swab deep into na¬sal cavity taking care not get in contact with the outside of the nostril. The swab was rotated hard enough on the inside of the nose to collect the sample. The swabs were transported at 2 - 8°C to the laboratory for microbiological processing and isolation of MRSA. The KRRH laboratory is a quality controlled labo-ratory that participates in the national quality assurance scheme conducted by the Central public Health Laboratory-Ministry of Health, Uganda (CPHL-MOH).
Microbiological Analysis
Phenotypic Characterization of S. aureus
Nasopharyngeal swabs collected from both humans and domes¬tic animals (cattle and swine) were inoculated onto Mannitol salt broth (MSB) (Oxoid™UK) and incubated aerobically at 37oC for 24 hours. Positive broth cultures were subcultured onto Manni¬tol salt agar (MSA) media (Oxoid™UK) and incubated at 37°C for 24 - 72 hours until appropriate growth was observed. Every new batch of MSB and MSA used to isolate S. aureus was qual¬ity controlled using control strains; S. aureus ATCC 6538 and Escherichia coli ATCC 8739 (MicrobiologicsTMUSA) as posi¬tive and negative controls respectively. Yellow bacterial colonies from MSA media were sub-cultured onto 5% sheep blood agar and incubated at 37°C for 24 hours. The colonies were evaluated by colonial morphology (yellow pigmentation), gram staining reaction, catalase and coagulase (free and bound) production as well as DNase test [8]. The same colonies were further identi¬fied using API® Staph (BioMérieux SA). All the phenotypically confirmed S. aureus isolates were further screened for methi¬cillin resistance using cefoxitin (30μg) disk (BioMérieux SA) diffusion where the isolates with zone diameter of ≤ 21mm and ≥ 22mm were labeled as MRSA and MSSA respectively following Clinical and Laboratory Institute Standard protocol [9]. All the phenotypic MRSA isolates were further confirmed using molec¬ular analysis by PCR.
Molecular Analysis
DNA Extraction
After three months of storage at -80°C, MRSA isolates were sub-cultured onto 5% sheep blood agar (SBA) and incubated at 37±2oC for 24 hours to obtain pure colonies. DNA was ex¬tracted in accordance with the protocol for PCR amplification of mecA, mecC (MECALGA251), spa and lukF-PV, lukS-PV genes as recommended by the EURL-AR2st version, September 2012 [10]. Briefly, about 1-2 pure colonies were suspended in 25µl of sterile distilled water and boiled at 100°C in a digital heat block (Thermo Scientific™) for 15 minutes followed by centrifugation at 15,000 × g for 15 minutes. The supernatant was removed and the pellet suspended 100µLof molecular biology-grade water (Eppendorf, Hamburg, Germany). This was further centrifuged at 15,000 × g for 10 minutes. The supernatant was eliminated and the remaining pellet was resuspended in 40μL of molecular biology-grade water and again subjected to boiling at 100°C in a water bath for 10 minutes. This was cooled on ice and centri-fuged at 15,000 × g for 10 seconds before freezing at -20°C.
PCR detection of the MRSA
Amplification for MRSA based on mecA gene was done ac-cording to previously published methods[10], [11]. The prim¬er sequences were: MecA F (5'-TCCAATTACAACTTCAC-CAGG-3' and MecA R (5'-CCACTTCATATCTTGTAACG3'”. A 50µl PCR reaction mixture was used which included; 45µl of master mix (Invitrogen, Carlsbad, CA, USA) containing PCR buffer (x1), dNTP mix (0.2mM of each), primer (0.5µM), Taq DNA polymerase (0.25U), and MgCl 2 (1.5mM) with 5µL of template DNA. PCR amplifications were performed under the following cycling conditions: Hot start at 94oC for 4 minutes, followed by 30 cycles of denaturation at 94oC for 45 seconds, annealing at 50oC for 45 seconds, and extension lead at 72oC for 1 minute and final extension lead at 72oC for 3 minutes. The PCR products were analyzed using electrophoresis on 2% aga¬rose gel (Invitrogen, Carlsbad, CA, USA) premixed with ethid-ium bromide dye (0.5µg/mL) and visualized using UV transillu-minator. Amplicon size of approximately 180bp was consistent with mecA gene amplification and was scored as MRSA posi¬tive. All samples were analyzed alongside; i) known MRSA Pos¬itive control (MRSA ATCC 43300); ii) negative control (MSSA contains a Methicillin Susceptible S. aureus strain ATCC 25923 and iii) negative control (all PCR components without the DNA template). All the confirmed MRSA positive strains were packed and preserved in cryovial tubes, containing 1.5ml of 30% glyc-erol mixed with brain heart infusion (BHI) broth (Oxoid™UK) and stored at -80oC for further antibiotic susceptibility testing and spa typing.
Spectrum and Antiprogram Testing
The MRSA isolates were subjected to duplicate antibiotic sus¬ceptibility test using Kirby Bauer’s disk diffusion techniques. The following antibiotics were used: Chloramphenicol (30μg), Trimethoprim – Sulfamethoxazole (1.25/23.75μg), linezol¬id (30 μg), Rifampin (5 μg), tetracycline (30 μg), gentamicin (10 μg), Ciprofloxacin (5 μg), and Clindamycin (2μg). The antibiotic selection and susceptibility scoring were based on the Clinical and Laboratory Standards Institute (CLSI) guide-lines[9]. All MRSA isolates were further tested for van A/B gene by film Arrays method using BioFire FilmArray Multiplex PCR System (BioMérieux. USA). S. aureus ATCC 29213 strains were used as vancomycin-susceptible controls and Enterococcus fae-calis ATCC 51299 as vancomycin resistant control. However, we could not confirm the resistance determinants due to limited testing capacity in our setting.
Spa Typing
All MecA positive isolates (n = 482) were further analyzed for spa typing where the polymorphic X region of the spa gene was amplified in a Rotor-Gene Q instrument (Qiagen), using Type-it HRM PCR Kit (QIAGEN®) and the melting temperature (Tm). The melting curve of every amplicon was analyzed in close tubes using Rotor-Gene ScreenClust HRM Software following standard protocol [12]. In brief, a 2.0-ml PCR reaction was set up, containing 0.8 ml Eva-Green, 1.0 ml SensiMix, 1 ml of each primer (100 mM; 1095 spa forward 5’- AGACGATCCTTCG-GTGAG-3’ and 1517spa reverse 5’-GCTTTTGCAATGCAA TGTCATTTACTG-3’, and 20ng of the template DNA; this was programmed as follows: a hold at 95oC for 10 min, followed by 35 cycles of 95oC for 20 seconds, 56oC for 20 seconds, and 72oC for 22 seconds. The high-resolution melting analysis of the amplicons was performed between 70oC and 95oC with a stepwise increase of 0.05oC/s with 25 acquisitions per degree. Extra DNA was added to the reaction mixture to acquire distinct melting curves per spa types and consequently improving as¬say performance. Optimal performance was achieved by adding 0.5ng DNA of spa type t003 and 0.5ng DNA of spa type t030 to the reaction mixtures. The melting temperatures (Tm) were determined by the negative derivative of decreased fluorescence over increased temperature (df/dt), using Rotor-Gene Screen-Clust HRM Software which also allowed visualization of the melting curves shapes. The identified spa types were recorded and distributed according to MRSA source and aggregated in accordance to their frequency of occurrence (Table 02). The spa types indicated by the melting temperatures were obtained from various publications [13-16]. However, the sequence-based spa typing and MLST were not performed.
Results
MRSA Carriage Rate A total of 3371 bacterial isolates were collected from outpatients (376), household contacts (1531), Cattle (1159) and Swine (305) as in Table 1. The prevalence of S. aureus in outpatients, house hold contact, cattle and swine were 33%, 30%, 84% and 44% respectively. Among the S. aureus positive isolates, 482 isolates were mecA positive and were designated as MRSA. MecA gene was detected in 27% (100/376) and 8% (123/1531) of outpa¬tients and household contacts respectively while among live¬stock, it was detected in 11% (132/1159) and 42% (127/305) of cattle and swine respectively as shown in Table 1 and figure 1.
Table 1: Prevalence of S. aureus and MRSA carriage among human and livestock Sources
|
Source |
Total (N = 3371) |
S.aureus (N = 1694) |
MecA Positive (N = 482) |
|
Outpatient, % (n/N) |
11.0 (376/3371) |
33.0 (124/376) |
27.0 (100/376) |
|
Household contacts, % (n/N) |
45.0 (1531/3371) |
30.0 (458/1531) |
8.0 (123/1531) |
|
Cattle, % (n/N) |
35.0 (1159/3371) |
84.0 (978/1159) |
11.0 (132/1159) |
|
Swine, % (n/N) |
9.0 (305/3371) |
44.0(134/305) |
42.0 (127/305) |
Note: % percent; N = total number of samples; n = number of cases of interest
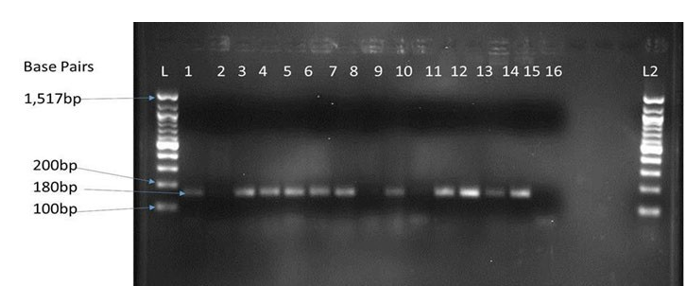
Figure 1: Gel electrophoresis of MecA gene amplified by PCR where an Amplicon of approximately 180bp was expected. L is 100bp ladder,1- Positive control (MRSA ATCC 43300), 2 – negative control (MSSA contains a Methicillin Susceptible S. aureus strain ATCC 25923) Lanes 3, 4,5,6,7,9,11,12,1314 and15 mecA positive (MRSA). From outpatients, were; 3,4, 5and 6 while HHC; 7 and 9, Cattle; 11 and 12 and swine; 14 and 15. Lane 8, 10 and 16 mecA negative (MSSA) isolates. L2 is the Negative amplification control.
Spa Type Diversity, Distribution and Coexistence between Human and Livestock
Twenty different spa types were identified among MRSA isolated from human and livestock (Figure 1). The spa types with corre¬sponding clonal complex(CC) according to were; t034(CC398), t4677(CC398), t108(CC398), t1451(CC398), t9377(CC45), t1081(CC45), t040(CC45) t701(CC6), t041(CC5), t002(CC5), t044(CC80), t037(CC8),t121(CC8), t127(CC1), t922(CC1), t032(CC22), t019(CC30), t018(CC30), t012(CC30) and t030(CC8/239). Among these spa types, the most prevalent, be-longs to CC398 i.e. t034 (109/482) and t4677 (53/482), CC45 i.e. were t9377 (63/482) and t1081 (53/482) Among the outpatients and respective household contacts, there were 19 and 17 different spa types respectively. However, 15 and 13 different spa types were identified among cattle and swine respectively (Table 2). The common spa types detected were; t034(n = 109), t9377(n = 63), t1081(n = 55), t4677(n = 53), t701(n = 30), t121(n = 41), t019(n = 28), t002(n = 23), t108(n = 20), t041(n = 11) and t018(n = 11). Other spa types were; t037 (n = 7), t127(n = 7), t1451(n = 7), t044(n = 5 ), t032(n = 3 ), t040(n = 3), t922(n = 2 ) and t012(n = 1). Spa type t044 (n = 5), t032 (n = 3), t030 (n = 3) and t012 (n = 1) were only observed among humans [17].
Table 2: Spa –types detected among MRSA isolated from Human, Cattle and Swine
|
Spa types (n = 20) |
Corresponding clonal complex (CC) |
Spa type distribution among human and livestock |
||||
|
No. of isolates (n =482) |
O u t p a -tients (n = 100) |
Household contacts (n = 123) |
Cattle (n = 132) |
Swine (n = 127) |
||
|
t1081 |
CC45 |
55 |
9 |
14 |
21 |
11 |
|
t701 |
CC6 |
30 |
24 |
2 |
1 |
2 |
|
t034 |
CC398 |
109 |
3 |
25 |
41 |
40 |
|
t041 |
CC5 |
11 |
4 |
5 |
2 |
0 |
|
t044 |
CC80 |
5 |
4 |
1 |
0 |
0 |
|
t037 |
CC8 |
7 |
3 |
2 |
0 |
0 |
|
t4677 |
CC398 |
53 |
11 |
15 |
14 |
16 |
|
t127 |
CC1 |
7 |
2 |
5 |
0 |
0 |
|
t1451 |
CC398 |
7 |
1 |
3 |
2 |
1 |
|
t9377 |
CC45 |
63 |
7 |
15 |
22 |
19 |
|
t032 |
CC22 |
3 |
1 |
2 |
0 |
0 |
|
t121 |
CC8 |
41 |
12 |
9 |
9 |
14 |
|
t922 |
CC1 |
2 |
1 |
1 |
0 |
0 |
|
t019 |
CC30 |
28 |
2 |
6 |
11 |
9 |
|
t108 |
CC398 |
20 |
4 |
6 |
2 |
8 |
|
t002 |
CC5 |
23 |
4 |
9 |
4 |
4 |
|
t030 |
CC8/239 |
3 |
3 |
1 |
0 |
0 |
|
t018 |
CC30 |
11 |
4 |
3 |
2 |
2 |
|
t040 |
CC45 |
3 |
01 |
1 |
1 |
1 |
|
t012 |
CC30 |
1 |
1 |
1 |
0 |
0 |
MRSA Antiprogram
The antibiotic susceptibility pattern of MRSA from humans (outpatients and household contacts) and livestock (cattle and swine) are as presented in Table 2. More than sixty percent of MRSA isolated from each of the sources (Outpatient, household contact, cattle and swine) were resistant to Chloramphenicol, Tetracycline, Gentamycin, Ciprofloxacin, and Trimethoprim–Sulfamethoxazole (Table 3). A high level of resistance to tet¬racycline among MRSA isolates from cattle (97%) and swine (100%) compared to those isolated from humans. Similarly, iso¬lates from human sources showed high resistance to rifampin (54%) as opposed to isolates from cattle (3%) and swine (9%). There was similar resistance pattern to Clindamycin resistance among the isolates from swine (56%), cattle (52%), household contacts (53%) and outpatients (49%). Antibiotic resistance against linezolid was less than 5% among isolates from both human and livestock. Antibiotic resistance against vancomycin was only seen in 2% of the MRSA isolates from swine.
Table 3: Antibiogram of MRSA isolates from humans and Livestock Sources
|
Antibiotics |
MRSA Isolates (n = 482) |
|||
|
Outpatients (n = 100) |
Household contact (n = 123 |
Cattle(n=132) |
Swine (n = 127) |
|
|
Chloramphenicol (30μg) |
67 (67%) |
90 (73%) |
106 (80%) |
114(90%) |
|
Tetracycline (30μg) |
89(89%) |
107(87%) |
128 (97%) |
127(100%) |
|
Gentamycin (10 μg) |
60(60%) |
107(87%) |
108(82%) |
89(70%) |
|
Ciprofloxacin (5 μg), |
78(78%) |
98(80%) |
112(85%) |
119(94%) |
|
Clindamycin (2 μg) |
49(49%) |
65(53%) |
69(52%) |
71(56%) |
|
TMP-SMX (1.25/23.75μg) |
89(89%) |
112(91%) |
95(72%) |
88(69%) |
|
Linezolid (30μg) |
02(2%) |
4(3%) |
01(1%) |
05(4%) |
|
Rifampin (5μg), |
43(43%) |
54(44%) |
12(9%) |
04(3%) |
|
Vancomycin (presence of vanA/B gene) |
00(0%) |
00(0%) |
00(0%) |
03(2%) |
Foot note: TMP/SMX = Trimethoprim/Sulfamethoxazole.
Discussion
This study is unique in its kind since it is the first to systemati-cally sample human and animals interface, investigating MRSA carriage rate, spa diversity and distribution while elaborating coexistence in our region. The presented data provide an insight into the MRSA distribution among outpatients to Household contacts and respective livestock particularly; Cattle and swine. We estimated the MRSA prevalence of 12% among humans and 18% among domestic cattle and swine. This is comparatively higher than prevalence of 1.2% reported in Hamburg, Germa¬ny among outpatients [18]. However, the prevalence of 24.7% reported among outpatients in Tanzania in much more higher than what is reported in our current study. Perhaps, these vari¬ations could be attributed to the differences in antibiotics usage among different settings [19]. Nevertheless, urgent Infection control at the outpatient units and among the healthcare workers should be over emphasized. In addition, our study reports prev¬alence of 42% among the swine population of Kabale region. In comparison, this is in agreement with the provenances of 41% and 41.4% reported in Kebbi, Northwestern Nigeria and Osona (Catalonia, Spain) respectively but significantly higher than prevalence of 29.9% reported in Ontario, Canada [20-22]. Differences in farm hygiene probably attributes to these vari¬ations in prevalence. However, emphasis should be put on to cattle and swine hygiene through constant kraal or pen cleaning, proper sanitation and disinfecting to reduce MRSA colonization. MRSA among livestock has been reported before and its signif¬icance in zoonotic transmissions should not be disregarded [22, 23].The diversity of MRSA is expanding, and detecting lineages of human origin in animals and vice-versa becoming more com¬mon and in our current study, we identified 20 spa types from both humans and Livestock. A similar partner of diversity has been reported in Serbia among community and livestock accord¬ing to [24, 25]. In addition, the current significant phenomenon of spa diversity among humans and respective household do¬mestic animals, particularly cattle and swine has been described before. This suggests significant increase in the diversity with heterogeneity representing imported and local clones among MRSA colonizing human and livestock. Distinguishing MRSA strains colonizing human and livestock plays a big role in the prevention and control of spread emphasizing several reservoirs [18, 20 and 26].
The twenty spa types observed among the human in our study agrees with other reports elsewhere showing presence of mul-tiple spa types among the MRSA isolates [25, 27]. However, differences has been observed in the spa types where we ob-served presence of; t1081, t701, t034, t041, t044, t037, t4677, t127, t1451, t9377, t032, t121, t922, t019, t108, t002, t030, t018, t040 and t012 in our region, while Vanessa et al reported t008, t020, t022, t104, t179, t718, t747, t910, t932, t1094, t2357, t5624, t10683 and t14933 in Portugal [24].This suggests diverse genetic backgrounds and multiple routes of their acquisition and spread. In addition, this is inconsistent with a systematic review of the global distribution of spa types which revealed that t064 and t037 were the most prevalent spa types in Africa yet t064 was not observed and t037 was not significantly high. Kateete et al had previously described t002 and t037 among the com- munity which is consistent with our study, even though we did not observe t4353 and t12939 as previously reported [28]. The diversity of MRSA strains is large and it seems to vary from region to region and may have consequences in the spread con¬trol of these strains between reservoirs. Interestingly, we confirm that LA-MRSA strains including t034, t4677, t1451/CC398 and t007, t019, t018, t012/ CC30 also occur among human MRSA isolates. Zoonotic transmission occurs probably via direct an¬imal contact, environmental contaminations or meat. Human -Livestock close contact possibly intensifies the exchange of bac¬teria between humans and animals resulting into anthroponotic and zoonotic transmissions [21, 22]. LA-MRSA infections among livestock animals and associated farmers are of great concern as these sources could potentially serve as reservoirs for zoonotic infections. Previously, surveillance of LA-MRSA among humans has been focusing on MRSA CC398 and finding of CC30 stresses that other MRSA clonal lineages associated with livestock exists. This is consistent with Kock et al in Ger¬man who reported CC398 and other clonal lineages were major cause for human infection [29]. Our findings should raise the awareness of the risk of transmission of LA-MRSA from animal to farmers in Kabale region. In addition, human spa types/clones has been identified among MRSA from livestock and indication of anthroponosis. Human related MRSA emerging as a frequent colonizer of animal populations is possibly favored by the large antibiotic use in animal husbandry and prolonged or frequent close contact [30]. The ability to prevent infection from animals to humans depends much on good hygiene practices in homes with livestock including safe meat and milk handling, and con¬sistent hand washing after close contact with animal can mini¬mize and control infection. Of concern, some of the spa types identified are related to healthcare-associated MRSA clones (HA –MRSA), particularly t1081, t040, t9377/CC45; t121, t037/ CC8; t041, t002/CC5; t922, t127/CC1 and t701/CC6. Kateete et al had previously reported existence of HA – MRSA clones among pastoral communities in rural western Uganda [28]. The of coexistence of MRSA clones is an interesting ecological and public health problem resulting from the interaction between CA-MRSA and HA-MRSA which may have epidemiological and clinical consequences. According to Kouyos et al, HA-MR-SA displays a broader resistance spectrum than CA-MRSA and very difficult to treat [31].
The association of MRSA with antimicrobial resistance profiles can provide useful information for the clinical treatment of in¬fection. While previous studies have reported high AMR prev¬alence among MRSA [31-34], little is known regarding AMR prevalence among MRSA isolates in Kabale region. We found both human and livestock isolates exhibited a higher AMR with general prevalence of 56% and 54% among MRSA isolates from human and livestock displaying resistance to all the 9 of the antibiotics selected and tested. Multidrug resistance (MDR) patterns similar to what we observed in our study have been reported around east African countries and elsewhere[28, 32]. We deduce that increased availability of over the counter anti¬biotic and their widespread use in the community are probable cause for the high levels of AMR. On the same note, the general AMR prevalence of 54% among the isolates from livestock is alarmingly high in our community. This pattern of resistance is consistent with antimicrobial use in the livestock farming in our region. Similar antibiotic resistance pattern has been previously reported in Morocco [35, 36]. Overuse and inappropriate pre-scription of antibiotics in livestock farming is probable driver of increased AMR. While antibiotic resistance is a natural phenom-enon, however, continuous introduced into the environment ex¬erts pressure on bacteria to resistant strains [37]. Local commu¬nity always use but not veterinary recommended antibiotics into their farms. Strict farm hygiene and judicious antibiotic usage in livestock is necessary reduce the prevalence and incidence of highly antibiotic resistance strains.
Our investigation had some limitations; the study did not use techniques such as PFGE, MLST and SCCmec typing which are more accurate. The unaffordability of these techniques coupled with lengthy turnaround time and result interpretation challeng¬es made us not to use them. In addition, the hospital patients and healthcare workers were not screened during the study, limiting assessment of their potential role in MRSA transmission into the hospital setting. Also, our investigation of the antimicrobial sus¬ceptibility of MRSA isolates did not compare community- with hospital-based resistance patterns.
Conclusion
In Kabale region, there is high diversity of spa types among the MRSA among which spa types; t034, t701, t9377, t4677, t121 are predominant. Similar spa types were identified circulating among humans and their respective livestock (Particularly, cattle and swine), which demonstrates the possibility of bidirection¬al transmission between humans and livestock. There was high level of multi drug resistance (MDR) MRSA which highlights the need for effective prevention and control of MRSA among livestock and in the community using One Health approach. We recommend periodic screening of human, animals and house hold farm workers and, decolonization measures to lower the risk of MRSA transmission.
Ethical Considerations
This study was approved by Institutional Review Board of Mbar¬ara University of science and technology (MUST) and Uganda National Council of Science and Technology (UNCST) study Number 13/08–15. All the protocols used in this study complied with the ethical standards of the committees on human experi¬mentation, and with the Helsinki Declaration of 1975 as revised in 2000.
Acknowledgments
The authors wish to thank the German academic exchange (DAAD) for financial support. We cannot forget to extend our sincere thanks to UNHLS management that provided reagents and space for this study. We also thank the staff of KRRH labo¬ratory that provided laboratory space and reagents.
Competiting Interest
We declare that there are no competing interests to this work.
Author Contributions
BA Conceived, designed the experiments, performed the exper¬iments and analyzed the data. BM: Contributed reagents/materi-als/analysis tools: BA, OP and JB: Wrote the paper.
Funding
We are grateful to the German academic exchange (DAAD) for funding this work.
Data Availability Statement
All data used in this work is available upon request.
References
- Shore, A. C., & Coleman, D. C. (2013). Staphylococcal cassette chromosome mec: recent advances and new insights. International Journal of Medical Microbiology, 303(6-7), 350-359.
- Andrew, B., Kagirita, A. A., Tonny, O., & Joel, B. (2018). High rates of methicillin-resistant Staphylococcus aureus colonization of domesticated swine of Kabale District–Southwestern Uganda.
- Peng, H., Liu, D., Ma, Y., & Gao, W. (2018). Comparison of community-and healthcare-associated methicillin-resistant Staphylococcus aureus isolates at a Chinese tertiary hospital, 2012–2017. Scientific reports, 8(1), 1-8.
- Graveland, H., Wagenaar, J. A., Bergs, K., Heesterbeek, H., & Heederik, D. (2011). Persistence of livestock associated MRSA CC398 in humans is dependent on intensity of animal contact. PloS one, 6(2), e16830.
- Choo, E. J. (2017). Community-associated methicillin-re-sistant Staphylococcus aureus in nosocomial infections. Infection & chemotherapy, 49(2), 158-159.
- David, M. Z., & Daum, R. S. (2010). Community-associ-ated methicillin-resistant Staphylococcus aureus: epidemiology and clinical consequences of an emerging epidemic. Clinical microbiology reviews, 23(3), 616-687.
- Kateete, D. P., Bwanga, F., Seni, J., Mayanja, R., Kigozi, E., Mujuni, B., ... & Joloba, M. L. (2019). CA-MRSA and HA-MRSA coexist in community and hospital settings in Uganda. Antimicrobial Resistance & Infection Control, 8(1), 1-9.
- P. H. England. (2015). “UK Standards for Microbiology Investigations,” Bacteriology, vol. B 55, no. 5.2, pp. 1–21.
- B. Limbago. (2001). “M100-S11, Performance standards for antimicrobial susceptibility testing,” Clin. Microbiol. Newsl., vol. 23, no. 6, p. 49.
- Ghaznavi-Rad, E., & Ekrami, A. (2018). Molecular characterization of methicillin-resistant Staphylococcus aureus isolates, isolated from a Burn Hospital in southwest Iran in 2006 and 2014. International journal of microbiology, 2018.
- Stegger, Á., Andersen, P. S., Kearns, A., Pichon, B., Holmes,M. A., Edwards, G., ... & Larsen, A. R. (2012). Rapid detection, differentiation and typing of methicillin-resistant Staphylococcus aureus harbouring either mecA or the new mecA homologue mecALGA251. Clinical Microbiology and Infection, 18(4), 395-400.
- Pfaller, M. A., & Castanheira, M. (2016). Nosocomial can-didiasis: antifungal stewardship and the importance of rapid diagnosis. Medical mycology, 54(1), 1-22.
- Fenner, L., Widmer, A. F., Dangel, M., & Frei, R. (2008). Distribution of spa types among meticillin-resistant Staphylococcus aureus isolates during a 6 year period at a low-prev-alence university hospital. Journal of medical microbiology, 57(5), 612-616.
- Strommenger, B., Kettlitz, C., Weniger, T., Harmsen, D., Friedrich, A. W., & Witte, W. (2006). Assignment of Staphylococcus isolates to groups by spa typing, SmaI macrore-striction analysis, and multilocus sequence typing. Journal of clinical microbiology, 44(7), 2533-2540.
- Hashemizadeh, Z., Bazargani, A., Kalantar-Neyestanaki, D., Mohebi, S., & Hadi, N. (2020). Determining spa-type of methicillin-resistant Staphylococcus aureus (MRSA) via high-resolution melting (HRM) analysis, Shiraz, Iran. BMC Research Notes, 13(1), 1-4.
- Fasihi, Y., Fooladi, S., Mohammadi, M. A., Emaneini, M., & Kalantar-Neyestanaki, D. (2017). The spa typing of methicillin-resistant Staphylococcus aureus isolates by high resolution melting (HRM) analysis. Journal of medical microbiology, 66(9), 1335-1337.
- Ruppitsch, W., Indra, A., StoÌ?ger, A., Mayer, B., Stadlbauer, S., Wewalka, G., & Allerberger, F. (2006). Classifying spa types in complexes improves interpretation of typing results for methicillin-resistant Staphylococcus aureus. Journal of clinical microbiology, 44(7), 2442-2448.
- Peters, C., Kleinmüller, O., Nienhaus, A., & Schablon, A. (2018). Prevalence and risk factors of MRSA colonisations: a cross-sectional study among personnel in outpatient care settings in Hamburg, Germany. BMJ open, 8(7), e021204.
- Moyo, S. J., Nkinda, L., Majigo, M., Mmbaga, E., Mbem-bati, N., Aboud, S., & Lyamuya, E. F. (2017). Prevalence of methicillin-resistant Staphylococcus aureus carriage on admission among patients attending regional hospitals in Dar es Salaam, Tanzania. BMC research notes, 10(1), 1-7.
- Gaddafi, M. S., Yakubu, Y., Junaidu, A. U., Bello, M. B., Garba, B., Bitrus, A. A., & Lawal, H. (2021). Nasal colonization of pigs and farm attendants by Staphylococcus aureus and methicillin-resistant Staphylococcus aureus (MRSA) in Kebbi, Northwestern Nigeria. The Thai Journal of Veterinary Medicine, 51(1), 119-124.
- Reynaga, E., Navarro, M., Vilamala, A., Roure, P., Quin-tana, M., Garcia-Nuñez, M., ... & Sabrià, M. (2016). Prevalence of colonization by methicillin-resistant Staphylococcus aureus ST398 in pigs and pig farm workers in an area of Catalonia, Spain. BMC infectious diseases, 16(1), 1-8.
- Mitevska, E., Wong, B., Surewaard, B. G., & Jenne, C. N. (2021). The prevalence, risk, and management of methicillin-resistant staphylococcus aureus infection in diverse populations across Canada: A systematic review. Pathogens, 10(4), 393.
- Cuny, C., Wieler, L. H., & Witte, W. (2015). Livestock-as-sociated MRSA: the impact on humans. Antibiotics, 4(4), 521-543.
- Salgueiro, V., Manageiro, V., Bandarra, N. M., Ferreira, E., Clemente, L., & Caniça, M. (2020). Genetic relatedness and diversity of Staphylococcus aureus from different reservoirs: humans and animals of livestock, poultry, zoo, and aquaculture. Microorganisms, 8(9), 1345.
- Cirkovic, I., Stepanovic, S., Skov, R., Trajkovic, J., Gr-gurevic, A., & Larsen, A. R. (2015). Carriage and genetic diversity of methicillin-resistant Staphylococcus aureus among patients and healthcare workers in a Serbian univer-sity hospital. PLoS One, 10(5), e0127347.
- Morschhäuser, J. (2016). The development of fluconazole resistance in Candida albicans–an example of microevolu-tion of a fungal pathogen. Journal of microbiology, 54(3), 192-201.
- Singh-Moodley, A., Lowe, M., Mogokotleng, R., & Perovic,O. (2020). Diversity of SCCmec elements and spa types in South African Staphylococcus aureus mecA-positive blood culture isolates. BMC Infectious Diseases, 20(1), 1-12.
- Asadollahi, P., Farahani, N. N., Mirzaii, M., Khoramrooz, S. S., Van Belkum, A., Asadollahi, K., ... & Darban-Sarokha-lil, D. (2018). Distribution of the most prevalent spa types among clinical isolates of methicillin-resistant and-suscep-tible Staphylococcus aureus around the world: a review. Frontiers in microbiology, 9, 163.
- Köck, R., Schaumburg, F., Mellmann, A., Köksal, M., Jurke, A., Becker, K., & Friedrich, A. W. (2013). Livestock-asso-ciated methicillin-resistant Staphylococcus aureus (MRSA) as causes of human infection and colonization in Germany. PloS one, 8(2), e55040.
- Bierowiec, K., PÅ?oneczka-Janeczko, K., & RypuÅ?a, K. (2016). Is the colonisation of Staphylococcus aureus in pets associated with their close contact with owners?. PLoS One, 11(5), e0156052.
- Kouyos, R., Klein, E., & Grenfell, B. (2013). Hospital-com-munity interactions foster coexistence between methicillin-resistant strains of Staphylococcus aureus. PLoS pathogens, 9(2), e1003134.
- Shrestha, A., Bhattarai, R. K., Luitel, H., Karki, S., & Bas-net, H. B. (2021). Prevalence of methicillin-resistant Staphylococcus aureus and pattern of antimicrobial resistance in mastitis milk of cattle in Chitwan, Nepal. BMC Veterinary Research, 17(1), 1-7.
- Kot, B., Wierzchowska, K., Piechota, M., & Grużewska, A. (2020). Antimicrobial resistance patterns in methicillin-re-sistant Staphylococcus aureus from patients hospitalized during 2015–2017 in hospitals in Poland. Medical Principles and Practice, 29(1), 61-68.
- Broens, E. M., Graat, E. A. M., Van der Wolf, P. J., Van de Giessen, A. W., Van Duijkeren, E., Wagenaar, J. A., .& De Jong, M. C. M. (2011). MRSA CC398 in the pig production chain. Preventive veterinary medicine, 98(2-3), 182-189.
- Wangai, F. K., Masika, M. M., Maritim, M. C., & Seaton,R. A. (2019). Methicillin-resistant Staphylococcus aureus (MRSA) in East Africa: red alert or red herring?. BMC infectious diseases, 19(1), 1-10.
- Mourabit, N., Arakrak, A., Bakkali, M., Zian, Z., Bakkach, J., & Laglaoui, A. (2020). Nasal carriage of Staphylococcus aureus in farm animals and breeders in North of Morocco. BMC infectious diseases, 20(1), 1-6.
- Dweba, C. C., Zishiri, O. T., & El Zowalaty, M. E. (2018). Methicillin-resistant Staphylococcus aureus: livestock-as-sociated, antimicrobial, and heavy metal resistance. Infection and Drug Resistance, 11, 2497.


