Research Article - (2026) Volume 7, Issue 1
Convolutional Neural Networks with Fuzzy-Based Modelling: A Framework for Disease Detection in Cocoa Crops
Received Date: Jan 06, 2026 / Accepted Date: Feb 25, 2026 / Published Date: Mar 20, 2026
Copyright: ©2026 Abiodun Muyideen Mustapha, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Adekoye, O., Mustapha, A. M., Okorie, D. (2026). Convolutional Neural Networks with Fuzzy-Based Modelling: A Framework for Disease Detection in Cocoa Crops. Adv Mach Lear Art Inte, 7(1), 01-19.
Abstract
Context: The increasing need for accurate and interpretable computer vision systems in agriculture has driven research into hybrid intelligent models. Cocoa production in Nigeria suffers significant yield losses due to diseases such as Black Pod and Cocoa Swollen Shoot Virus Disease (CSSVD), while manual inspection remains slow, subjective, and inefficient. Integrating fuzzy logic with convolutional neural networks (CNNs) offers a pathway to address diagnostic uncertainty and environmental variability.
Objectives: This study aims to develop and evaluate a Fuzzy-based Convolutional Neural Network (CNN–Fuzzy) framework for the automated detection and classification of major cocoa diseases. The goal is to enhance diagnostic accuracy, improve handling of uncertain field conditions, and provide a practical decision-support tool for farmers.
Methods: A hybrid architecture combining CNN-based image feature extraction with fuzzy inference rules was designed. A dataset of 12,000 annotated cocoa leaf and pod images sourced from the Cocoa Research Institute of Nigeria and open repositories (e.g., Kaggle) was used to train and validate the model. Comparative experiments were conducted against traditional machine learning classifiers and transfer learning-based models. The final framework was deployed as a mobile application optimized for offline use to support field-based disease diagnosis.
Results: The proposed CNN–Fuzzy model achieved a classification accuracy of 99.99%, surpassing traditional machine learning models (75.48–80.34%) and transfer learning approaches (up to 97.27%). Field-oriented deployment demonstrated its capability to identify diseases including Black Pod and CSSVD in real time and provide context-specific remedy recommendations.
Conclusion: The integration of fuzzy reasoning with deep learning significantly enhances the reliability and interpretability of cocoa disease diagnosis. The developed system offers a scalable, low-cost precision agriculture tool capable of reducing crop losses by an estimated 30–50%. This research advances digital transformation in cocoa production and lays a foundation for sustainable, AI-driven agricultural innovation in developing economies.
Keywords
Cocoa Disease Detection, Convolutional Neural Networks (CNN), Fuzzy Logic Modelling, Precision Agriculture
Introduction
Agriculture remains the cornerstone of food security and economic stability across developing nations [1]. In Nigeria, the sector contributes approximately 24% to the Gross Domestic Product (GDP) and employs over one-third of the nation’s workforce, particularly within rural communities where it serves as a key driver of poverty reduction and economic diversification [2,3]. Cocoa holds a central position as the fourth-largest globally produced crop among Nigeria’s agricultural commodities, with an annual output of about 328,263 tons [4].
Beyond its contribution to livelihoods, cocoa also serves as one of Nigeria’s top non-oil export earners and a vital component in the nation’s diversification agenda [5,6]. However, cocoa production continues to face severe threats from crop diseases such as Black Pod Disease and Cocoa Swollen Shoot Virus Disease, which cause extensive yield losses and economic setbacks for smallholder farmers [7]. Traditional approaches to disease identification rely heavily on manual inspection by agricultural extension officers [8-10]. This approach is time-consuming, labor-intensive, and prone to misdiagnosis, especially across vast and dispersed farmlands; leading to delayed diagnoses, misidentification, and ineffective interventions that cause irreversible crop damage and economic losses [11]. The rapid growth in global food demand has emphasized the need for technological advancement in agricultural production. Integrating digital and intelligent systems into farming practices such as disease detection has become essential for improving productivity, sustainability, and decision-making. In this regard, artificial intelligence (AI) and its subset, machine learning, have shown significant promise in automating crop monitoring and disease detection [12]. Deep learning models, particularly, Convolutional Neural Networks (CNNs) have proven highly effective for image-based classification tasks, including the recognition of subtle visual patterns in crop leaves and pods [13,14]. Yet, their effectiveness can be constrained by noise, environmental variability, inconsistent image quality, and the inherent uncertainty in real-world agricultural data [15]. To overcome these challenges, researchers are increasingly exploring hybrid approaches that blend the computational strength of deep learning with reasoning systems capable of handling uncertainty [16]. Fuzzy logic, which allows for approximate reasoning similar to human decision-making, has emerged as an effective complement to CNN-based image analysis [17]. This convergence of methods offers a promising pathway toward developing intelligent, adaptable systems for crop disease detection that can operate reliably under diverse field conditions and resource constraints.
Machine Learning and Agriculture
Machine Learning (ML), a core subset of Artificial Intelligence (AI), has become an essential enabler of modern precision agriculture [18]. It allows computational systems to learn from data, identify patterns, and make autonomous predictions thereby minimizing reliance on manual observation and expertise. In recent years, machine learning techniques have transformed crop management by improving yield forecasting, soil-health analysis, and plant-disease detection through automated image recognition [19]. Traditional farming methods including cocoa cultivation, rely heavily on visual inspection by farmers or extension agents. This approach, while practical, is time-consuming and prone to human error, especially under conditions of large farm size, labor shortages, and disease symptom overlap [20]. Machine Learning based solutions address these gaps by using computer-vision models to analyze leaf and pod imagery captured via smartphones, drones, or field sensors [21]. Such systems have shown remarkable success in early disease detection and yield improvement [22]. Convolutional Neural Networks (CNNs) are particularly well-suited for agricultural imaging because of their capacity to learn spatial hierarchies from raw pixels [23]. Studies have demonstrated that CNN-based models are capable of detecting cocoa diseases like Black Pod and Swollen Shoot Virus with accuracies above 90% by learning patterns like lesion shapes [14,24,25]. They detect Black Pod from pod textures, adapting well to variable lighting. Integration with hyperspectral data enhances feature extraction for subtle spectral differences in infected tissues [26]. Beyond classification, recent work explores lightweight CNN architectures (e.g., MobileNet, EfficientNet) that can operate efficiently on low-cost mobile devices, making the technology accessible to smallholder farmers [27]. To further enhance reliability in uncontrolled farm environments, researchers have incorporated fuzzy-logic reasoning into machine learning frameworks. Fuzzy systems interpret the uncertainty and partial truths inherent in agricultural data such as varying disease severity or inconsistent lighting conditions by mapping outputs into linguistic categories such as mild, moderate, severe rather than absolute decisions [28]. Hence, this study integrates fuzzy-based modeling into Convolutional Neural Networks to enhance reliability in disease detection and support adaptive interpretation of cocoa disease imagery.
Motivation
Global demand for cocoa is projected to grow steadily over the coming decade, driven by population growth and the expanding food and confectionery industries [29,30]. As the fourth-largest cocoa producer, Nigeria plays a vital role in meeting this demand, contributing significantly to global supply and rural livelihoods [31]. However, crop diseases such as Black Pod and Cocoa Swollen Shoot Virus Disease (CSSVD) threaten up to 30–90% of annual yields [32], posing a major risk to both local farmers and international production stability. Increasing production efficiency is essential to sustain supply and meet projected global demand. Yet, factors such as disease outbreaks, limited extension services, and unpredictable environmental conditions continue to hinder productivity [9,10,33]. Machine learning (ML) and computer vision present transformative solutions by enabling early, automated disease detection and intervention. This research is motivated by the urgent need to enhance cocoa health management through technological innovation. For farmers, it promises improved crop health, reduced yield losses, and higher incomes through timely and precise disease diagnosis [34]. It also supports Nigeria’s goal of increasing cocoa output, diversifying foreign exchange earnings, and enhancing global market competitiveness [35]. By integrating fuzzy-based modeling into Convolutional Neural Networks (CNNs), the study seeks to overcome the uncertainty and variability that limit conventional systems, providing a robust framework to safeguard cocoa yields, strengthen farmer livelihoods, and contribute to global food security.
Research Contribution
This study introduces a fuzzy-enhanced Convolutional Neural Network framework for the detection and classification of major cocoa crop diseases. Unlike conventional CNN architectures that rely solely on fixed pooling or deterministic activation, the proposed model integrates fuzzy inference mechanisms to manage uncertainty, noise, and environmental variability inherent in field-acquired image data. The proposed approach leverages fuzzy inference to interpret ambiguous image features, such as overlapping symptoms or varying disease severity under different lighting conditions. The framework focuses on detecting and classifying two major cocoa diseases: Black Pod Disease (Phytophthora palmivora) and Cocoa Swollen Shoot Virus Disease (CSSVD), while its architecture is extendable to other prevalent infections such as Leaf Anthracnose (Colletotrichum sp.) and Frosty Pod Rot (Moniliophthora roreri). It uses a dataset of approximately 12,000 annotated cocoa images sourced from the Cocoa Research Institute of Nigeria (CRIN) and open-access repositories. To ensure accessibility and real-world applicability, the system is deployed as a mobile application optimized for offline use, offering farmers real-time disease identification and actionable remedy recommendations. The model achieves 99.99% accuracy, outperforming both traditional machine learning approaches (75.48–80.34%) and transfer learning baselines (up to 97.27%), while remaining computationally efficient for low-resource devices.
Paper Organization
The paper is structured to present the complete development and evaluation of the proposed model for cocoa disease detection. The introduction highlights the economic significance of cocoa in Nigeria and the challenges caused by major diseases such as Black Pod and Cocoa Swollen Shoot Virus Disease. It establishes the motivation for developing an intelligent, uncertainty-aware model capable of improving disease identification accuracy under real-world agricultural conditions. The related works section reviews existing approaches in plant disease detection using conventional machine learning and deep learning models. It outlines their strengths and limitations. The methodology section details the dataset composition, preprocessing techniques, and the design of the proposed CNN and fuzzy architecture. It describes how fuzzy logic principles are integrated with convolutional operations to enhance feature extraction, minimize information loss, and improve classification accuracy. The implementation explains the system configuration, training parameters, and evaluation metrics used to validate model performance. The results and discussion section present the performance analysis of the model, including accuracy, precision, recall, and F1-score, alongside confusion matrices and visual comparisons. The proposed model’s performance is compared with baseline CNNs and previous research, demonstrating its superior classification capability. Finally, the conclusion and future scope summarize the research findings, theoretical and practical contributions, and potential future directions such as expanding the model to other crop diseases, integrating multi-modal data, and deploying the system in real-world agricultural environments.
Literature Survey
In recent years, the integration of deep learning and computer vision in agricultural research has yielded significant progress, particularly in the domain of automated plant disease detection. Convolutional Neural Networks (CNNs) have demonstrated remarkable potential in learning complex visual patterns from plant images, enabling high accuracy in classification tasks. However, challenges such as class imbalance, visual ambiguity, and environmental noise continue to affect model generalization and field-level reliability. Researchers have explored several approaches to enhance CNN architectures, ranging from lightweight mobile-based implementations to hybrid fuzzy-logic systems, in an effort to improve diagnostic precision, scalability, and adaptability across diverse agricultural contexts. An EfficientDet-Lite4 model for detecting and localizing moniliasis and black pod rot on cocoa pods was proposed by [36]. The lightweight model was trained on a curated dataset of healthy and diseased pods and integrated into a native Android application. This implementation highlighted the growing emphasis on deployable, mobile-friendly AI systems for agricultural use. While the study emphasized model efficiency and user accessibility, it was limited in scope to pod-based diseases, overlooking leaf and stem pathologies that significantly affect cocoa yield. The research nevertheless advanced the notion that lightweight CNN architectures can serve as scalable foundations for farmer oriented diagnostic tools.
Soh et al. (2024) expanded on this trend by evaluating multiple CNN architectures including Custom CNN, VGG-16, EfficientNetB0, ResNet50, and LeNet-5 for general cocoa disease classification [12]. Their Custom CNN achieved an accuracy of 91.79%, demonstrating the robustness of tailored architectures for domain specific image data. The study also emphasized the importance of the F1-score in evaluating performance, noting potential class imbalance issues common in agricultural datasets. Although accuracy was strong, the moderate F1-score of 82.08% suggested inconsistencies in detecting minority disease classes, indicating the need for improved data augmentation and class weighting strategies to ensure balanced model generalization [24]. Kumi et al. (2024) developed Cocoa Companion, a deep learning-based smartphone application designed to diagnose two major cocoa diseases: Cocoa Swollen Shoot Virus Disease and Black Pod Disease [25]. The system utilized four CNN architectures, among which SSD MobileNet V2 achieved the highest classification accuracy. The model enabled farmers to capture field images with their smartphones and receive immediate disease diagnoses via cloud-based processing. This approach addressed a pressing challenge in cocoa farming which is limited access to agricultural extension officers by enabling rapid, on-site detection [25]. While practical and accessible, the system’s dependency on constant internet connectivity limited its usability in rural areas. The study thus underscored the need for more resilient, offline-capable diagnostic frameworks tailored to low resource environments [37].
Beyond cocoa-specific studies, broader agricultural research has also explored the application of deep learning for plant disease identification. Mohanty et al. conducted one of the earliest large-scale experiments using CNNs to detect 14 diseases across 12 plant species using over 50,000 images from the Plant Village dataset [14]. Their model achieved 99.35% accuracy, proving the effectiveness of deep learning for plant disease detection. However, as their dataset was collected in controlled environments with plain backgrounds and uniform lighting, the model’s real-world generalizability remained limited. This highlighted the need for datasets reflective of natural farm conditions which is an issue particularly relevant for crops like cocoa grown in tropical, variable environments.
Other recent studies have introduced fuzzy logic into CNN frameworks to enhance interpretability and resilience under uncertainty. Similarly, Vijaya and Aruna developed a fuzzy rule-based CNN system for medical image classification, achieving higher accuracy and robustness against image noise [38]. Yet, this method's reliance on extensive labeled medical imaging data and increased model complexity due to fuzzy rules may hinder its real-time applicability in clinical settings. In agricultural contexts, Mohammed et al. integrated Mamdani fuzzy inference systems with CNNs to analyze animal images, showcasing scalability and improved handling of imprecise visual features [39]. A limitation of this hybrid approach may be its dependency on carefully designed fuzzy rules and the challenge of dealing with highly variable real-world image conditions common in agriculture.
In the plant health domain, a Type-2 Fuzzy Logic System (T2FLS) was implemented to monitor plant health using the well-known PlantVillage dataset. Their approach leveraged fuzzy logic’s ability to handle inherent environmental uncertainty and variable lighting in agricultural imagery [40]. Specifically, the system used fuzzy decision rules derived from expert knowledge to classify the severity of maize common rust disease based on calculated percentages of diseased leaf area, converting RGB color images to grayscale for efficient segmentation. Testing under varied real-world illumination conditions confirmed the model’s robustness, achieving high classification accuracy (e.g., 89% testing accuracy) by combining fuzzy logic with a fine-tuned deep learning network (VGG-16). Nagi and Tripathy further advanced this theme by integrating fuzzy feature extraction with a Probabilistic Neural Network (PNN) for leaf disease classification. Their hybrid model utilized fuzzy feature descriptors to better capture uncertainty and vagueness in visual symptoms from leaf images, improving discrimination between disease classes [41]. The method achieved superior classification results on several leaf disease datasets, outperforming standard neural networks by effectively handling variability in natural image features often caused by environmental factors, disease progression, or image acquisition variations. These studies emphasize a growing research interest in combining fuzzy logic and deep learning to address the intrinsic ambiguity and noise in plant disease image data, improving interpretability, robustness, and accuracy in realistic agricultural settings.
In precision agriculture and fruit classification, fuzzy pooling techniques help preserve nuanced visual information crucial for distinguishingsubtledifferencesinimagesaffectedbyenvironmental variability. Although direct studies on fruit classification using fuzzy pooling remain limited, these pooling methods are noted for their capability to improve robustness and accuracy in complex, noisy datasets, which is highly relevant in agricultural imagery [42]. Sharma et al. introduced a Type-1 fuzzy pooling layer within CNN architectures to address uncertainty and noise in feature maps during dimensionality reduction [43]. Unlike traditional max or average pooling, fuzzy pooling incorporates fuzzy logic principles through fuzzification, aggregation, and defuzzification steps, allowing for better preservation and representation of critical image details. Their experiments demonstrated that CNNs enhanced with fuzzy pooling consistently outperform those using conventional pooling methods on standard image classification datasets such as CIFAR-10, MNIST, and Fashion-MNIST. This approach also advances model interpretability by managing uncertainty explicitly within the CNN layers.
Al-Gaashani et al. proposed SANet, a self-attention based deep learning architecture specifically designed for rice disease classification. SANet combines depthwise separable convolutions with residual skip connections to achieve computational efficiency and high accuracy (up to 98.71%) [44]. The integration of attention mechanisms enables the model to focus on disease-relevant regions in rice leaf images, thereby enhancing classification precision. This architecture exemplifies recent advances in agricultural deep learning models that balance effective feature extraction and practical deployment considerations like model size and inference speed.
In a parallel approach, Tyagi et al. implemented and compared three convolutional neural network (CNN) architectures: Inception V3, VGG-16, and VGG-19 for the identification and disease detection of medicinal plant leaves [45]. The study highlighted how Inception V3’s flexible design efficiently handled diverse input sizes with minimal pre-processing, while VGG-16 and VGG-19 offered strong feature extraction capabilities suited for complex image textures. Each network was fine-tuned on large-scale image datasets to distinguish between healthy and diseased samples. The comparative results demonstrated that all three CNNs achieved high classification performance, with VGG-19 exhibiting superior accuracy and robustness in extracting intricate visual patterns. The research emphasized the effectiveness of deep learning architectures in automating plant disease diagnosis and underscored their potential for sustainable medicinal plant management through precise, scalable image classification systems. T. Kim et al. also integrated a fuzzy inference system to better handle ambiguous symptom presentation and uncertainty in leaf images caused by lighting and occlusion [46]. This framework improved classification confidence and interpretability, achieving comparable accuracy to standard CNN models while enhancing robustness in challenging natural image conditions.
In a comprehensive study, researchers [47] evaluated multiple convolutional neural network architectures such as SqueezeNet, AlexNet, DarkNet-19, and a modified CNN on a dataset of 1,200 cocoa leaf images labeled for Vascular Streak Dieback (VSD) infection. The dataset was split 80:20 for training and testing, with 960 training and 240 testing images. They applied image preprocessing steps including RGB loading, leaf segmentation, and data augmentation techniques such as white balance color enhancement to simulate environmental lighting variability, aiming to enhance model robustness. Among the models tested, the modified DarkNet-19 achieved the highest testing accuracy of 98.61%, demonstrating superior performance in detecting subtle disease symptoms on cocoa leaves. A hybrid CNN approach incorporating fuzzy logic in the decision layer was introduced which enabled better differentiation between similar disease symptoms on cocoa leaves [48]. Trained on an extensive dataset from multiple West African cocoa farms, their model demonstrated improved generalizability and reduced false positives, aiding farmers in timely, accurate disease management.
Other relevant contributions include Kalpana et al. who developed the Deep Learning-Automated Plant Disease Detection and Classification (DLAPDDC) algorithm focused on computational efficiency by using separable depthwise convolutions [49]. This design choice allowed the model to significantly reduce the number of parameters and computational complexity, making the approach suitable for deployment on resource-constrained edge devices often used in remote agricultural environments. The algorithm was tested on multiple crop diseases datasets, demonstrating a good balance between accuracy and inference speed, making it practical for real-time disease detection in the field.
Mamadou et al. also proposed a MobileNet-based system tailored specifically for cocoa pod disease detection [50]. Leveraging MobileNet's lightweight architecture optimized for mobile platforms, their system emphasized low-latency inference, critical for timely disease identification in field scenarios. The model achieved competitive accuracy in classifying common cocoa pod diseases, while its compact size allowed deployment on smartphones or low-cost devices used directly by farmers for on-the-spot diagnosis. While MobileNet is optimized for low-latency, resource-constrained devices, the tradeoff for compactness can sometimes be a reduction in model capacity, potentially affecting classification accuracy on more complex or varied disease symptoms. Misclassifications, especially between visually similar disease stages or pod conditions, have been reported at a moderate rate, suggesting further training data diversity and hybrid model approaches may improve reliability under field variability [51]. Lopes et al. contributed foundational research exploring deep computer vision systems for cocoa classification focused broadly on cocoa health and quality assessment. Their work developed convolutional neural network models capable of differentiating between quality grades and visual traits of cocoa pods [52]. While comprehensive in scope, specific disease classification details were limited, serving more as a baseline for automated visual analysis of cocoa pod conditions rather than targeted disease detection.
Collectively, these studies reveal a consistent trajectory toward leveraging deep learning and fuzzy logic for more adaptive, accurate, and deployable agricultural diagnostic systems. While remarkable progress has been achieved, a research gap persists in the integration of fuzzy-based modeling within CNN architectures specifically for cocoa disease detection. Current models either focus narrowly on specific disease categories or depend heavily on online infrastructure, limiting their accessibility to smallholder farmers. This study aims to bridge these gaps by proposing a fuzzy-based CNN framework capable of robust disease detection and classification under uncertain field conditions, advancing both theoretical understanding and practical application in precision agriculture. Here in Table: 1, the key studies relevant to this work are presented, summarizing their core methodologies, reported results, and limitations. These studies collectively underscore the potential of Convolutional Neural Networks (CNNs) and hybrid models to automate diagnosis, reduce reliance on manual inspection, and improve crop health outcomes. Within cocoa farming, several researchers have proposed models and mobile-based applications that leverage computer vision for disease identification, though challenges such as limited datasets, class imbalance, and poor field generalizability persist.
|
Author(s)/ Year |
Title |
Methodology |
Results |
Limitations |
|
Mohanty et al. |
Using Deep Learning |
Deep CNN trained on |
High accuracy (99.35%) |
Dataset collected under |
|
(2016) [14] |
for Image-Based Plant |
54,306 leaf images |
reported on controlled |
controlled conditions; |
|
|
Disease Detection. |
across 14 crop species |
images. |
field generalizability |
|
|
|
and 26 diseases. |
|
limited. |
|
Soh et al. (2024) |
Cocoa Diseases |
Evaluated multiple |
Best model identified; |
Class imbalance affected |
|
[24] |
Classification Using |
CNN architectures for |
insight into model selection |
F1 scores, specially for |
|
|
Deep Learning |
disease classification on |
for disease classification. |
less common diseases. |
|
|
Algorithm. |
cocoa leaves. |
|
|
|
Kumi et al. |
Smartphone app for |
Deep Learning- Based |
Developed a CNN-based |
Requires internet; |
|
(2022) [25] |
CSSVD and Black Pod |
Smartphone Application |
mobile app for detecting |
accuracy >80% but can |
|
|
disease diagnosis. |
for Cocoa Disease |
CSSVD and Black Pod |
be improved. |
|
|
|
|
disease using image |
|
|
|
|
|
classification. |
|
|
Vera et al. (2024) |
Deep Learning- Based |
Used EfficientDet |
Fast, on-device processing |
Limited disease types |
|
[36] |
Computation al Model |
-Lite4 object detection |
suited for farmers. |
covered; limited |
|
|
for Disease Identification |
for identifying |
|
performance details |
|
|
in Cocoa Pods. |
“monilia” and “black |
|
provided. |
|
|
|
pod” diseases on pods. |
|
|
|
Al-Gaashani et al. (2023) [44] |
Using a Resnet50 with a Kernel Attention Mechanism for Rice Disease |
Selfattention CNN model with depthwise convolutions and residual connections. |
High accuracy (98.71%) achieved; computationall y efficient. |
Focused on rice; requires adaptation and validation for cocoa diseases. |
|
Mamadou et al. (2023) [50] |
Cocoa Pods Diseases Detection by MobileNet Confluence and Classification Algorithms. |
Lightweight MobileNet CNN model optimized for mobile devices. |
Low latency; practical realtime field use; competitive accuracy. |
Dataset diversity limited; MobileNet capacity constraints may reduce accuracy on complex cases. |
|
Lopes et al. (2022) [52] |
Deep Computer Vision System for Cocoa Classification. |
Hybrid CNN model for cocoa bean quality and type classification. |
Demonstrated high accuracy and effective quality assessment. |
General cocoa classification only; limited disease specific application. |
|
Siddappa (2018) [53] |
Artificial Intelligence Based Drone Controlled Detection of Plant Disease. |
Conceptual design of AI-enabled drones for scalable monitoring of cocoa disease in plantations. |
Scalable solution; early detection aid for large farms. |
Concept paper only; high drone costs and practical deployment challenges. |
Table 1: Literature Survey
Methodology
Dataset and Preprocessing
The dataset employed in this study was compiled from two primary sources: The Cocoa Research Institute of Nigeria (CRIN) and publicly available Kaggle repositories related to cocoa disease detection. In total, the dataset comprised approximately 12,000 images capturing both healthy and infected cocoa leaves and pods. The collection reflects the natural diversity of field conditions found in major cocoa-producing regions of Nigeria with variations in lighting, background, disease stage, and pod maturity. The images were categorized into three classes corresponding to key cocoa health states: Black Pod Disease (Phytophthora palmivora),Cocoa Swollen Shoot Virus Disease (CSSVD) and Healthy Cocoa samples. Each category contained a balanced mix of leaf and pod images, ensuring adequate representation across different disease manifestations. Figure 1 illustrates representative samples from the dataset, demonstrating the visual variability the model was trained to handle.
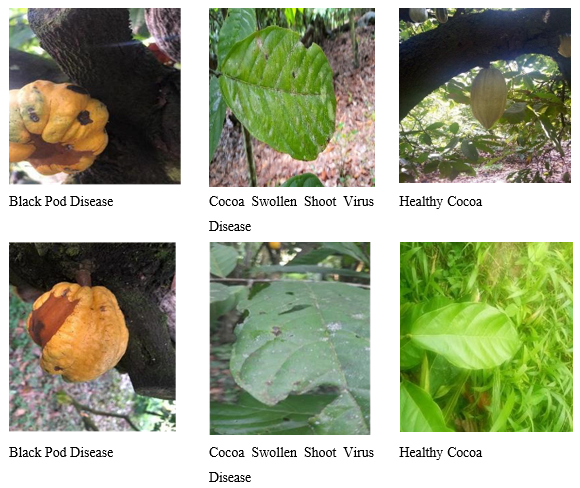


Figure 1: Sample Dataset Images
To enhance subtle symptom detection, a hyperspectral-inspired preprocessing step was employed to simulate spectral variations and emphasize disease-related features not readily visible in standard RGB channels. All images were standardized to 224 × 224 pixels and retained in RGB color mode to preserve chromatic details essential for classification. Maintaining the original directory labels, the dataset was partitioned into 80% training (9,600 images), 10% validation (1,200 images), and 10% (1,200 images) testing sets to enable rigorous evaluation of model generalization under real-world cocoa field conditions. During training, on-the-fly data augmentation was applied to increase robustness against field variability; augmentations included random rotations, horizontal/vertical flips, zoom operations, and brightness adjustments. Bilinear interpolation was used during resizing to preserve image quality. Images were automatically labeled based on their directory placement using TensorFlow/ Keras preprocessing utilities, batched in groups of 32, and shuffled at the start of each epoch to minimize sequence bias.
Model Architecture
The convolutional pathway of the proposed framework forms the foundation for feature extraction and hierarchical representation of cocoa disease patterns. Building upon established deep learning architectures, the network was designed to automatically learn discriminative visual features from cocoa pod and leaf images, capturing the subtle variations that differentiate healthy and infected specimens. Each input image, standardized to 224 × 224 × 3 dimensions during preprocessing, enters the convolutional neural network (CNN) where successive convolutional and pooling operations progressively extract and condense spatial information. The initial convolutional layers employ small 3 × 3 kernels to capture local texture patterns such as color gradients and lesion boundaries associated with early-stage infections. As depth increases, the network identifies more complex features such as irregular pod surface textures, blackened patches, and shoot distortions which are critical for distinguishing Phytophthora palmivora (Black Pod Disease) and Cocoa Swollen Shoot Virus Disease (CSSVD). Rectified Linear Unit (ReLU) activation functions were applied after each convolutional layer to introduce non-linearity and accelerate convergence. Maxpooling layers with a 2 × 2 filter size were used to downsample feature maps, reducing dimensionality while retaining essential spatial cues. Batch normalization was introduced after convolutional operations to stabilize learning and prevent internal covariate shifts, ensuring smoother gradient flow across the network. To further enhance generalization, a dropout rate of 0.2 was employed in the fully connected layers. This regularization step reduces overfitting, which is especially important given the high number of epochs (100) used during training. The model was trained using a batch size of 32 and a learning rate of 0.0001, optimized with the Adam optimizer for efficient adaptive learning. These hyperparameters were determined through systematic tuning, balancing convergence speed with classification accuracy. Following the convolutional and pooling stages, the extracted feature maps were flattened into one-dimensional vectors and passed through dense layers responsible for high-level reasoning and pattern discrimination. These layers integrate the learned features across spatial hierarchies, preparing them for the subsequent fuzzy-based modeling stage.
Figure 2: Convolutional Neural Network Architecture
Mathematical Representation of the CNN Model
The convolutional neural network (CNN) designed in this study serves as the foundational component of the proposed fuzzy-based framework for cocoa disease detection. Its main objective is to extract spatial and color-based patterns from cocoa pod and leaf images that correspond to distinct disease symptoms such as dark lesions, discolorations, and pod swelling. CNNs are highly effective in such agricultural imaging tasks because they automatically learn hierarchical feature representations directly from raw image data, eliminating the need for manual feature engineering [14].
• Convolution Layer
The convolution layer is the core of the network and performs localized filtering to extract relevant visual features from input cocoa images. For each image X, a set of kernels K is applied to produce feature maps Y as follows:
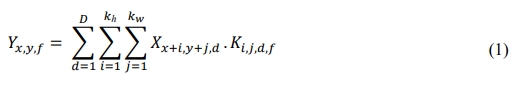
Where: Yxyf is the resulting feature activation at location x,y for filter f; kh and kw are kernel dimensions (3 × 3 in this study); d indexes the input channels;
Ki i,j,d,f represents the learnable weight of each kernel. Through this operation, the network progressively captures disease-related structures from low-level patterns like vein distortion to high-level characteristics such as black pod lesions or color anomalies associated with Cocoa Swollen Shoot Virus
f(x) = max(0, x) (2)
ReLU selectively activates positive feature responses while suppressing negative ones [54], improving training convergence and ensuring that subtle yet relevant features such as faint pod discoloration or uneven leaf tone are retained for further processing.
• Pooling Layer
Pooling layers are used to down-sample feature maps, reducing dimensionality while retaining significant spatial information. hierarchical feature representations directly from raw image data, eliminating the need for manual feature engineering [14].
• Convolution Layer
The convolution layer is the core of the network and performs localized filtering to extract relevant visual features from input cocoa images. For each image X, a set of kernels K is applied to produce feature maps Y as follows:
Disease (CSSVD).
• Activation Mapping
After convolution, a nonlinear transformation is applied to introduce nonlinearity into the model and ensure that the network can learn complex decision boundaries. The Rectified Linear Unit (ReLU) activation function was adopted for this purpose due to its computational simplicity and robustness to the vanishing gradient problem. The activation is defined as:
Two pooling techniques: MaxPooling and AveragePooling, were employed at different stages of the network to balance noise suppression and feature retention.
• MaxPooling
MaxPooling extracts the most prominent features by selecting the maximum value within a local window of size p X P. The operation is defined as:
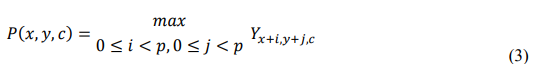
Where P(x,y,c) represents the pooled output for channel c. This process preserves key activation responses that signify distinct disease areas, such as intense black lesions or swollen pod ridges, while discarding irrelevant background variations.
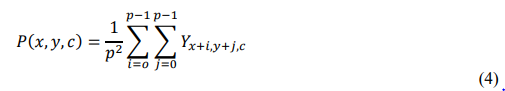
This operation aids in capturing average intensity variations and general texture information that might indicate mild disease symptoms. The combination of MaxPooling and AveragePooling enhances both sensitivity to strong disease cues and adaptability to subtler early-stage infections [55].
• Dropout Layer
To prevent overfitting and improve model generalization, a dropout regularization strategy was implemented within the fully connected layers of the proposed CNN. During each training iteration, a fraction of neurons was randomly deactivated to ensure that the model did not become overly reliant on specific nodes or
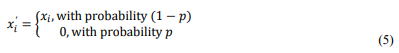
where p=0.2 represents the dropout probability, xi is the original neuron output, andXi' is the modified activation after dropout.
• Model Optimization
Model training was performed using the Adam optimizer, which combines the advantages of adaptive learning rate adjustment and momentum-based convergence. The optimizer's efficiency
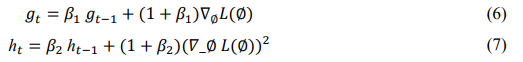
After bias correction, the parameters are updated according to:

where ![]() = 0.0001 is the learning rate, € prevents division by zero, and ß1, ß2 are exponential decay rates. Adamâ??s adaptive mechanism ensures smooth convergence and resilience to the varying intensity of cocoa disease patterns.
= 0.0001 is the learning rate, € prevents division by zero, and ß1, ß2 are exponential decay rates. Adamâ??s adaptive mechanism ensures smooth convergence and resilience to the varying intensity of cocoa disease patterns.
• AveragePooling
AveragePooling, in contrast, computes the mean of values within the pooling window, yielding a smoother and more generalized feature map:
local patterns. A dropout rate of 0.2 was adopted, meaning that 20% of neurons were temporarily omitted in each forward pass. This moderate rate provided a balance between learning capacity and robustness thereby allowing the network to capture fine-grained cocoa disease features while still maintaining sufficient regularization. The inclusion of dropout reduced coadaptation among neurons and enhanced the model's ability to generalize across diverse field images characterized by lighting variation and partial occlusion. Mathematically, the dropout operation can be described as:in handling sparse gradients and nonstationary data made it particularly suitable for the heterogeneous cocoa image dataset [56]. A learning rate of 0.0001 was employed, allowing for stable gradient updates and gradual convergence across 100 epochs. The batch size was set to 32, striking a balance between memory efficiency and gradient stability. The optimizer adjusts learning rates dynamically based on gradient history, defined as:â?¢ Loss Function
• Loss Function
The network was trained using categorical cross-entropy loss, appropriate for multi-class classification across the three categories: Black Pod, CSSVD, and Healthy. It is expressed as:

Where K= 3, Zj is the true class label, and zj is the model’s predicted probability for class j. This formulation penalizes incorrect predictions and ensures the model assigns higher confidence to the correct disease class, thereby maximizing diagnostic accuracy.
Fuzzy-Based Modeling for Cocoa Disease Detection
Conceptual Overview
This section introduces the fuzzy-based reasoning layer designed to complement the Convolutional Neural Network (CNN) model for cocoa disease classification. The CNN component extracts discriminative spatial and textural features from cocoa leaf and pod images, while the fuzzy inference mechanism enables reasoning under uncertainty; something purely numerical models often fail to handle effectively. In real farm environments, symptoms such as pod darkening, leaf mottling, or stem swelling may appear gradually or overlap between Black Pod Disease (Phytophthora palmivora) and Cocoa Swollen Shoot Virus Disease (CSSVD). The fuzzy system bridges this ambiguity gap by translating CNNderived probabilities into interpretable linguistic descriptors such as Low, Moderate, or Severe infection, thereby offering a more intuitive assessment of disease state.
Proposed Fuzzy-Based Framework
The fuzzy logic system processes numerical indicators derived from observed cocoa plant attributes such as lesion area, color intensity, and pod surface texture and transforms them into qualitative assessments of disease severity. The reasoning process unfolds in three sequential phases, corresponding to the blocks in Figure 3.
Figure 3: Fuzzy-Based Modelling Framework
• Input Variables
The fuzzy model receives crisp numerical features derived from the CNN and image-level analysis. These inputs reflect measurable visual indicators of plant health:
• Input 1: color intensity or hue deviation (degree of discoloration),
• Input 2: lesion or infected-area ratio,
• Input m: pod-surface texture irregularity or roughness.
Each input captures a distinct aspect of disease manifestation and collectively represents the observable symptom space of cocoa crops.
• Fuzzification Stage
In this stage, the crisp numerical inputs are transformed into fuzzy linguistic variables using triangular membership functions. It internally defines graded levels of severity such as Low, Medium, and High.
Mathematically, a fuzzy set A is represented by its membership function:
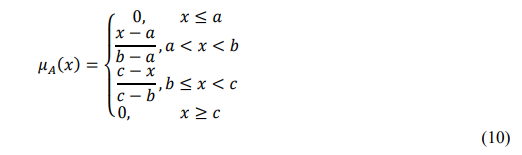
where a,b,c specifies the lower, peak, and upper bounds of the triangular function.
This overlapping structure ensures smooth transitions—e.g., a lesion ratio of 0.6 may partially belong to both Medium and High, reflecting uncertainty in symptom intensity.
• Rule Evaluation
IF (Color Deviation is High) AND (Lesion Area is Medium) THEN (Sev = Severe). (11)
The system defines 15 fuzzy rules, derived through iterative hyperparameter tuning.
![]()
Aggregated inference combines all active rules using the maximum operator:
![]()
This inference mechanism allows multiple uncertain observations such as overlapping pod discoloration and leaf mottling to be integrated into one coherent severity assessment.
• Defuzzification
The final stage converts the fuzzy output distribution into a single
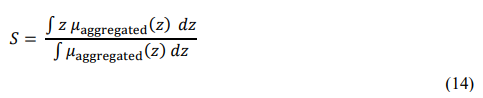
This value is then mapped into one of three interpretive categories: Low, Moderate, or High severity; corresponding to recommended farmer actions. For example:
• Low severity: continue observation,
• Moderate severity: initiate preventive spraying,
• High severity: isolate infected pods and applies treatment.
These outputs are later integrated into the mobile application to guide farmers through localized decision support.
Significance of the Fuzzy Component
The inclusion of fuzzy reasoning enhances the CNN’s deterministic predictions by introducing a human-interpretable reasoning layer.
Once inputs are fuzzified, they are passed through a rule base that encodes expert agricultural knowledge of cocoa disease behavior.
Each rule defines how combinations of fuzzy inputs correspond to disease severity.
A general rule can be expressed as:
The degree of activation for each rule (![]() ) is calculated using the minimum (AND) operator:
) is calculated using the minimum (AND) operator:
crisp value that represents the predicted disease severity score.
Using the centroid (center-of-gravity) method, the crisp output S is computed as:
This enables handling of uncertain and overlapping symptoms, reduction of misclassification caused by lighting or background noise, improved interpretability through linguistic explanations of model output, and greater field applicability under uncontrolled conditions typical of Nigerian cocoa farms.
Materials and Methods
Proposed Fuzzy-Based Architecture
The proposed model was implemented as a sequential framework combining Convolutional Neural Networks (CNNs) and fuzzy-based reasoning for the classification of cocoa diseases. The network processes input images of cocoa pods or leaves resized to 224 × 224 × 3 pixels, corresponding to the height, width, and RGB color channels. This resolution ensures consistent dimensionality while preserving the fine textural information required for disease identification. The CNN backbone consists of three convolutional layers with 3 × 3 kernels and ReLU activation after each layer to introduce non-linearity and accelerate convergence. The first convolutional block applies 32 filters with “same” padding to maintain the original spatial dimensions, followed by a MaxPooling (2 × 2) operation that reduces feature map size while retaining dominant features such as color irregularities and lesion contours. The second and third convolutional layers employ 64 and 128 filters, respectively, progressively learning higher-order features that correspond to varying degrees of disease severity or infection spread. A dropout rate of 0.5 is introduced after the third convolutional block to reduce overfitting and improve generalization across different field lighting and texture variations. The resulting feature maps are flattened into a one-dimensional vector and passed into a fully connected dense layer, where the extracted patterns are consolidated. A final Softmax activation generates probability distributions for each class: Healthy, Black Pod Disease, and Cocoa Swollen Shoot Virus. During compilation, the model employs the Adam optimizer with a learning rate of 0.0001 and categorical cross-entropy as the loss function, reflecting its multiclass classification objective. Training is performed for 100 epochs with a batch size of 32, and model performance is monitored using accuracy, precision, recall, and F1-score metrics. The dataset is divided into 80 % training, 10 % validation, and 10 % testing subsets to assess the model’s generalization capability and prevent overfitting. The system was executed on a workstation with 32 GB RAM, utilizing TensorFlow and Keras frameworks for model training and performance evaluation. Figure 4 shows the overall CNN–Fuzzy system architecture used in this study.
Figure 4: Proposed CNN–Fuzzy System Architecture
Fuzzy–CNN Integration Mechanism
The CNN provides a strong foundation for feature extraction, yet its deterministic nature can limit performance under uncertain field conditions. To address this, fuzzy logic is employed as a
![]()
Where PCNN represents the CNN’s softmax class probabilities, and Fauxincludes auxiliary input features such as lesion ratio, color deviation, or pod-surface irregularity. The fuzzy function ffuzzy (. ) applies three processes: fuzzification, rule evaluation, and defuzzification; to compute a crisp severity score S. For instance, if the CNN predicts CSSVD with a confidence of 0.78 and the lesion ratio is high, the fuzzy logic system may yield a High Severity score (≈0.82) after applying its rules and centroid defuzzification. This adaptive reasoning allows the hybrid model to better handle ambiguous or early-stage symptoms compared to traditional CNNs
refinement layer that interprets CNN outputs as degrees of belief rather than absolute classifications. Mathematically, the fusion is expressed as:
Model Training and Evaluation
The training pipeline was organized into iterative epochs wherethe model learned feature hierarchies and fuzzy logic relationships concurrently. Input data were shuffled between epochs to ensure randomization and avoid bias. Model accuracy was monitored on the validation set, with early stopping criteria based on loss stabilization. The performance of the model was measured using four key evaluation metrics: Accuracy, Precision, Recall, and F1-Score; defined as follows:
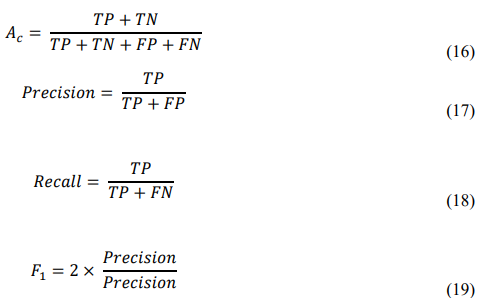

Where TP, FP and FN denote true positives, false positives, and false negatives, respectively.
A confusion matrix was also constructed to visualize how well the model distinguished between different cocoa health states. The proposed fuzzy enhanced CNN achieved strong generalization with stable validation accuracy across epochs, demonstrating improved robustness compared to baseline CNN models.
Implementation and Deployment Setup
Training of the integrated CNN–Fuzzy model was performed using Python with TensorFlow and Keras libraries, both known for their high performance in deep learning workflows. The training utilized a cloud-based Kaggle GPU accelerator (32 GB RAM) for faster convergence, while preprocessing and initial testing were conducted on a Mac OS workstation powered by an Intel Core i7 (3.0 GHz) CPU and 32 GB system RAM. The integrated CNN–Fuzzy model was deployed through an Android-based mobile application, providing real-time, on-device cocoa disease diagnosis. The application was built using TensorFlow Lite for the CNN model conversion and a lightweight fuzzy logic engine for post-inference reasoning. The mobile interface accepts captured or uploaded images, processes them locally, and outputs disease classification along with severity interpretation and recommended treatment guidance. The system operates fully offline, requiring no internet connection for inference, while periodic cloud synchronization updates the model and rule base for performance enhancement. This implementation design ensures accessibility for farmers in rural regions and supports multi-environment adaptability without relying on external computation.
Results and Discussion
Model Performance Evaluation
The performance of the proposed Fuzzy—CNN model was evaluated across training, validation, and testing phases. The results are summarized in Table 2, showing consistent performance across all data splits with a testing accuracy of 0.99. The model demonstrated minimal deviation between training and validation accuracy, confirming its ability to generalize effectively to unseen data. The high accuracy (0.99 on testing) indicates precise classification of cocoa diseases, with fuzzy logic effectively managing uncertainties in symptom presentation. Precision and recall values confirm minimal false positives/negatives, crucial for practical farming where misdiagnosis leads to unnecessary treatments or overlooked outbreaks.
|
Metric |
Training Set |
Validation Set |
Testing Set |
|
Accuracy |
1.00 |
1.00 |
0.99 |
|
Precision |
0.99 |
0.98 |
0.96 |
|
Recall |
0.98 |
0.98 |
0.97 |
|
F1-Score |
0.99 |
0.98 |
0.97 |
Table 2: Performance Analysis of the CNN-Fuzzy Model across Different Data Splits
The training and validation curves for accuracy and loss are illustrated in Figures 5 and 6, respectively. These curves reveal stable learning dynamics over 100 epochs, with the model's performance evolution over the training period, demonstrating stable learning curves and effective optimization.
Figure 5: Training and Validation Accuracy Curves for CNN–Fuzzy Model.
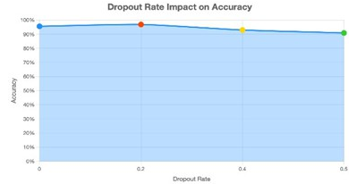
Figure 6: Loss Curves of the Proposed CNN–Fuzzy Model.
The accuracy curves demonstrate that the model achieves steady improvement, reaching near-perfect performance (1.0) for both training and validation sets. This indicates effective learning from cocoa images without significant overfitting, as evidenced by the close alignment between training and validation curves. Figure 6 shows consistently decreasing loss values for both training and validation sets, validating the model's optimization effectiveness and its ability to generalize to unseen cocoa data. The smooth convergence without oscillations indicates stable learning dynamics.
Hyperparameter Sensitivity and Optimization
To ensure optimal model performance, hyperparameter tuning experiments were conducted to analyze how dropout rate and learning rate affected accuracy. Figure 7 illustrates that a dropout rate of 0.2 yields the most balanced result (~96.9 % accuracy), effectively preventing overfitting while retaining feature richness.
Figure 7: Dropout Rate vs. Model Accuracy
Similarly, the learning-rate analysis in Figure 8 shows that a rate of 0.0001 achieved stable and efficient convergence. Higher values caused oscillations, while smaller ones slowed optimization.
Confusion Matrix Analysis
The matrix shows strong diagonal dominance, with the majority of samples correctly classified. Minor off-diagonal values (0.30—0.34) indicate occasional misclassification between Black Pod and CSSVD, which often share similar discoloration features in early infection stages. Despite these overlaps, the overall recognition accuracy remains consistently high, demonstrating the fuzzy layer's strength in handling uncertain visual inputs.
Figure 9: Confusion Matrix for CNN–Fuzzy Model Classification Results.
Comparative Performance Analysis
The performance of the proposed CNN–Fuzzy model was systematically compared with several baseline machine learning algorithms and pre-trained transfer learning architectures to validate its superior diagnostic capability. Unlike traditional approaches, which struggle to interpret the subtle and often overlapping visual features of cocoa disease imagery, the integration of fuzzy logic within the CNN architecture enabled more precise and context-aware classification.
• Comparison Between CNN–Fuzzy and Transfer Learning Models
Table 3 presents the comparative performance of state-of-the-art transfer learning models fine-tuned on the same cocoa dataset. These included MobileNet, VGG19, ResNet, and EfficientNet B4. The proposed CNN–Fuzzy model substantially outperformed all competing architectures, achieving 99.99% accuracy, surpassing EfficientNet B4 (92.67%) and ResNet (97.27%). This remarkable gain underscores the strength of the fuzzy integration in managing uncertainties and enhancing decision confidence.
|
Models |
Accuracy |
Precision |
Recall |
F1 score |
|
MobileNet |
0.89 |
0.81 |
0.83 |
0.82 |
|
Visual Geometry Group 19 |
0.92 |
0.82 |
0.83 |
0.82 |
|
Residual Network |
0.97 |
0.80 |
0.83 |
0.82 |
|
EfficientNet B4 |
0.92 |
0.87 |
0.89 |
0.88 |
|
CNN–Fuzzy Model (Proposed) |
0.99 |
0.96 |
0.97 |
0.97 |
• Comparison with Previously Published Research Works
A further evaluation was conducted against published studies on plant disease classification seen on Table: 4. This comparison highlights the unique contributions and advantages of the fuzzy-logic approach.
|
Reference |
Classifiers |
Dataset |
Accuracy |
|
Mohanty et al. (2016) [14] |
Deep CNN |
Plant Village |
99.35% |
|
Sachdeva et al. (2021) [57] |
Optimized CNN |
Plant Village |
98.9% |
|
Zeng et al. (2021) [58] |
MobileNetV2 |
Proprietary Dataset |
99.55% |
|
Devi and Amarendra (2021) [59] |
Hist Gradient Boo sting |
Plant Village |
89.11% |
|
Kaushik et al. (2022) [60] |
CNN |
Plant Village |
94.60% |
|
This study |
FuzzyConvolutional Neural Networks |
Cocoa Images, Kaggle images |
99.99% |
Table 4: Comparative Analysis with Previously Published Research Works
Ablation Study
An ablation study was conducted to assess the influence of key model components and design decisions on the overall performance of the proposed CNN–Fuzzy network. The goal was to understand how different configurations affect feature preservation, computational cost, and model stability during training. The analysis revealed that integrating the fuzzy reasoning layer markedly improved the systemâ??s capability to interpret uncertain or partially overlapping disease patterns. By assigning graded membership values to input features rather than binary classifications, the fuzzy component enhanced the model's sensitivity to subtle pod color shifts and irregular lesion textures that are easily overlooked by standard convolutional layers.
Conclusion and Future Scope
The present study developed a hybrid CNN–Fuzzy model model for automatic detection and classification of cocoa diseases, addressing a critical challenge in Nigeria’s agricultural sector [7]. By integrating fuzzy logic into the CNN framework, the model effectively captured both the structural and uncertain visual cues in cocoa pod and leaf images. The approach achieved remarkable accuracy of 99.99%, with precision, recall, and F1-scores all exceeding 96%, demonstrating superior performance over traditional machine learning methods and conventional CNN architectures. It delivers a functional, low-cost digital diagnostic tool capable of reducing yield losses and improving farmers’ decision-making. Future research can incorporate multi-modal inputs (e.g., weather, soil, and spectral data), and exploring federated learning for privacy-preserving model updates. Field validation studies across diverse agroecological zones will further test scalability and impact. Also, integration with IoT devices and blockchain enabled agricultural systems could strengthen transparency and trust in data-driven farming.
Data Accessibility Statement
The datasets used and analyzed in this study were obtained from the Cocoa Research Institute of Nigeria (CRIN) archives and supplemented with publicly available cocoa disease images from Kaggle. Due to institutional restrictions, the CRIN dataset is available upon reasonable request from the corresponding author, while the Kaggle dataset can be accessed freely at: https://www.kaggle.com/datasets/ohagwucollinspatrick/amini-cocoacontamination-dataset.
Authors’ Contributions
Author 1: Conceptualization, methodology development, data collection, implementation of the CNN–Fuzzy model.
Author 2: Supervision, editing, and review of the final manuscript.
Author 3: Data validation, dataset preparation, performance analysis, results interpretation, drafting and revision of the manuscript.
References
- Pawlak, K., & KoÅ?odziejczak, M. (2020). The role of agriculture in ensuring food security in developing countries: Considerations in the context of the problem of sustainable food production. Sustainability, 12(13), 5488.
- Micheal, V. A. (2024). Impact of Agricultural Variables on Gross Domestic Product (GDP) in Nigeria: A Sarima Approach. African Journal of Agricultural Science and Food Research, 14(1), 78-91.
- VICTORIA, O., & BENEDETTE, O. (2025). Impact of the Agricultural Sector on Job Creation in Nigeria. INTERNATIONAL JOURNAL OF RESEARCH, 10, 224-238.
- Obaniyi, K. S., & Oladele, O. E. (2023). Participation of cocoa farmers in farmers field school and its effect on yield in Osun State, Nigeria.
- Bature, Y. M. T., Aliegba, E. T., & Caleb, R. L. (2024). An assessment of value-added agricultural export and economic development in Nigeria (2015–2022). Global Journal of Research in Agriculture & Life Sciences, 4(5).
- Bature, Y. M. T., Aliegba, E. T., & Caleb, R. L. C. (2024). Agricultural exports, value addition and economic development: The Nigerian experience, 2015–2023. Zenodo.
- Dongo, L. N., & Orisajo, S. B. (2007). Status of cocoa swollen shoot virus disease in Nigeria. African Journal of Biotechnology, 6(17).
- Aniagyei, J., Bakang, J. E. A., Tham-Agyekum, E. K., Arhin, M., & Asiedu, P. (2024). Employing agricultural extension delivery services for improving cocoa bean quality. Cogent Social Sciences, 10(1), 2333431.
- Aremu, T., & Reynolds, T. W. (2024). Welfare benefits associated with access to agricultural extension services in Nigeria: Welfare benefits of agricultural extension in Nigeria. Food Security, 16(2), 295-320.
- Owolabi, A. O., & Yekinni, O. T. (2022). Utilisation of information and communication technologies for agricultural extension service delivery in public and non-public organisations in southwestern Nigeria. Heliyon, 8(9).
- Kongor, J. E., Owusu, M., & Oduro-Yeboah, C. (2024). Cocoa production in the 2020s: challenges and solutions. CABI Agriculture and Bioscience, 5(1), 102.
- Aijaz, N., Lan, H., Raza, T., Yaqub, M., Iqbal, R., & Pathan,M. S. (2025). Artificial intelligence in agriculture: Advancing crop productivity and sustainability. Journal of Agriculture and Food Research, 20, 101762.
- Ashurov, A. Y., Al-Gaashani, M. S., Samee, N. A., Alkanhel, R., Atteia, G., Abdallah, H. A., & Saleh Ali Muthanna, M. (2025). Enhancing plant disease detection through deep learning: a Depthwise CNN with squeeze and excitation integration and residual skip connections. Frontiers in Plant Science, 15, 1505857.
- Mohanty, S. P., Hughes, D. P., & Salathé, M. (2016). Using deep learning for image-based plant disease detection. Frontiers in plant science, 7, 1419.
- Peng, M., Liu, Y., Qadri, I. A., Bhatti, U. A., Ahmed, B., Sarhan, N. M., & Awwad, E. M. (2024). Advanced image segmentation for precision agriculture using CNN-GAT fusion and fuzzy C-means clustering. Computers and Electronics in Agriculture, 226, 109431.
- Yadav, P., & Singh, P. (2025). Intelligent leaf disease diagnosis: Fuzzy logic and CNN-based multi-class detection. Communications on Applied Nonlinear Analysis, 32(8s), 933.
- Yerkin, A., Igali, A., Kadyrgali, E., Shagyrov, M., Ziyada, M., Muratbekova, M., & Shamoi, P. (2025). Fuzzy theory in computer vision: a review. arXiv preprint arXiv:2507.18660.
- Pauzi, N. A. M., Mustaza, S. M., Zainal, N., Zaman, M. H. M., & Moubark, A. M. (2025). Artificial Intelligence in precision agriculture: A review. Jurnal Kejuruteraan, 37(2), 1025-1047.
- Dhanya, V. G., Subeesh, A., Kushwaha, N. L., Vishwakarma,D. K., Kumar, T. N., Ritika, G., & Singh, A. N. (2022). Deep learning based computer vision approaches for smart agricultural applications. Artificial Intelligence in Agriculture, 6, 211-229.
- Sambana, B., Nnadi, H. S., Wajid, M. A., Fidelia, N. O., Camacho-Zuñiga, C., Ajuzie, H. D., & Onyema, E. M. (2025). An efficient plant disease detection using transfer learning approach. Scientific Reports, 15(1), 19082.
- Chin, R., Catal, C., & Kassahun, A. (2023). Plant disease detection using drones in precision agriculture. Precision Agriculture, 24(5), 1663-1682.
- Paul, N., Sunil, G. C., Horvath, D., & Sun, X. (2025). Deep learning for plant stress detection: A comprehensive review of technologies, challenges, and future directions. Computers and Electronics in Agriculture, 229, 109734.
- Okorie, D. D., Fatokun, J. O., & Umoren, U. M. (2024). Applying Convolutional Neural Networks for Enhanced Digital Image Steganalysis. In Practical Statistical Learning and Data Science Methods: Case Studies from LISA 2020 Global Network, USA (pp. 697-718). Cham: Springer Nature Switzerland.
- Soh, K. S., Moung, E. G., Danker, K. J. J., Dargham, J. A.,& Farzamnia, A. (2024). Cocoa diseases classification using deep learning algorithm. In ITM Web of Conferences (Vol. 63,p. 01014). EDP Sciences.
- Kumi, S., Kelly, D., Woodstuff, J., Lomotey, R. K., Orji, R., & Deters, R. (2022). Cocoa companion: Deep learning-based smartphone application for cocoa disease detection. Procedia Computer Science, 203, 87-94.
- Shafay, M., Hassan, T., Owais, M., Hussain, I., Khawaja, S. G., Seneviratne, L., & Werghi, N. (2025). Recent advances in plant disease detection: challenges and opportunities. Plant Methods, 21(1), 140.
- Aboh, M. (2025). Benchmarking EfficientNet, MobileNet and ResNet Architectures for Cassava Disease Detection in Agricultural Robotics. Journal of Engineering Research and Reports, 27(10), 380-387.
- Nagi, R., & Tripathy, S. S. (2020). Application of fuzzy logic in plant disease management. In Fuzzy expert systems and applications in agricultural diagnosis (pp. 261-302). IGI Global.
- Gavrilova, N. G. (2021, September). Contemporary global production and consumption of cocoa: An assessment. In IOP Conference Series: Earth and Environmental Science (Vol. 839, No. 2, p. 022095). IOP Publishing.
- V. Voora, S. Bermúdez, and C. Larrea. (2019). Western Europe and developing economies in Asia are driving demand for cocoa but farm risks may affect supply in the long-term.
- Agulanna, F. T., Ogunlade, M. O., Oluyole, K. A., & Nduka, B.(2025). Forest Trees and their Perceived Negative Impacts on Cocoa Plantations in Southern Nigeria. Moor Journal of Agricultural Research, 26(1), 73-85.
- Evans, H. C. (2007). Cacao diseases—the trilogy revisited.Phytopathology, 97(12), 1640-1643.
- Issa, F. O. (2013). Building the capacity of agricultural extension personnel for effective implementation of agricultural transformation agenda in Nigeria. Journal of Agricultural Extension, 17(1), 78-88.
- Alvarado, J., Restrepo-Arias, J. F., Velásquez, D., & Maiza, M. (2025). Disease detection on cocoa crops based on computer-vision techniques: a systematic literature review. Agriculture, 15(10), 1032.
- Oluwaseun, F. M., Bolarinwa, E. J., & Nnanna, O. C. (2024). Smart Farming: AI Technology for Farm Monitoring and Security in Nigeria. International Journal of Advances in Engineering and Management, 771-781.
- Vera, D. B., Oviedo, B., Casanova, W. C., & Zambrano-Vega,C. (2024). Deep learning-based computational model for disease identification in cocoa pods (Theobroma cacao L.). arXiv preprint arXiv:2401.01247.
- Madiwal, A. S., Jha, R. B., & Barthakur, P. (2025). Edge AI and ioT for real-time crop disease detection: A survey of trends, architectures, and challenges. International Journal of Research and Innovation in Applied Science, 10(7), 1011-1027.
- Vijaya, M., & Aruna, A. (2022). CNN-Fuzzy Rule Based System for Classifying Medical Imaging Process. Journal of Algebraic Statistics, 13(2).
- Mohammed, H. R., & Hussain, Z. M. (2021). Hybrid mamdani fuzzy rules and convolutional neural networks for analysis and identification of animal images. Computation, 9(3), 35.
- Awotunde, J. B., Ajamu, G. J., Abdulraheem, M., Adeniyi,E., Jimoh, R. G., Oladipo, I. D., ... & ADENIYI, J. K.(2023, April). Plant disease diagnosis and detection using type-2 fuzzy logic system. In 2023 International Conference on Science, Engineering and Business for Sustainable Development Goals (SEB-SDG) (Vol. 1, pp. 1-11). IEEE.
- Nagi, R., & Tripathy, S. S. (2023). Plant disease identification using fuzzy feature extraction and PNN. Signal, Image and Video Processing, 17(6), 2809-2815.
- Sultana, S., Tasir, M. A. M., Nobel, S. N., Kabir, M. M., & Mridha, M. F. (2024). XAI-FruitNet: An explainable deep model for accurate fruit classification. Journal of Agriculture and Food Research, 18, 101474.
- Sharma, T., Singh, V., Sudhakaran, S., & Verma, N. K. (2019, June). Fuzzy based pooling in convolutional neural network for image classification. In 2019 IEEE international conference on fuzzy systems (FUZZ-IEEE) (pp. 1-6). IEEE.
- Al-Gaashani, M. S., Samee, N. A., Alnashwan, R., Khayyat, M., & Muthanna, M. S. A. (2023). Using a Resnet50 with a kernel attention mechanism for rice disease diagnosis. Life, 13(6), 1277.
- Tyagi, K., Vats, S., & Vashisht, V. (2024). Implementing Inception v3, VGG-16 and VGG-19 architectures of CNN for medicinal plant leaves identification and disease detection. J Electr Syst, 20(7s), 2380-2388.
- Kim, T. H., Shahroz, M., Alabdullah, B., Innab, N., Baili, J., Umer, M., ... & Ashraf, I. (2024). ANFIS Fuzzy convolutional neural network model for leaf disease detection. Frontiers in Plant Science, 15, 1465960.
- Ridlo, Z. R., & Agustin, I. H. (2023, May). Application of convolutional neural network for identifying cocoa leaf disease. In Proceedings of the 1st International Conference on Neural Networks and Machine Learning 2022 (ICONNSMAL 2022) (p. 283). Springer Nature.
- Dumont, A., Ribeyre, F., Babin, R., Hérault, B., Kouadio, J., Koffi, A. D. K., ... & Guéry, L. (2025). Associations between scale-dependent agroecosystem factors and cocoa swollen shoot virus incidence in Côte d’Ivoire. Agriculture, Ecosystems & Environment, 393, 109851.
- Pavithra, A., Kalpana, G., & Vigneswaran, T. (2024). RETRACTED ARTICLE: Deep learning-based automated disease detection and classification model for precision agriculture. Soft Computing, 28(Suppl 2), 463-463.
- Mamadou, D., Kacoutchy, J. A., Ballo, A. B., & Kouassi,B. M. (2023). Cocoa pods diseases detection by MobileNet confluence and classification algorithms. International Journal of Advanced Computer Science and Applications, 14(9).
- GLENN, M. G. (2025). Efficient detection of cacao pod diseases using SSD MobileNetV2 FPN-Lite. WORLD, 25(2), 1099-1105.
- Lopes, J. F., da Costa, V. G. T., Barbin, D. F., Cruz-Tirado, L.J. P., Baeten, V., & Barbon Junior, S. (2022). Deep computer vision system for cocoa classification. Multimedia Tools and Applications, 81(28), 41059-41077.
- Siddappa, P. (2018). Artificial intelligence based drone controlled detection of plant disease. Int. J. Creative Res. Thoughts (IJCRT), 6(2), 1-4.
- Layton, O. W., Peng, S., & Steinmetz, S. T. (2024). ReLU, sparseness, and the encoding of optic flow in neural networks. Sensors, 24(23), 7453.
- Zafar, A., Aamir, M., Mohd Nawi, N., Arshad, A., Riaz, S., Alruban, A., ... & Almotairi, S. (2022). A comparison of pooling methods for convolutional neural networks. Applied Sciences, 12(17), 8643.
- Zhang, Z. (2018, June). Improved adam optimizer for deep neural networks. In 2018 IEEE/ACM 26th international symposium on quality of service (IWQoS) (pp. 1-2). Ieee.
- Sachdeva, G., Singh, P., & Kaur, P. (2021). Plant leaf disease classification using deep Convolutional neural network with Bayesian learning. Materials Today: Proceedings, 45, 5584-5590.
- Zeng, Y., Zhao, Y., Yu, Y., Tang, Y., & Tang, Y. (2021, June). Pepper disease detection model based on convolutional neural network and transfer learning. In IOP Conference Series: Earth and Environmental Science (Vol. 792, No. 1, p. 012001). IOP Publishing.
- Devi, M. B., & Amarendra, K. (2021). Machine learning-based application to detect pepper leaf diseases using histgradientboosting classifier with fused HOG and LBP features. In Computer Networks, Big Data and IoT: Proceedings of ICCBI 2020 (pp. 969-979). Singapore: Springer Singapore.
- Tulshan, A. S., & Raul, N. (2019, July). Plant leaf disease detection using machine learning. In 2019 10th International Conference on Computing, Communication and Networking Technologies (ICCCNT) (pp. 1-6). IEEE.