Research Article - (2023) Volume 1, Issue 1
Comparison of Antimicrobial Susceptibility of Escherichia Coli Isolated from Fecal Poultry and Bovine Housed in Tunisian Farms; Phylogroup Diversity and Detection of Tetracycline and Sulfonamides Resistant Genes With Integron Class1
2Faculty of Medicine, LR99ES09 Laboratory of Antibiotic Resistance, University of Tunis El Manar, Tun, Tunisia
3Department of Microbiology, Hospital of Charles Nicolle, Tunis, Tunisia
4Emergency Center for Transboundary Animal Diseases (FAOSNE), Tunisia
Received Date: Apr 15, 2023 / Accepted Date: May 20, 2023 / Published Date: Jun 23, 2023
Copyright: ©Â©2023 Kilani Hajer, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Hajer, K., Mohamed Salah, A., Rim, D., Bouraoui, J., Riadh, M., et al. (2023). Comparison of Antimicrobial Suscepti- bility of Escherichia Coli Isolated from Fecal Poultry and Bovine Housed in Tunisian Farms; Phylogroup Diversity and Detection of Tetracycline and Sulfonamides Resistant Genes With Integron Class1. Transl Med OA, 1(1), 17-26.
Abstract
Little detailed documentation researched the excessive use of antimicrobials such as tetracycline and sulfonamides in vet- erinary medicine in Tunisia and more studies are needed. A total of 58 of commensal Escherichia coli isolates recovered from fecal samples of healthy poultry (n=31) and bovin (n=27) recovered from farms in Tunisia were examinated for 20 antimicrobial as well as, researched the presence of integron, variable regions (VRs), phylogroupes, tetracycline (tetA, tetB et tetC) and sulfonamides(sul1, sul2, sul3) resistance genes. The most frequently resistance in poultry origin were to tetracycline (94.3%), sulfonamide (70.69%), nalidixic acid (61.29), amoxicillin (58%), to trimthetoprim-sulfamethoxazole and streptomy- cin with the same rate (64.51%), ticarcilline (58%). Whereas, the bovine isolates were most resistant to streptomycin (55.5%), to amoxicillin (18.5), to tetracycline (37%), and have a moderate same rates to kanamycine, to trimthetoprim-sulfamethoxaz- ole 11.11, to nalidixic acid and to sulfonamide 7.4%, For poultry and bovine class 1 integron were detected in 20, 6 isolates, respectively as well as class 2 integron were found in 2 and 1 isolates, respectively. Class 1 integrons were significantly asso- ciated with poultry origin (p=0.001). For poultry sul1, sul2, and sul3 genes were detected in 14 (46.2 %), 7 (23.8 %), and 4 (8.9 %) resistant isolates, respectively. Whereas, for bovine 5 isolates were resistant to sulfonamide and sul1 and sul2 genes were detected in 4 and 1 isolates with absence of sul3 genes. and tetracycline genes tetA, tetB genes were observed in 27 (84.37 %) and 8 (25%) resistant isolates, respectively while, TetC was not detected amongst our isolates. Seven arrangement gene cassette were detected; dfrA1-satA1-aadA1 in one identical DNA fragments with approximate size of 2000 bp and six arrangements of resistance gene cassettes of class 1 integron were detected; dfrA1+aadA1 (5isolates); for dfrA17+aadA1, dfrA12+ orfF+aadA2 each one two 2isolates; one isolate for aadA1 and dfrA5, respectively. In poultry, 16 isolates were found to belong to phylogroup A (sub- groupA1: 12, sub- groupA0: 4); 9 to B1, 1 to B2 and 5 to phylogroup D. However, in bovine 9 isolates have the phylogroups A1, 7 isolates B1, 4 isolates B2, and 3 isolates found to phylogroup D. Our results showed that the prevalence of resistance in E. coli isolates from poultry was much higher than that in bovin.
Keywords
Escherichia Coli, Antimicrobial Resistance, Integrons, Poultry, Bovin.
Introduction
Escherichia coli can cause a variety of diarrheal and other ex- tra-intestinal infections in humans and animals. The emergence of E. coli isolates with multiple antibiotic resistance phenotypes, has been previously reported and is considered as a serious health concern [1,2]. In Enterobacteriaceae and particularly in E. coli, resistance to beta-lactams due to Extended Spectrum Beta-Lactamases (ESBL), quinolones, and aminoglycosides have drawn considerable attention worldwide [3]. ESBL-pro- ducing isolates are usually resistant to other antibiotics such as aminosides, tetracycline, chloramphenicol, trimethoprim, sulfonamides, or quinolones, often due to the presence of dif- ferent resistance genes on transferable elements such as plas- mids, transposons, or integrons [4-8]. In the last decade, it has been observed that ESBL-producers and multi-drug resistant E. coli isolates are frequently detected in food-producing animals or food products, and thus health authorities are worried about the potential transmission of these resistant microorganisms to humans through the food chain. Likewise, in Tunisia many re- ports have highlighted a high prevalence of antibiotic resistance and ESBL production in E. coli isolates from food-producing animals or food products, a situation that requires more investi- gations and vigilance to monitoring large dissemination of such resistant isolates [9-12].
Mobile genetic elements such as plasmids, transposons, and in- tegrons are able to disseminate genes encoding antibiotic resis- tance by horizontal transfer, and play important role in the evolu- tion and dissemination of multidrug resistance in Gram-negative bacteria [13,14]. Five classes of integrons related to antibiotic resistance have been described based on the homology of their integrase genes; however, class 1 and 2 were the most prevalent integrons in Enterobacteriaceae [15-17]. Class 1 integron com- prises two conserved segments, the 5’conserved segment (5’CS) and the 3’ conserved segment (3’CS), and an internal variable region. Variable regions are able to contain many gene cassettes and up to 6 cassettes have been reported, which explain, in part, the multiresistance trait of some reported isolates [18].
The aim of this study was to investigate the presence of integrons in commensal E. coli isolates from animal origins (poultry and bovine) housed in farms located in four different governorates of Tunisia (Sousse, Tunis, Bizerte and Gbeli), with a focus on antimicrobial resistance of the isolates and phylogroups as well as genetic characterization of genes encoding sulphonamides, tetracycline- resistance (owing to their high rates of resistance in our collection and their excessive use in veterinary medicine in Tunisia).
Materials and Methods
Bacterial Strains and Epidemiological Background
Fifty-eight fecal E. coli isolates were collected from poultry (31 isolates, 4 farms (two farms in Sousse in the Central-East re- gion of Tunisia, farm in borej cédri, farm in Tunis) and bovine (27 isolates, 2 farms; farm in Bizerte ( in the north of Tunisia) and farm in Gbeli (the southern of Tunisia) during 2012. All animals were healthy and in on each farm, fecal samples were obtained from different flocks that contained from 1000 to 1500 and immediately after collection were cultivated on Mac-Conk- ey agar and incubated overnight at 37°C. One colony with typi- cal E. coli trait from each sample was picked and re-isolated on Mac-Conkey agar, and followed by biochemical identification using Api20E (Bio-Mérieux, France).
Antimicrobial Susceptibility Testing
Antimicrobial susceptibility testing was carried out by the agar disk diffusion method on Mueller–Hinton agar plates according to recommendation of Clinical and Laboratory Standards Insti- tute guidelines (CLSI, 2012). The antibiotics tested were the fol- lowing: amoxicillin (25 μg), amoxicillin + clavulanic acid (20 μg + 10 μg), ticarcillin (75 μg), ticarcillin + clavulanic acid (75 μg + 10 μg), piperacillin (75 μg), cefoxitin (30 μg), cefotaxime (30 μg), ceftazidime (30 μg), imipenem (10 μg), streptomycin (10 UI), gentamicin (10 UI), kanamycin (30 UI), tobramycin (10 μg), tetracycline (30 UI), nalidixic acid (30 μg), ofloxacin (5 μg), pefloxacin (5 μg), ciprofloxacin (5 μg), sulfonamide (200 µg), trimethoprim/sulfamethoxazole (1,25 μg + 23,75 μg) (Bio- Rad). E. coli ATCC 25922 was used as a control strain. For the screening of ESBL production The Double-Disc Synergy Test (DDST) with cefotaxime or ceftazidime in the proximity to amoxicillin-clavulanic acid was used [19].
Detection and Characterization of Integron Class 1 and 2
For integrons of class 1 and 2, the 3’CS of class 1 integrons, and the variable regions of integrons class 1 and 2 were amplified by PCR as previously reported and randomly 16 chosen poultry and bovine isolates of class 1 and 2 integrons were sequenced [20].
Detection of the Genes Encoding Tetracycline and Sul- fonamide Resistance
The presence of the genes encoding tetracycline (tetA, tetB, and tetC) and sulfonamides-resistance (sul1, sul2, and sul3) was in- vestigated by PCR for all resistant isolates to these markers as reported by Saénz et al., [20].
Phylogenetic Analysis of E.coli Isolates
E. coli phylogenetic grouping was determined by a triplex PCR assay as reported by Clermont et al, (2000). The combination of PCR products obtained allowed classification of E. coli strains into one of the four major E. coli phylogenetic lineages: A, B1, B2 or D. Appropriate positive and negative controls were includ- ed in the assay.
Statistical Analysis
Statistical testing was done using Epi Info (version 6.04) soft- wares to check if there were a significance relation with; origin- phylogroups, origin-integron and phylogroups-integrons. Com- parisons of proportions were determined using the chi-square test or Fisher's exact test.
Results
Susceptibility to Antibiotics
E. coli isolates were collected from poultry (31 isolates), bo- vine (27 isolates). Among 31 poultry E. coli isolates (Table 1), high rates of resistance were observed for the tetracycline (90.32%), sulfonamide (70.51%), trimethoprim/sulfamethoxaz- ole and streptomycin with same rate (64.51%), nalidixic acid (61.29%), amoxicillin and ticarcillin with same rate (58 %). Moderate frequencies of resistance were noted for ciprofloxacin (32.25%) as well as a lowly rate were observed to amoxicillin/ clavulanic acid, cefotaxim and ceftazidime with a same rate (3.2 %). In addition, for bovin E. coli isolates high rates of resis- tance were observed for the streptomycin (55.5%), tetracycline (37%), Moderate lowly frequencies of resistance was noted for amoxicillin (18.5%), trimthoprim/sulfamethoxazole (11.11%), and sulfonamide, nalidixic acid with same rate (7.4%). Lowely frequencie of resistance was noted for ciprofloxacin (3.7%). No resistance to imipenem was detected. No ESBL producing E. coli isolate was detected. However, it was remarkable that the majority of the resistant isolates were of poultry origin (Table 1).We found that 10 (32.25%) poultry isolates were (resistant to five antibiotic families or more) multiresistants
|
Reference of isolate |
Source / Farm |
Phy- logroups |
Resistance profile |
Int* |
VRs** des intégrons (bp) / arrangements of gene Cassettes*** |
tet |
sul |
Qac-sul1 |
|
1 |
Poultry / farm Sousse |
B1 |
AMX, TIC, S, SXT, SSS, TE, AN, CIP |
1 |
1000+280/ dfrA12+orfF+aadA2+cm- lA1+aadA1+qacH+IS440+- sul3 |
A |
Sul1- 3 |
- |
|
2 |
|
B1 |
AMX, TIC, S, SXT, SSS , TE, AN |
1 |
1500/ dfrA1+aadA1 |
A |
Sul1-2-3 |
+ |
|
3 |
|
A1 |
AMX, TIC, S, SXT, SSS ,TE, AN |
1 |
1500+700+280/ dfrA1+aa- dA1 |
A |
Sul 2-3 |
+ |
|
4 |
|
A1 |
AMX, TIC, S, SXT, SSS, TE, AN |
1 |
1500+700+280/ dfrA12+orfF+aadA2 |
B |
Sul1-2 |
+ |
|
5 |
|
D2 |
S, SXT, SSS, TE |
1 |
280/ dfrA1+aadA1 |
A |
Sul1 |
+ |
|
6 |
|
A1 |
AMX, TIC, SSS , TE, AN, CIP, |
1 |
1500/ dfrA1+aadA1 |
- |
Sul1 |
+ |
|
7 |
|
B22 |
TE, AN |
- |
- |
A |
- |
- |
|
8 |
|
A1 |
AMX, TIC, S, SXT, SSS, TE |
1 |
1500/ dfrA5 |
- |
Sul2 |
+ |
|
9 |
|
A0 |
AMX, TIC, S, SXT, SSS, TE |
1 |
1500/ dfrA17+aadA1 |
A |
Sul2 |
+ |
|
10 |
|
B1 |
S, GM, SSS, TE, AN |
1 |
1000/ aadA1 |
- |
Sul1 |
+ |
|
11 |
|
D1 |
AMX, TIC, S, SXT, SSS, TE |
1 |
1000+280+600 |
- |
- |
- |
|
12 |
|
A1 |
AMX, TIC, S, SXT, SSS, TE |
- |
- |
A |
- |
- |
|
13 |
|
B1 |
SXT, SSS , TE, AN, CIP |
1 |
350+1700 |
- |
Sul1 |
+ |
|
14 |
|
D1 |
AMC, TIC, S, SSS, SXT, TE, AN |
1 |
- |
A |
- |
- |
|
15 |
|
B1 |
AMX, TIC, S, TE, SSS, AN, CIP |
- |
- |
A |
- |
- |
|
16 |
|
B1 |
AMX, AMC, S, TE, AN, CIP, CAZ, CTX |
1 |
- |
B |
- |
- |
|
17 |
|
B1 |
AMX, TIC, S, SSS, SXT, TE, AN, CIP |
1 |
1500 |
- |
Sul1 |
+ |
|
18 |
|
D2 |
AMX, TIC, S, SXT, TE, AN, CIP |
1 |
300 |
A |
- |
- |
|
19 |
|
B1 |
AMX, TIC, S, SXT, , SSS, TE, AN |
1 |
- |
A |
Sul2 |
- |
|
20 |
Poultry / Farm Sousse (2) |
A1 |
TE |
- |
- |
A |
- |
- |
|
21 |
|
A1 |
TE |
- |
- |
B |
- |
- |
|
22 |
|
A0 |
SENSIBLE |
- |
- |
- |
- |
- |
|
23 |
|
A1 |
TE |
1 |
1500 |
A |
- |
- |
|
24 |
|
A0 |
SXT, SSS |
- |
- |
- |
Sul1 |
+ |
|
25 |
|
A0 |
SENSIBLE |
- |
- |
- |
- |
- |
|
26 |
Poultry/ Borej cedria |
B1 |
TIC, S, SXT, SSS, TE,AN, CIP |
1 |
1500 dfrA12+orfF+aadA2 |
A-B |
Sul1-2 |
+ |
|
27 |
|
A1 |
AMX, TIC, SXT, SSS, TE,AN, CIP |
1 |
1000 dfrA1+aadA1 |
A |
Sul1 |
+ |
|
28 |
|
A1 |
S, SXT, SSS, TE,AN, CIP |
- |
1500/ dfrA17+aadA1 |
B |
Sul1 |
+ |
|
29 |
Poultry/ tunis |
A1 |
AM, TIC, S, SXT, SSS, TE |
2 |
2000/dfrA1-satA1-aadA1 |
A-B |
Sul1-3 |
+ |
|
30 |
|
D1 |
K, TE,AN |
2 |
2000/ dfrA1-satA1-aadA1 |
A |
- |
- |
|
31 |
|
A1 |
AM, TIC, K, S, SXT, SSS, TE,AN |
1 |
1500 |
B |
Sul1 |
+ |
|
1 |
bovin/ Far- momElhani Bizerte |
B1 |
K, S, SXT, SSS, TE |
1 |
1500/ |
A |
Sul1 |
+ |
|
2 |
|
B1 |
K, TE |
- |
- |
A |
- |
- |
|
3 |
|
A1 |
SXT, TE |
1 |
- |
A |
- |
- |
|
4 |
|
B1 |
AMX, AMC, TE, AN, CIP |
1 |
- |
A |
- |
- |
|
5 |
|
B1 |
AMX, AMC, K, S, TE, SXT, SSS, AN |
1 |
2000+1500+1200+500+280/ dfrA1+aadA1 |
A-B |
Sul 1 |
+ |
|
6 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
7 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
8 |
|
D1 |
S, TE |
- |
- |
A |
- |
- |
|
9 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
10 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
11 |
|
B1 |
S |
- |
- |
- |
- |
- |
|
12 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
13 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
14 |
|
B1 |
S |
- |
- |
- |
- |
- |
|
15 |
|
B22 |
S |
- |
- |
- |
- |
- |
|
16 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
17 |
|
A0 |
SENSIBLE |
- |
- |
- |
- |
- |
|
18 |
|
D1 |
SENSIBLE |
- |
- |
- |
- |
- |
|
19 |
|
B22 |
S,TE |
- |
- |
A |
- |
- |
|
20 |
|
B22 |
AMX, S, TE |
- |
- |
A |
- |
- |
|
21 |
|
B22 |
AMX, TE |
1 |
- |
A |
Sul1 |
+ |
|
22 |
Bovin / Farm Gbeli |
A1 |
AMX, S, TE |
2 |
2000 / dfrA1-satA1-aadA1 |
A |
Sul2 |
- |
|
23 |
|
D1 |
S |
- |
- |
- |
- |
- |
|
24 |
|
A1 |
S |
1 |
280 |
- |
Sul1 |
- |
|
25 |
|
A0 |
S |
- |
- |
- |
- |
- |
|
26 |
|
A1 |
S |
- |
- |
- |
- |
- |
|
27 |
|
B1 |
SENSIBLE |
- |
- |
- |
- |
|
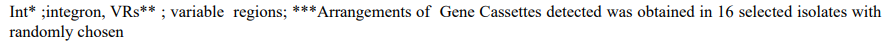
Table 1: Phenotypic and genotypic characteristics of fecal poultry (N=31) and bovine ( N=27) isolates.
Occurrence of Integrons and Determination of the Vari- able Region of the Integron Class 1 and 2 by PCR and Oc- currence of Arrangements of Resistance Gene Cassettes De- tected By Sequencing
For poultry E.coli isolates class 1 and 2 integrons were detected in 22 isolates; twenty-six isolates harbored the class 1 integrons and only in three isolates was found class 2 integron,. Accord- ing to the origin of integron-positive isolates, 22 were poultry isolates and 7 bovine isolates. The 3’CS (qacEΔ- sul1) mainly present in the typical class 1 integron were found only in 17 among 22 class 1 integron-positive poultry isolates, whereas, 3 isolates (37 %) among 7 class 1 integron-positive isolates were detected in bovin origin. It is interesting to note that isolates containing integrons were more resistant to antimicrobial agents than integron-negatives isolates (Tables 1). Both of class 1 and 2 integrons were not found in our isolates. Statistical analysis showed that class 1 integrons were significantly associated with poultry origin (p=0.001).
The VRs were detected in twenty four isolates (82.75%) amongst twenty nine class 1 and 2 integron. The size were from 280bp to 1700 in eighteen poultry int1-positive isolates; 1500 (9 isolates), 1000+700+280 (2 isolates) and for six isolates everyone have a alone or combination of VRs likes; 1000pb+280pb+600pb, 1700pb+350pb, 1000pb+280pb, 1000pb, 300pb and 280pb re- spectively. Whereas, in bovine origin the VRs of class1 integron were detected in 3 isolates with a size of 1500pb and in one isolate with 280pb. While, the combination of VRs; 1200pb+1500pb+ 1200pb+500pb+ 280pb was detected only one in isolate with the arrangements of resistance gene cassettes detected dfrA12+ orfF+aadA2+ cmlA1+aadA1+qacH+IS440+sul3. The VRs of class2 integron were detected in poultry and bovin in 2 and 1 isolates, respectively, and showed the arrangement gene cas- sette; dfrA1-satA1-aadA1 in one identical DNA fragments with approximate size of 2000 bp: Six arrangements of resistance gene cassettes of class 1 integron were detected; dfrA1+aadA1 (5isolates); for dfrA17+aadA1, dfrA12+ orfF+aadA2 each one two 2isolates; one isolate for aadA1 and dfrA5, respectively.
Genes Encoding Tetracycline-, Sulphonamide- Resistance and ESBL Production
Thirty eight were resistant to tetracycline. Genes of tet-type were carried in 32 devised in 22 (78.75%) poultry isolates and 10 bovin isolates. The genes tetA, tetB genes were observed in 27 (84.37 %) and 8 (25%) resistant isolates, respectively. TetA-tetB genes were simultaneously found in 3 isolates, while tetC was not detected in our collection. Among thirty one poultry isolates, twenty two (70.96%) isolates were sulfonamide resistant and sul-type genes were detected in eighteen isolates while qac-sul1 was detected in 16 isolates.
The genes sul1, sul2, and sul3 were detected in 14 (77.77 %), 7 (38.88%), and 4 (22.22%) isolates, respectively. The combina- tions of following genes were identified (number of isolates): sul1+sul2: (2); sul1+sul3 (2); sul2+sul3 (1) and sul1+sul2+sul3 (1). In addition, sulfonamide resistant encoded by sul1 and sul2 genes in 4 and 1 bovin isolates, respectively with absence of sul3-type. We showed a positive relation between the presence of integrons and the detection of genes encoding resistance to tetracycline and sulfonamides (Table 1).
Determination of Phylogroups
In poultry, 16 isolates were found to belong to phylogroup A (sub- groupA1: 12, sub- groupA0: 4); 9 to B1, 1 to B2 and 5 to phylogroup D. However, in bovine 9 isolates have the phy- logroups A1, 7 isolates B1, 4 isolates B2, and 3 isolates found to phylogroup D.
Discussion
We collected fifty eight E. coli strains isolated from healthy poultry and bovins recovred from farms which located in four different governorates of Tunisia (Sousse, Tunis, Bizerte and Gbeli). We studied the antibiotic susceptibility and the results showed that among 31 poultry E. coli isolates high rates of re- sistance were observed for the tetracycline (90.32%), nalidixic acid (61.29%), sulfonamide (70.96%), and trimethoprim-sulfa- methoxazole (64.51%). Our results were in agreement with oth- er Tunisia studie of 136 fecal poultry E. coli isolated from thirty six different farms which showed high rates resistance of tet- racycline (94%), nalidixic acid (89.5%) and trimethoprim-sul- famethoxazole (73. 1%) [11]. Moreover, our previous study suggest that poultry isolates were considerate trimethoprim-sul- famethoxazole resistance rate, it is similar to our previous results [21].
However, moderate frequencies of resistance were noted for amoxicillin and ticarcillin with same rate (58%), ciprofloxacin (32.25%), as well as, a lowly rate were observed to amoxicil- lin/clavulanic acid, cefotaxim and ceftazidime with a same rate (3.2 %). In addition, for bovin E. coli isolates, high rates of re- sistance were observed for the streptomycin (55.5%), tetracy- cline (37%). Moderate frequencie of resistance was noted for amoxicillin (18.5%) such as lowly rate of resistance were noted to trimethoprim-sulfamethoxazole (11.11%), nalidixic acid and sulfonamides with same rate (7.4%) and ciprofloxacin (3.7%). Our results were in agreement with other studies (Jiang et al., 2009; Lei et al., 2010) which showed high rates of tetracycline resistance (90.8 %- 95.2 %) and streptomycin (46 %).
Our study presented high rates of the two antimicrobial tetracy- cline and streptomycine in poultry and bovin E. coli isolates, this was having been mainly reported from farmed animals world- wide [2,22,23]. However, the prevalence to streptomycin resis- tance in E. coli was variable according to geographic area and husbandary conditions [2,22,23]. In addotion, high rate of resis- tance against trimethoprim-sulfamethoxazole and sulfonamides were observed in poultry origin among our collection, in con- trast lowly prevalences of resistance to the same two antimicro- bials were found in bovin isolates. Also, this finding is similar to that reported in other countries for E. coli strains isolated from healthy animals and food-products of animal origins [24-28].
No ESBL-producing isolate was identified amongst our col- lection, while, similary Soufi et al., have not identified any ES- BL-producing E. coli isolate from a collection of 164 isolates; therefore, in our work the ESBL-producer population might be underestimated owing to the randomly selection of one colo- ny per sample grown on unselective media [21,29]. Recently, ESBL-producing E. coli isolates have been reported in animal worldwide especially from poultry origin [30-33]. In Tunisia, the presence of ESBL producers has been previously reported in E. coli from poultry, pets, dromedary, and meat of various animals [9,31,10-12,34]. In those studies, ESBL production was detected by using a selective protocol.
Amongst fifty eight isolates class 1 and class 2 integrons were found in twenty-nine isolates of E. coli. Integrons of class 1 were found in twenty six isolates while class 2 only in three isolates, according to the origin of integron-positive isolates, 22 were poultry isolates and 7 bovine isolates, our results were in agree- ment with other works which showed the dominance of integron class 1 in animal-derived and human E. coli isolates [21,29]. However, the presence of integrons were most frequently in poultry origin then bovin origin and this was confirmed by the statistical analysis which found that there is a correlation be- tween poultry origin and class 1 integron (p=0.001).
Class 1 integron is a dynamic genetic system encoding a func- tional integrase protein enabling integration of several gene cas- settes in the variable regions, which were all expressed due to the presence of a common promoter. This genetic trait, explain in part the high dominance of class 1 integrins in Enterobacte- riaceae, particularly in resistant isolates and in a rich-antibiotic environment. The 3’CS (qacEΔ- sul1) found generally in class 1 integron was detected only in 17 poultry among 22 class 1 inte- gron-positive isolates, whereas, 3 isolates (37%) among 7 class 1 integron-positive isolates were detected in bovin origin. This finding was in agreement with other recent studies reporting the absence of this region in class 1 integron [21,29,31,35].
The size of VRs detected by PCR were from 280bp to 1700pb in int1-positive isolates and an approximate size of 2000pb in class 2 integrons which unable to integrate new gene cassettes into the variable region and was not functional [5].The variable region of class2 integron mainly carries dfrA1 (encoding trimethoprim resistance), sat1 (encoding streptomricin resistance) and aadA1 (encoding streptomycin/spectinomycin resistance) [5,21,29]. The fragment of the VR of class 1 integron sized 280pb cannot correspond to any gene cassette. In a genetic point of view it is plausible that currently a phenomenon of crucial genetic rear- rangements in class 1 integron is happening. This shifting might argument for the evolution toward more successful structures in capitation and dissemination of resistance. Indeed, this region was replaced in classical class 1 integron by a 'transposon-like' structure, a qacHΔ-IS440-sul3, which could facilitate the dis- semination of class 1 integron by a mechanism of transposition [36].
We choose randomly for sequencing of VRs for poultry origin 12, 2 isolates which harbored class 1 integrons and 2, respec- tively and 2 isolates for bovine each one harbored different class of integron. In 15 class 1-positive isolates, the VR showed the presence of six arrangements of gene cassettes; we notice that the gene cassette array dfrA1-aadA1 was the most frequently de- tected, found in ten isolates (5/15, 33.33 %). Furthermore, there were one isolates harbouring long arrangement of gene cassettes with un-classical 3' CS: dfrA12-orf F-aadA2-cmlA1-aadA1-qa- cHΔ-IS440-sul3. This structure was also reported by other au- thors [29,37]. This phenomenon of "substitution» (qacEΔ-sul1/ qacHΔ-IS440-sul3) might be for perfection rather than 'change of function’. Indeed, the qacΔH and sul3 genes inserted code for same functions as the genes qacEΔ (resistance the quaternary ammonium) and sul1 (resistance in sulfonamides), respectively. Genetically, this could be explained by the continuous use of the ammonium-quaternary and sulfonamides till the current days in avian industries. The variable regions in class 2 integrons, pre- sented in three isolates, were identical, dfrA1-sat1-aadA1. This structure is mainly common in all class 2 integrons identified worldwide [35].
We showed a positive relation between the presence of integrons and the detection of genes encoding resistance to tetracycline and sulfonamides. Tetracycline resistance was observed in 32 out of 58 isolates (84.2 %), a rate closer to that reported in our previous work [21]. The absence of detection of resistance genes in some of our resistant isolates could indicate that they might contain other non-tested or unknown resistance genes. Genes of tet-type were carried in 32 devised in 22 (78.75%) poultry isolates and 10 bovin isolates. The genes tetA, tetB genes were observed in 27 (84.37 %) and 8 (25%) resistant isolates, respectively. Te- tA-tetB genes were simultaneously found in 3 isolates, while tetC was not detected in our collection. The genes tetA and tetB were simultaneously found in six isolates. The predominance of tetA gene was also reported by other studies while the tetC gene is scarcely reported [38-40]. A genetic linkage on conjugative plasmids of tetA gene and class 1 integron was reported by Sun- de and Norstrom, this is in agreement with our results where the majority of isolates containing tetA also harboured the integron of class 1 [27]. This suggests the possibility of co-localization of the genes of tet-type and integrons; however, more thorough genetic studies were necessary to confirm this hypothesis.
Among thirty one poultry isolates, twenty two (70.96%) isolates were sulfonamide resistant and sul-type genes were detected in eighteen isolates, resistance in the remaining isolates might be due to chromosome mutation in dihydropteroatic synthetase DHPS [41].
The genes sul1, sul2, and sul3 were detected in 14 (77.77 %), 7 (38.88%), and 4 (22.22%) isolates, respectively while qac- sul1 was detected in 16 isolates. The combinations of follow- ing genes were identified (number of isolates): sul1+sul2: (2); sul1+sul3 (2); sul2+sul3 (1) and sul1+sul2+sul3 (1). In addition, sulfonamide resistant encoded by sul1 and sul2 genes in 4 and 1 bovin isolates, respectively with absence of sul3-type.
In agreement with our results, the gene sul1 remains by far the most frequently found gene followed by the gene sul2, while sul3 generally less frequent [42,43]. However, in one of the previous studies of our laboratory, the sul3 gene was the most prevalent one in avian E. coli isolates followed by sul1 and sul 2, respectively [29]. It is interesting to note that the majority of integron containing isolates also harboured the sul genes, that might be linked to the structure of class 1 integron which contain sul1 gene in classical 3 'CS and also sul3 in some isolates. For the distribution of isolates according to different phylogroups 29 isolates were found to belong to phylogroup A and 16 to phy- logroups B1, in the literature groupsA and B1 in human E.coli strains are considered non- virulent commensal strains contrary to animal E.coli isolates belonged to phylogroup A and B1 such as the ESBL strains, which mainly virulent [11,44,34]. Gener- ally, many studies noted that pathogenic strains producing ex- tra-intestinal infections (ExPEC) belong mainly to the B2 group and D group. Among our collection we detected only 5 isolates belonged to phylogroup B2 and 8 to phylogroup D [45-50].
Taken together, these findings highlight the importance of poul- try E. coli as reservoir of antibiotic resistance that certainly linked to the excessive use of antibiotic especially for tetracy- cline and sulfonamide in avian husbandry in Tunisia more than in bovine farms. This dramatically situation is certainly not spe- cific to Tunisia. Therefore, this is worrisome for global human health, especially with the increasing consumption of poultry meat in Tunisia and in other part of the world owing to its rela- tively lower cost comparing to red meat
Acknowledgements
Kilani hajer has a fellowship from the Tunisian Ministry of Higher Education and Scientific Research.
Conflict of interest
statement None of the authors of this paper has a financial or personal re- lationship with other people or organizations that could inappro- priately influence or bias the content of the paper.
References
- Wassenaar, T. M., & Silley, P. (2008). Antimicrobial resis- tance in zoonotic bacteria: lessons learned from host-spe- cific pathogens. Animal Health Research Reviews, 9(2), 177-186.
- Maynard, C., Fairbrother, J. M., Bekal, S., Sanschagrin, F., Levesque, R. C., Brousseau, R., ... & Harel, J. (2003). An- timicrobial resistance genes in enterotoxigenic Escherich- ia coli O149: K91 isolates obtained over a 23-year period from pigs. Antimicrobial agents and chemotherapy, 47(10), 3214-3221.
- Smet, A., Martel, A., Persoons, D., Dewulf, J., Heyndrickx, M., Herman, L., ... & Butaye, P. (2010). Broad-spectrum β-lactamases among Enterobacteriaceae of animal origin: molecular aspects, mobility and impact on public health. FEMS microbiology reviews, 34(3), 295-316.
- Baquero, F. (2004). From pieces to patterns: evolutionary engineering in bacterial pathogens. Nature Reviews Micro- biology, 2(6), 510-518.
- Partridge, S. R., Recchia, G. D., Stokes, H. W., & Hall, R.M. (2001). Family of class 1 integrons related to In4 from Tn 1696. Antimicrobial agents and chemotherapy, 45(11), 3014-3020.
- Preston, K. E., Graffunder, E. M., Evans, A. M., & Venezia,R. A. (2003). Survey of plasmid-associated genetic markers in Enterobacteriaceae with reduced susceptibilities to ceph- alosporins. Antimicrobial agents and chemotherapy, 47(7), 2179-2185.
- Villa, L., Pezzella, C., Tosini, F., Visca, P., Petrucca, A., & Carattoli, A. (2000). Multiple-antibiotic resistance mediatedby structurally related IncL/M plasmids carrying an extend- ed-spectrum β-lactamase gene and a class 1 integron. Anti- microbial agents and chemotherapy, 44(10), 2911-2914.
- Wang, M., Tran, J. H., Jacoby, G. A., Zhang, Y., Wang, F., & Hooper, D. C. (2003). Plasmid-mediated quinolone re- sistance in clinical isolates of Escherichia coli from Shang- hai, China. Antimicrobial agents and chemotherapy, 47(7), 2242-2248.
- Jouini, A., Vinué, L., Slama, K. B., Saenz, Y., Klibi, N., Hammami, S., ... & Torres, C. (2007). Characterization of CTX-M and SHV extended-spectrum β-lactamases and as- sociated resistance genes in Escherichia coli strains of food samples in Tunisia. Journal of Antimicrobial Chemothera- py, 60(5), 1137-1141.
- Ben Sallem, R., Ben Slama, K., Sáenz, Y., Rojo-Bezares, B., Estepa, V., Jouini, A., ... & Torres, C. (2012). Prevalence and characterization of extended-spectrum beta-lactamase (ESBL)–and CMY-2–producing Escherichia coli isolates from healthy food-producing animals in Tunisia. Foodborne pathogens and disease, 9(12), 1137-1142.
- Mnif, B., Ktari, S., Rhimi, F. M., & Hammami, A. (2012). Extensive dissemination of CTXâ?Mâ?1â?and CMYâ?2â?produc- ing Escherichia coli in poultry farms in Tunisia. Letters in applied microbiology, 55(6), 407-413.
- Grami, R., Mansour, W., Dahmen, S., Mehri, W., Haenni, M., Aouni, M., & Madec, J. Y. (2013). The bla CTX-M-1 IncI1/ST3 plasmid is dominant in chickens and pets in Tuni- sia. Journal of Antimicrobial Chemotherapy, 68(12), 2950- 2952.
- Carattoli, A. (2001). Importance of integrons in the diffu- sion of resistance. Veterinary research, 32(3-4), 243-259.
- Liebert, C. A., Hall, R. M., & Summers, A. O. (1999). Trans- poson Tn 21, flagship of the floating genome. Microbiology and molecular biology reviews, 63(3), 507-522.
- Arakawa, Y., Murakami, M., Suzuki, K., Ito, H., Wacharo- tayankun, R., Ohsuka, S., ... & Ohta, M. (1995). A novel integron-like element carrying the metallo-beta-lactamase gene blaIMP. Antimicrobial agents and chemotherapy, 39(7), 1612-1615.
- Biskri, L., & Mazel, D. (2003). Erythromycin esterase gene ere (A) is located in a functional gene cassette in an unusual class 2 integron. Antimicrobial agents and chemotherapy, 47(10), 3326-3331.
- Rowe-Magnus, D. A., & Mazel, D. (2002). The role of in- tegrons in antibiotic resistance gene capture. International Journal of Medical Microbiology, 292(2), 115-125.
- Domingues, S., da Silva, G. J., & Nielsen, K. M. (2012). In- tegrons: vehicles and pathways for horizontal dissemination in bacteria. Mobile genetic elements, 2(5), 211-223.
- Bradford, P. A. (2001). Extended-spectrum β-lactamases in the 21st century: characterization, epidemiology, and detec- tion of this important resistance threat. Clinical microbiolo- gy reviews, 14(4), 933-951.
- Sáenz, Y., Vinué, L., Ruiz, E., Somalo, S., Martínez, S., Ro- jo-Bezares, B., ... & Torres, C. (2010). Class 1 integrons lacking qacEΔ1 and sul1 genes in Escherichia coli isolates of food, animal and human origins. Veterinary microbiolo- gy, 144(3-4), 493-497.
- Soufi, L., Abbassi, M. S., Sáenz, Y., Vinué, L., Somalo, S.,Zarazaga, M., ... & Torres, C. (2009). Prevalence and diver- sity of integrons and associated resistance genes in Esche- richia coli isolates from poultry meat in Tunisia. Foodborne Pathogens and Disease, 6(9), 1067-1073.
- Boerlin, P., Travis, R., Gyles, C. L., Reid-Smith, R., Heather Lim, N. J., Nicholson, V., ... & Archambault, M. (2005). Antimicrobial resistance and virulence genes of Escherichia coli isolates from swine in Ontario. Applied and environ- mental microbiology, 71(11), 6753-6761.
- Gyles, C. L. (2008). Antimicrobial resistance in selected bacteria from poultry. Animal health research reviews, 9(2), 149-158.
- Guerra, B., Junker, E., Schroeter, A., Malorny, B., Lehmann, S., & Helmuth, R. (2003). Phenotypic and genotypic char- acterization of antimicrobial resistance in German Esche- richia coli isolates from cattle, swine and poultry. Journal of Antimicrobial Chemotherapy, 52(3), 489-492.
- Kang, H. Y., Jeong, Y. S., Oh, J. Y., Tae, S. H., Choi, C. H.,Moon, D. C., ... & Lee, J. C. (2005). Characterization of an- timicrobial resistance and class 1 integrons found in Esche- richia coli isolates from humans and animals in Korea. Jour- nal of Antimicrobial Chemotherapy, 55(5), 639-644.
- Meyer, E., Lunke, C., Kist, M., Schwab, F., & Frank, U. (2008). Antimicrobial resistance in Escherichia coli strains isolated from food, animals and humans in Germany. Infec- tion, 36, 59-61.
- Sunde, M., & Norström, M. (2006). The prevalence of, as- sociations between and conjugal transfer of antibiotic re- sistance genes in Escherichia coli isolated from Norwegian meat and meat products. Journal of Antimicrobial Chemo- therapy, 58(4), 741-747.
- Van, T. T. H., Moutafis, G., Tran, L. T., & Coloe, P. J. (2007). Antibiotic resistance in food-borne bacterial con- taminants in Vietnam. Applied and environmental microbi- ology, 73(24), 7906-7911.
- Soufi, L., Sáenz, Y., Vinué, L., Abbassi, M. S., Ruiz, E., Zarazaga, M., ... & Torres, C. (2011). Escherichia coli of poultry food origin as reservoir of sulphonamide resistance genes and integrons. International journal of food microbi- ology, 144(3), 497-502.
- Blanc, V., Mesa, R., Saco, M., Lavilla, S., Prats, G., Miró, E., ... & Llagostera, M. (2006). ESBL-and plasmidic class C β-lactamase-producing E. coli strains isolated from poul- try, pig and rabbit farms. Veterinary microbiology, 118(3-4), 299-304.
- Slama, K. B., Jouini, A., Sallem, R. B., Somalo, S., Sáenz, Y., Estepa, V., ... & Torres, C. (2010). Prevalence of broad-spectrum cephalosporin-resistant Escherichia coli isolates in food samples in Tunisia, and characterization of integrons and antimicrobial resistance mechanisms impli- cated. International journal of food microbiology, 137(2-3), 281-286.
- Cortés, P., Blanc, V., Mora, A., Dahbi, G., Blanco, J. E., Blanco, M., ... & Llagostera, M. (2010). Isolation and char- acterization of potentially pathogenic antimicrobial-resis- tant Escherichia coli strains from chicken and pig farms in Spain. Applied and environmental microbiology, 76(9), 2799-2805.
- Trott, D. (2013). β-lactam resistance in gram-negativepathogens isolated from animals. Current pharmaceutical design, 19(2), 239-249.
- Kilani, H., Abbassi, M. S., Ferjani, S., Mansouri, R., Sghai- er, S., Ben Salem, R., ... & Boubaker, I. B. B. (2015). Oc- currence of bla CTX-M-1, qnr B1 and virulence genes in avian ESBL-producing Escherichia coli isolates from Tuni- sia. Frontiers in cellular and infection microbiology, 5, 38.
- Chah, K. F., Agbo, I. C., Eze, D. C., Somalo, S., Estepa, V., & Torres, C. (2010). Antimicrobial resistance, integrons and plasmid replicon typing in multiresistant clinical Escherich- ia coli strains from Enugu State, Nigeria. Journal of Basic Microbiology, 50(S1), S18-S24.
- Antunes, P., Machado, J., & Peixe, L. (2007). Dissemina- tion of sul3-containing elements linked to class 1 integrons with an unusual 3′ conserved sequence region among Sal- monella isolates. Antimicrobial agents and chemotherapy, 51(4), 1545-1548.
- Sunde, M., Solheim, H., & Slettemeås, J. S. (2008). Genetic linkage between class 1 integrons with the dfrA12-orfF-aa- dA2 cassette array and sul3 in Escherichia coli. Veterinary Microbiology, 130(3-4), 422-425.
- Bryan, A., Shapir, N., & Sadowsky, M. J. (2004). Frequency and distribution of tetracycline resistance genes in geneti- cally diverse, nonselected, and nonclinical Escherichia coli strains isolated from diverse human and animal sources. Ap- plied and environmental microbiology, 70(4), 2503-2507.
- Zhang, T., Wang, C. G., Lv, J. C., Wang, R. S., & Zhong,X. H. (2012). Survey on tetracycline resistance and antibi- otic-resistant genotype of avian Escherichia coli in North China. Poultry Science, 91(11), 2774-2777.
- Koo, H. J., & Woo, G. J. (2011). Distribution and transfer- ability of tetracycline resistance determinants in Escherich- ia coli isolated from meat and meat products. International journal of food microbiology, 145(2-3), 407-413.
- Sköld, O. (2001). Resistance to trimethoprim and sulfon- amides. Veterinary research, 32(3-4), 261-273.
- Hammerum, A. M., Sandvang, D., Andersen, S. R., Sey-farth, A. M., Porsbo, L. J., Frimodt-Møller, N., & Heuer, O.E. (2006). Detection of sul1, sul2 and sul3 in sulphonamide resistant Escherichia coli isolates obtained from healthy hu- mans, pork and pigs in Denmark. International journal of food microbiology, 106(2), 235-237.
- Trobos, M., Jakobsen, L., Olsen, K. E., Frimodt-Møller, N., Hammerum, A. M., Pedersen, K., ... & Olsen, J. E. (2008). Prevalence of sulphonamide resistance and class 1 integron genes in Escherichia coli isolates obtained from broilers, broiler meat, healthy humans and urinary infections in Den- mark. International journal of antimicrobial agents, 32(4), 367-369.
- Huber, H., Zweifel, C., Wittenbrink, M. M., & Stephan, R. (2013). ESBL-producing uropathogenic Escherichia coli isolated from dogs and cats in Switzerland. Veterinary mi- crobiology, 162(2-4), 992-996.
- Cantón, R., González-Alba, J. M., & Galán, J. C. (2012). CTX-M enzymes: origin and diffusion. Frontiers in micro- biology, 3, 110.
- Clinical and Laboratory Standards Institute (CLSI), (2012). Perforance Standards for Antimicrobial Susceptibility Test- ing; Twenty-second Informational Supplement. Clinicaland Laboratory Standards Institute Document M100-S22. Clinical and Laboratory Standards Institute, Wayne, PA.
- Jiang, H. X., Lü, D. H., Chen, Z. L., Wang, X. M., Chen, J.R., Liu, Y. H., ... & Zeng, Z. L. (2011). High prevalence and widespread distribution of multi-resistant Escherichia coli isolates in pigs and poultry in China. The veterinary journal, 187(1), 99-103.
- Lei, T., Tian, W., He, L., Huang, X. H., Sun, Y. X., Deng, Y.T., ... & Liu, J. H. (2010). Antimicrobial resistance in Esche- richia coli isolates from food animals, animal food products and companion animals in China. Veterinary microbiology,146(1-2), 85-89.
- Pitout, J. D., & Laupland, K. B. (2008). Extended-spectrum β-lactamase-producing Enterobacteriaceae: an emerging public-health concern. The Lancet infectious diseases, 8(3), 159-166.
- Lim, S. K., Lee, H. S., Nam, H. M., Cho, Y. S., Kim, J.M., Song, S. W., ... & Jung, S. C. (2007). Antimicrobial re- sistance observed in Escherichia coli strains isolated from fecal samples of cattle and pigs in Korea during 2003–2004. International journal of food microbiology, 116(2), 283-28



