Research Article - (2022) Volume 6, Issue 1
Comparative Analysis of Orthologous Mast Cell Chymases Active Sites Using Bioinformatics Tools
2Institute of Biological Sciences, Sarhad University of Science and Information Technology, Peshawar, Pakistan
Received Date: Mar 24, 2022 / Accepted Date: Apr 06, 2022 / Published Date: Apr 18, 2022
Copyright: ©Copyright: Ã?©2022 Jawairia Kiran. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Jawairia Kiran, Sadaf Chiragh, (2022). Comparative Analysis of Orthologous Mast Cell Chymases Active Sites Using Bioinformatics Tools. Stem Cell Res Int 6(1), 21-30.
Abstract
Mast cells, neutrophils, basophils, natural killer cells, and cytotoxic T cells are among the hematopoietic cell lines originating from immune cells. The importance of mammalian mast cells in innate immunity has piqued the scientific community's interest. cytoplasmic granules of mast cells. Histamine, proteoglycans, proteins, and cytokines are examples of compounds (mediators) that produce mast cell cytoplasmic granules. When external stimuli are received, these granules are released, causing degranulation. Mast cells generate a lot of proteases including chymase, tryptase, and type A carboxypeptidase, among other things. The structure of chymase cma1 was unknown in mice, but it shares 74 percent of its human sequence with chymase CMA1, which was used as a reference. The human CMA1 structure was used to establish and interpret the mouse cma1 structure. Significant residues from the active site, such as catalytic triads and binding residues, were examined. The position of a catalytic triad in human CMA1 chymase and mouse cma1 chymase has been discovered to be similar. Val 175 and Val 197, respectively, replaced two Ala 192 and Gly 214 residues in active site binding residues in target, according to active site residue analysis. We may deduce from this substitution that the cleavage specificity of mouse cma1 chymase varies from that of human CMA1 chymase
Keywords
Chymase, Mast Cell, Proteases, Mus Musculus, Sequence Alignment, Catalytic Triads
Introduction
Mast cells (MC) are immature cells that develop in the bone mar- row and circulate in the blood, maturing in various tissues under the influence of various factors to play a significant role in the first line of defences [1]. And they shield their hosts from pathogens including bacteria, parasites, and even snake venom [2-4].
One of the most distinguishing characteristics of MCs is the pres- ence of secretory cytoplasmic granules that secrete a variety of extracellular compounds known as "MC mediators." Biogenic amines (histamine and serotonin), cytokines, serglycine proteogly- can, lysosomal enzymes, and protease are among these substances [5]. Antigen binding to the IgE antibody, attached to the highly specific IgE receptor FC-RI [6]. Anaphylotoxin penetration, stem cell factor, endothelin-1, and various neuropeptides may all induce MC degranulation. Mc proteases are the main component of MC granules, accounting for over 25% of MC protein Both MC-con- taining proteases are kept as active enzymes [7].
Tryptase, chymase, and MC-CPA are the three forms of MC-spe- cific protaeses. Tryptase and chymase are serine proteases, while MC-CPA is a zinc-dependent metalloprotease [8]. In humans, MCs are divided into two subclasses based on their protease con- tent: MCt, which expresses only tryptase, and MCtc, which ex- presses all forms of MC proteases, including tryptases, chymases, and MC-CPA. Connective tissue MC (CTMC) and mucosal MC are the two subclasses of MC in mice (MMC). Two forms of chy- mases (mouse mast cell protease-4, mMCP-4 and mMCP-5), two tetrameric tryptases (mMCP-6 and mMCP-7) and an MC-CPA are all expressed by CTMC. The MMC, on the other hand, expresses two forms of chymases (mMCP-1 and mMCP-2) [9].
Chymase cleaves after aromatic amino acid residues in the same way as chymotrypsin does. Chymases, unlike tryptase, are mono- meric enzymes. MC chymases are divided into two classes based on their phylogenetic relationships: the α- and the β- classes [10]. The human MC-chymase gene (CMA1) is found on chromosome 14q11.2, a 30kb region [11]. This gene is found at one end of a four-gene cluster. The cathepsin G gene is specific to neutrophils, while the granzyme H and B genes are specific to T cells. This locus' expansion varies by species and can be separated into chy- mase and granzyme loci [12]. The MC chymase genes are found on chromosome 14C3, which spans 160kb in the mouse. The tran- scriptional orientation of the MCP-5 gene (cma1/ mcpt5) is iden- tical to that of the mMCP-4, -8, and -9 genes (mcpt4, 8, and 9) in humans [13].
Chymases are known to have a variety of anti-inflammatory properties. The injection of chymase into the skin of guinea pigs resulted in the aggregation of both neutrophils and eosinophils, according to one report. Human chymases are chemotactic for neutrophils and monocytes in vitro and play a significant role in inflammatory cell recruitment. The role of chymase in the patho- genesis of allergic skin reactions is supported by studies that link the presence of chymase to disease progression [14].
Because of the discovery that chymase may transform ANG-I to ANG-II and the potential for this operation in terms of cardiac function, several studies have suggested that chymase can play a role in heart disease [15]. Chymases are also involved in the fibrotic process. In animal models of experimental diabetes and autoimmune liver fibrosis the existence of chymases is associated with fibrosis; however, chymase inhibitors may minimise fibrosis [16-18]. The exact mechanism by which chymase causes fibrosis is unknown [19].
Although MCs play a role in angiogenesis, their exact function is unknown. Several studies have shown that chymase is involved [20]. Chymase's effect on angiogenesis can result in both patho- logical and physiological outcomes. The presence of chymase has been linked to both wound healing and tumour angiogenesis [21]. The structure of mouse cma1 was constructed and analysed in the current research, which used the structure of human CMA1 to describe the enzymatic properties. Significant residues including catalytic triads and active site binding residues were investigated.
Material and methods
Template collection and retrieval of sequences
Our target sequence, Mus musculus (mouse) chymase 1, was ob- tained from the Uniprot database and saved in fasta format. The Basic Local Alignment Search Tool (BLAST) accessible at www. ncbi.nlm.nih.gov/blast, is used to find homology between sequenc- es of known three-dimensional structures and the target sequence [22, 23]. After changing the database to Protein Data Bank, the target sequence was blast in the Uniprot database for template se- lection (PDB). The protein with the greatest similarity was chosen as the prototype. The crystal structure coordinates of chymase 1 (PDB ID: 1NN6) from Homo sapiens were used to model chymase 1 from Mus musculus [24]. The series of template human CMA1 was also saved from SWISS-PROT.
Building and refining a three-dimensional model
First, the template's layout was downloaded in PDB format from the RCSB-PDB databank (http://www.rcsb.org/pdb). The three-di- mensional homology model of target (chymase 1) was then deter- mined using crystal coordinates of template (chymase 1) based on alignment between target and template [25]. The three-dimension- al homology model of target (chymase 1) was determined using crystal coordinates of template based on alignment between target and template (chymase 1). The homology models were created us- ing MODELLER 9v7.
The three-dimensional structure of protease (Mus musculus) was constructed using modeller 9v7 and the crystal structure of 1NN6. MODELLER generates a model with low energy from a template pdb file obtained from the Protein Data Bank (PDB). Many mod- els were created using MODELER, and each model was tested by ProSa to select the models that satisfied the stereo chemical properties.
Model visualisation
For the alignment and simulation of variable surface loops and structural investigations, the newly developed models were visu- alised using the software DS Viewer [26]. The visualised structure aids in the study of atom and residue spatial association.
Model evaluation
Geometry analysis, stereochemistry, and energy distribution in models are all part of model evaluation. The models' reliability was tested using the software procheck version 3.4 [27]. This soft- ware is used to assess if the modelled model's structural features are appropriate in terms of protein structures. Procheck compares the target model's various parameters to the ideal parameters of a previously known sequence. Procheck provides a graphical repre- sentation of the result. Procheck produces a number of output files, each of which includes unique details about the protein structure. Model No. 14 was chosen from the thirty models based on sum- mary files and a Ramachandran plot developed by Procheck as the final model that met the stereo chemical requirements.
Estimation of Energy
The energy measurement and stability of the target three-dimen- sional structures were calculated using online ProSA, which can be used without having to instal the software on the device [28]. An internet connection is needed for ProSA site. The result can be retrieved via the ProSA site to determine the atomic coordinates of the model that has been uploaded in PDB format. The Z-score indicates the model's consistency. This importance can be seen in the plot, which includes the z-scores of all the protein chains that have been calculated experimentally. This plot can be used to see whether the target structure's z-score falls within the scoring range of native proteins of similar sizes.
Superimposition
Superimposition is a term used to describe a situation in which the target and prototype are superimposed to compare the variability of the newly constructed target structure to that of the template. DS Viewer was used to superimpose the images. The target file was inserted from the file menu after opening the template file in DS Viewer version 3.1. Both files were chosen and superimposed on top of each other; the display style and colour of these files could also be modified, and the result was saved as a PDB file.
Result and Discussion
BLAST result (Sequence Comparison)
The comparison of the model with other recognised proteins is the first step in homology modelling [29]. Our target's swiss prot accession number (p23946) was used to extract the primary se- quence in swiss prot. The sequence had 247 amino acids, but 21 were omitted because it was in Propeptide shape, and we needed active protein form. The sequence was downloaded as a FASTA file and matched against the PDB using the NCBI server, which can be found at http://www.ncbi.nlm.nih.gov/blast.
According to the blast result, both sequences have 74 percent iden- tity and 81 percent positives, indicating that the template and target have the same amino acids (Fig. 01). The first identical amino acid in both sequences is Methionine (M), and positive residues indi- cate amino acids that are biochemically similar. The estimated bit score is 970, while the E-value, or predicted value, is 2.6e-132, or 2.610-132.
Human Chymase 1 is identical to mouse Chymase 1 and can be used as a basis for model construction, according to BLAST re- sults. These proteins belong to the same family. As a result, we chose Homo sapiens Chymase 1 as a template for model building of the target protein (Chymase 1).
Figure 1: BLAST result of mouse cma1 with human CMA1
3D model generation
The three-dimensional structure of the target (mouse cma1) was generated from its primary sequence using the three-dimensional model of the prototype (human Chymase 1); the modeller version used was 9v7. The modeller programme was used to create thirty energy-minimized models, as well as Het atoms and other ligands, which were attached to the template. The procheck programme was used to test all thirty models, and the best one was chosen for further study. There were no dent variations between the sample and the template when they were superimposed. (See Figure 02)
Figure 2: Solid ribbon representation of target and template superimposed on each other, blue color shows template (CMA 1) and red color shows target (cma1).
3D Model Evaluation
Procheck was used to assess the newly created three-dimensional structure of the goal. Different output files are generated for each of the thirty models after Procheck has been run. On the basis of these output files, the most suitable model was chosen. For each model, ten plots were developed, with plot 1 being the Ramachan- dran plot. A software called GhostScript view (GS view) was used to view these plots. The Ramachandran plot is divided into four regions: preferred, permitted, generously allowed, and disallowed. In the case of our findings, model 14 was chosen based on the Ra- machandran plot and energy minimization. It reveals that the mod- el includes 224 residues, with 171 residues in the preferred region accounting for 88.1 percent, 21 and 2 residues in the allowed re- gion accounting for 10.8 percent, and generously allowed regions accounting for 1.0 percent. The 0.0 percent was accounted for by the disallowed area, which contains 0 residues (Fig. 03). The tem- plate (human CMA1) is seen in a Ramachandran storey (Fig. 04).
Plot statistics
|
Residues in most favored region [A, B, L] |
171 |
88.1% |
|
Residues in additional allowed region [a, b, l, p] |
21 |
10.8% |
|
Residues in generously allowed region[a, b, l, p] |
2 |
1.0% |
|
Residues in disallowed regions |
0 |
.0% |
|
Number of non glycine and non protein residues |
194 |
100.0% |
|
Number of N-residues (excl, Gly and pro) |
1 |
|
|
Number of glycine residues (shown as triangles) |
16 |
|
|
Number of proleine residues |
13 |
|
|
Total number of residues |
224 |
|
Figure 3: RAMACHANDRAN Plot by procheck for mouse chymase 1(target)
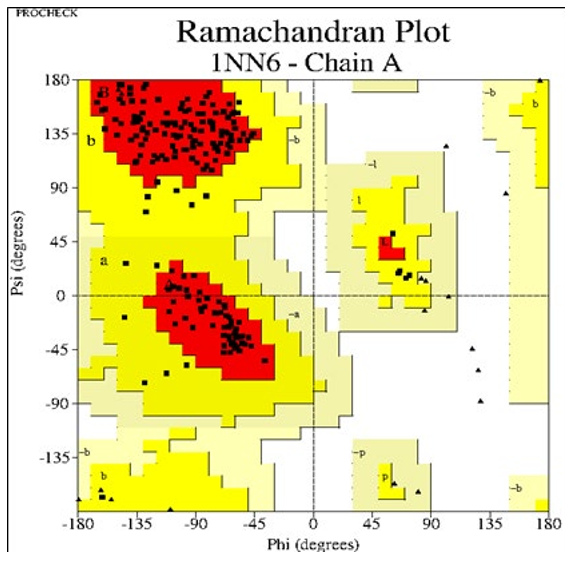
Plot statistics
|
Residues in most favored region [A, B, L] |
166 |
90.2% |
|
Residues in additional allowed region [a, b, l, p] |
18 |
9.8% |
|
Residues in generously allowed region[a, b, l, p] |
0 |
.0% |
|
Residues in disallowed regions |
0 |
.0% |
|
Number of non glycine and non protein residues |
184 |
100.0% |
|
Number of N-residues (excl, Gly and pro) |
3 |
|
|
Number of glycine residues (shown as triangles) |
20 |
|
|
Number of proleine residues |
15 |
|
|
Total number of residues |
222 |
|
Figure 4: RAMACHANDRAN Plot by procheck for mouse chymase 1(target)
Pairwise Sequence Alignment
Different methods were used to predict the evolutionary history of our target proteins in comparison to that of other similar proteins using pairwise sequence alignment. We can learn about the nucle- otide and amino acid sequences, as well as the conserved regions, by using various methods, such as BioEdit for alignment. Then, using this analogy, we can predict a protein's structural and func- tional relationships.
The alignment file of mouse cma 1 with that of human CMA1 is shown in Fig. 05. The N-terminus region has more differences than the C-terminus region; this is due to the removal of peptide sequences from the N-terminus during secretion, and it plays no role in protein function.
Figure 5: Pairwise sequence alignment of target (mouse cma1) with template (human CMA1)
Superposition
By superposing the newly constructed model (mouse cma1) with the prototype (human CMA1), we can predict the variability of the newly constructed model (mouse cma1). There was no difference between the two structures after superposition and elimination of 21 amino acids from the N-terminus, implying that our target (cma1) is nearly identical to the template (CMA1) structure. (See Figure 6)
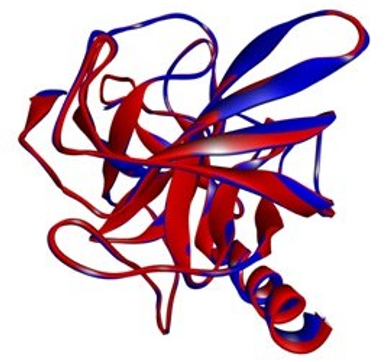
Figure 6: Superposition of the target (mouse cma1) over the tem- plate ( human CMA1
Target and Template Superposition in Propeptide Form
Chymase is synthesised as an inactive “zymogen” with an N-ter- minal pro region that prevents the zymogen from transitioning to an activated conformation, similar to the other proteases. The first 23 amino acids in our target code for the N-terminal propeptide. (Figure 7) The N-terminus region has more differences than the C-terminus region; this is due to the removal of peptide sequences from the N-terminus during secretion, and it plays no role in pro- tein function.
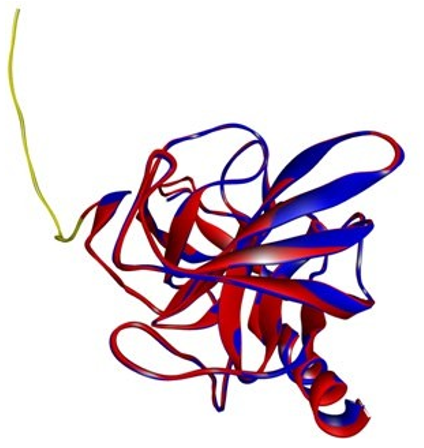
Figure 7: Shows the superimposed figure of target and template with N-terminal propeptide. (Yellow color indicates propeptide, blue indicates template while the red color indicates target.)
The Catalytic Triads of Human and Mouse Cma1: Struc- tural Review
With Ser195 on one hand and Asp102 and His57 on the other, the catalytic triads span the active site cleft. The catalytic triad (Asp102-His57-Ser195) is part of a broad hydrogen bonding net- work [30, 31]. The catalytic triad of human chymase cma1 aligns with that of mouse chymase cma1, and the catalytic triad is ar- rayed in a geometry similar to that of human chymase 1 (cma1) (Fig. 08). The three-dimensional structure of mouse chymase 1 was determined for the first time, and it provided new insight into the mechanism of action of serine proteases [32]. Because of the geometric relationship between Asp102, His57, and Ser195, it was hypothesised that His57 is involved in proton transfer from Ser195 to Asp102 in a charge relay process [30].
Serine proteases that use Ser-His-Asp in the form of the triad or charge relay system are classified into four subfamilies: chymo- trypsin, subtilisin, carboxypeptidase T, and Clp protease families [33]. The catalytic triads residues in mouse chymase 1 are con- served, and it is a serine protease, whose role is based on the inter- action of a neucleophilic Ser residue with an acidic Asp and a His residue [34]. (Fig. 9)
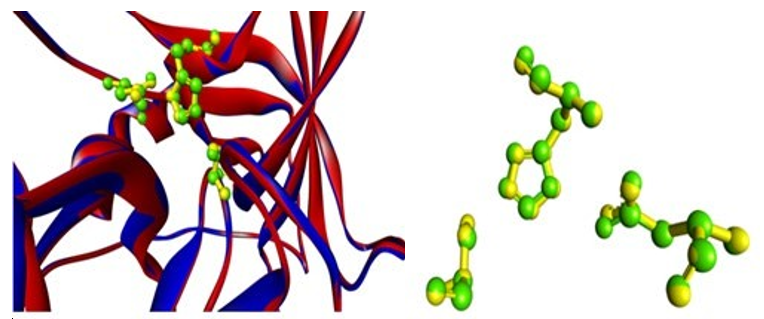
Figure 8: Superimposed Figure with Catalytic Triads of Human Chymase 1(Yellow) and Mouse Chymase1 (Green)
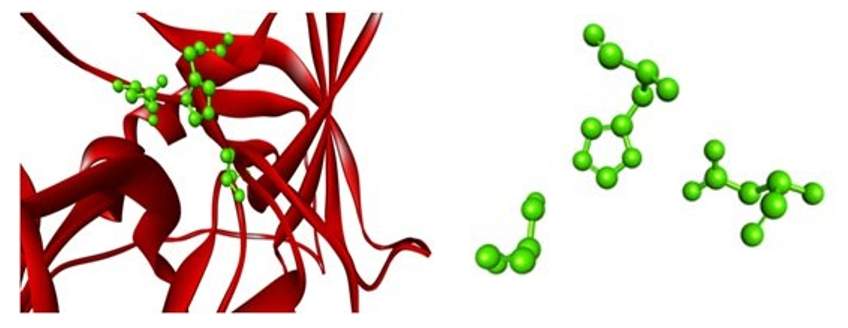
Figure 9: Mouse Chymase 1 (Target) with Catalytic Residues (Green)
The Substrate Binding Specificity of Human and Mouse
Cma1: A Structural
Study Serine proteases are graded as either trypsin chymase or elastase depending on the primary specificity of the cleavage. The prote- ases' distinctive cleavage is visible in the S1 pocket structure. S1 pocket is made up of residues 189-192, 214-216, and 224-228 of these, residues 189, 216, and 226 are of particular interest [35]. The S1 pocket base contains the amino acid Ala189, while the S1 pocket contains the amino acids Gly216 and Ala226 [36].
In our case, mouse chymase cma1 differs slightly from human chy- mase in terms of substrate binding residues, with Val at position 189 instead of Ala and Val at position 216 instead of Gly showing the same amino acid at position 226 as Ala. (Figures 10 and 11)
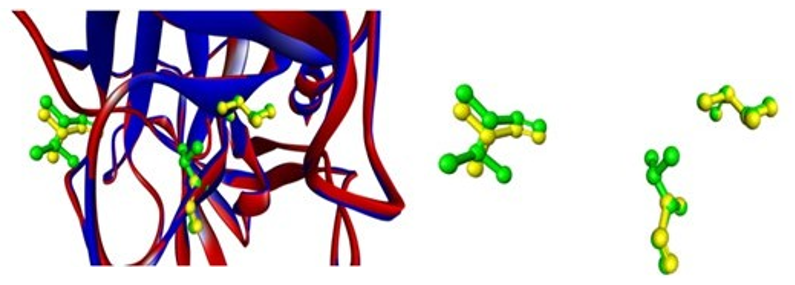
Figure 10: Superimposed figure showing substrate binding residues in target (green) and template (yellow)
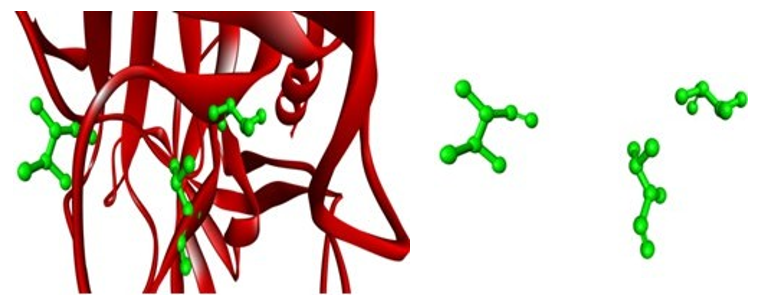
Figure 11: Showing target (mouse cma1) with substrate binding residues (green)
Results of ProSA
(Fig. 12) shows the energy graph for cma1 (target) in terms of z-score (-6.6), which indicates the overall quality of protein struc- ture. The energy graph of CMA1 (template) with a z-score (-6.53) is shown in the (Fig.13). Since our goal value is in the native vali-dation range, any positive values could be problematic for protein modelling. With the exception of a few peaks, total energy resi- dues are negative [37].
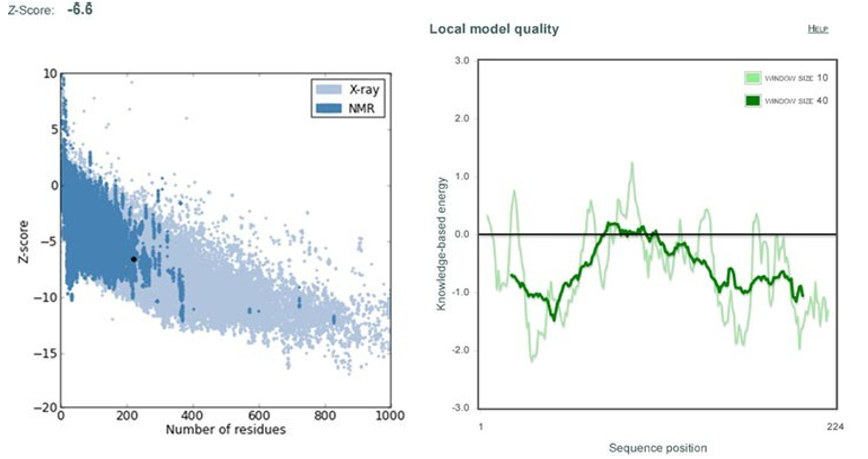
Figure 12: Shows Z-score and energy peaks of Mouse Chymase 1 (cma1) obtained through ProSa web
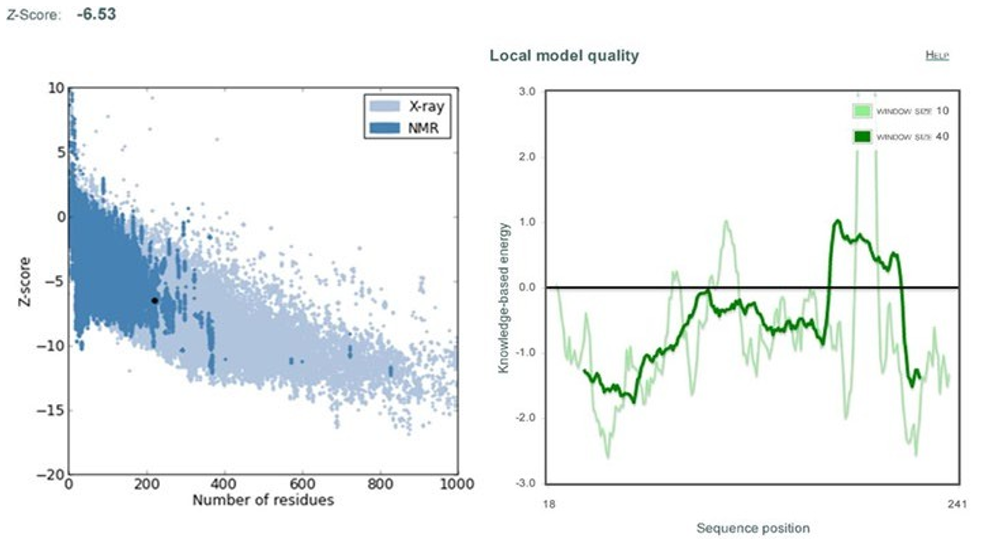
Figure 13: Shows Z-score and energy peaks of Human Chymase 1 (cma1) obtained through ProSa web
Conclusion
We concluded from our research that the catalytic traids of both the template (human chymase) and target (mouse chymase) pro- teins are the same, suggesting that both are serine proteases with catalytic traids of histidine, asparagine, and serine. The primary sequence of the structure suggests that the primary sequence of mouse CMA 1 has different amino acids, namely alanine in hu- man and valine in mouse, and glycine in human at another posi- tion, while mouse CMA1 has valine at another position and both proteins have the same amino acid, namely valine. This suggests that the catalytic activity of mouse CMA1 may vary from that of human CMA1.
It is suggested that further research be conducted to create a re- combinant protein of human CMA1 and to create a 3D structure of the protein using X-ray crystallographic and NMR techniques, which can reveal the protein's specificity as well as its catalytic activity.
References
- Yu, T., Liu, B., He, Z., Yang, M., Song, J., Ma, C., ... & Li, J.(2018). Short-term in vitro culture of purity and highly func- tional rat bone marrow-derived mast cells. In Vitro Cellular & Developmental Biology-Animal, 54(10), 705-714.
- Wernersson, S., & Pejler, G. (2014). Mast cell secretory gran- ules: armed for battle. Nature Reviews Immunology, 14(7), 478-494.
- Mukai, K., Tsai, M., Saito, H., & Galli, S. J. (2018). Mast cells as sources of cytokines, chemokines, and growth factors. Immunological reviews, 282(1), 121-150.
- Siebenhaar, F., Redegeld, F. A., Bischoff, S. C., Gibbs, B. F., & Maurer, M. (2018). Mast cells as drivers of disease and therapeutic targets. Trends in immunology, 39(2), 151-162.
- Douaiher, J., Succar, J., Lancerotto, L., Gurish, M. F., Orgill, D. P., Hamilton, M. J., ... & Stevens, R. L. (2014). Develop- ment of mast cells and importance of their tryptase and chy- mase serine proteases in inflammation and wound healing. Advances in immunology, 122, 211-252.
- Lundequist, A., & Pejler, G. (2011). Biological implications of preformed mast cell mediators. Cellular and molecular life sciences, 68(6), 965-975.
- Vitte, J. (2015). Human mast cell tryptase in biology and med- icine. Molecular immunology, 63(1), 18-24.
- Johnzon, C. F., Rönnberg, E., & Pejler, G. (2016). The role of mast cells in bacterial infection. The American journal of pathology, 186(1), 4-14.
- Carlos, B. M. (2008). Unveiling the biological role of sergly- cin proteoglycans (Vol. 2008, No. 2008: 14).
- Fu, Z., Thorpe, M., & Hellman, L. (2015). rMCP-2, the ma- jor rat mucosal mast cell protease, an analysis of its extended cleavage specificity and its potential role in regulating intesti- nal permeability by the cleavage of cell adhesion and junction proteins. PLoS One, 10(6), e0131720.
- Kasperkiewicz, P., Altman, Y., D’Angelo, M., Salvesen, G. S., & Drag, M. (2017). Toolbox of fluorescent probes for parallel imaging reveals uneven location of serine proteases in neu- trophils. Journal of the American Chemical Society, 139(29), 10115-10125.
- Ohneda, K., Ohmori, S. Y., & Yamamoto, M. (2019). Mousetryptase gene expression is coordinately regulated by GATA1 and GATA2 in bone marrow-derived mast cells. International journal of molecular sciences, 20(18), 4603.
- Trias, E., King, P. H., Si, Y., Kwon, Y., Varela, V., Ibarburu, S.,... & Barbeito, L. (2018). Mast cells and neutrophils mediate peripheral motor pathway degeneration in ALS. JCI insight, 3(19).
- Dell’Italia, L. J., Collawn, J. F., & Ferrario, C. M. (2018). Multifunctional role of chymase in acute and chronic tissue injury and remodeling. Circulation research, 122(2), 319-336.
- Pichler, R., Afkarian, M., Dieter, B. P., & Tuttle, K. R. (2017). Immunity and inflammation in diabetic kidney disease: translating mechanisms to biomarkers and treatment targets. American Journal of Physiology-Renal Physiology, 312(4), F716-F731.
- Pinto, C., Ninfole, E., Benedetti, A., Maroni, L., & Marzi- oni, M. (2020). Aging-related molecular pathways in chronic cholestatic conditions. Frontiers in Medicine, 332.
- Kosanovic, D., Luitel, H., Dahal, B. K., Cornitescu, T., Jans- sen, W., Danser, A. J., ... & Schermuly, R. T. (2015). Chy- mase: a multifunctional player in pulmonary hypertension associated with lung fibrosis. European Respiratory Journal, 46(4), 1084-1094.
- Chen, H., Xu, Y., Yang, G., Zhang, Q., Huang, X., Yu, L., & Dong, X. (2017). Mast cell chymase promotes hypertrophic scar fibroblast proliferation and collagen synthesis by acti- vating TGF β1/Smads signaling pathway. Experimental and therapeutic medicine, 14(5), 4438-4442.
- de Souza Junior, D. A., Santana, A. C., da Silva, E. Z. M., Oliver, C., & Jamur, M. C. (2015). The role of mast cell spe- cific chymases and tryptases in tumor angiogenesis. BioMed Research International, 2015.
- Tellechea, A., Bai, S., Dangwal, S., Theocharidis, G., Nagai, M., Koerner, S., ... & Veves, A. (2020). Topical application of a mast cell stabilizer improves impaired diabetic wound heal- ing. Journal of Investigative Dermatology, 140(4), 901-911.
- Aponte-López, A., Fuentes-Pananá, E. M., Cortes-Muñoz, D., & Muñoz-Cruz, S. (2018). Mast cell, the neglected member of the tumor microenvironment: role in breast cancer. Journal of immunology research, 2018.
- Adeolu, M., Alnajar, S., Naushad, S., & Gupta, R. S. (2016). Genome-based phylogeny and taxonomy of the ‘Enterobac- teriales’: proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pec- tobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. International journal of systematic and evolutionary mi- crobiology, 66(12), 5575-5599.
- Sayers, E. W., Agarwala, R., Bolton, E. E., Brister, J. R., Canese, K., Clark, K., ... & Ostell, J. (2019). Database re- sources of the national center for biotechnology information. Nucleic acids research, 47(Database issue), D23.
- Korkmaz, B., Caughey, G. H., Chapple, I., Gauthier, F., Hirschfeld, J., Jenne, D. E., ... & Thakker, N. S. (2018). Ther- apeutic targeting of cathepsin C: from pathophysiology to treatment. Pharmacology & therapeutics, 190, 202-236.
- Xu, Y. Y., Yao, L. X., & Shen, H. B. (2018). Bioimage-based protein subcellular location prediction: a comprehensive re- view. Frontiers of Computer Science, 12(1), 26-39.
- Zhao, L., Hanrahan, M. P., Chakravarty, P., DiPasquale, A. G., Sirois, L. E., Nagapudi, K., ... & Rossini, A. J. (2018). Charac- terization of pharmaceutical cocrystals and salts by dynamic nuclear polarization-enhanced solid-state NMR spectroscopy. Crystal Growth & Design, 18(4), 2588-2601.
- Elfiky, A. A. (2021). SARS-CoV-2 RNA dependent RNA polymerase (RdRp) targeting: An in silico perspective. Journal of Biomolecular Structure and Dynamics, 39(9), 3204-3212.
- Tang, Y., Huang, Y. J., Hopf, T. A., Sander, C., Marks, D. S., & Montelione, G. T. (2015). Protein structure determination by combining sparse NMR data with evolutionary couplings. Nature methods, 12(8), 751-754.
- Parker, K. R. (2013). Metabolic network construction based on the genome of the marine diatom Thalassiosira pseudonana and the analysis of genome-wide transcriptome data to inves- tigate triacylglyceride accumulation. San José State Univer- sity.
- Lundblad, R. L. (2004). Chemical reagents for protein modi- fication. CRC press.
- Perras, F. A., Wang, Z., Naik, P., Slowing, I. I., & Pruski, M. (2017). Natural abundance 17O DNP NMR provides precise O− H distances and insights into the Brønsted acidity of het- erogeneous catalysts. Angewandte Chemie, 129(31), 9293-9297.
- Gokulan, K., Khare, S., Cerniglia, C. E., Foley, S. L., & Var- ughese, K. I. (2018). Structure and inhibitor specificity of L, D-transpeptidase (LdtMt2) from Mycobacterium tuberculosis and antibiotic resistance: calcium binding promotes dimer formation. The AAPS journal, 20(2), 1-14.
- Niphakis, M. J., & Cravatt, B. F. (2014). Enzyme inhibitor discovery by activity-based protein profiling. Annual review of biochemistry, 83, 341-377.
- Kreyling, W. G., Abdelmonem, A. M., Ali, Z., Alves, F., Geis- er, M., Haberl, N., ... & Parak, W. J. (2015). In vivo integrity of polymer-coated gold nanoparticles. Nature nanotechnolo- gy, 10(7), 619-623.
- Veillard, F., Troxler, L., & Reichhart, J. M. (2016). Drosophi- la melanogaster clip-domain serine proteases: Structure, func- tion and regulation. Biochimie, 122, 255-269.
- Ghosh, A. K., & Brindisi, M. (2015). Organic carbamates in drug design and medicinal chemistry. Journal of medicinal chemistry, 58(7), 2895-2940.
- Webb, B., & Sali, A. (2016). Comparative protein structure modeling using MODELLER. Current protocols in bioinfor- matics, 54(1), 5-6.


