Research Article - (2025) Volume 10, Issue 1
Bibliometric Analysis of Breast Cancer and Circular RNA Research Trends from 2002 to 2022
Received Date: Dec 19, 2024 / Accepted Date: Jan 16, 2025 / Published Date: Jan 22, 2025
Copyright: ©©2024 Xiuli Zhang, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Zhong, S., Zhang, X., Li, Y., Zhang, M., Cao, H. (2025). Bibliometric Analysis of Breast Cancer and Circular RNA Research Trends from 2002 to 2022. Int J Cancer Res Ther, 10(1), 01-14.
Abstract
A bibliometric analysis aims to evaluate the research hotspots and emerging trends in breast cancer and circular RNA, offering new perspectives for researchers in future studies.Methods: This study utilizes bibliometric analysis software, such as VOSviewer and CiteSpace, to analyze literature on breast cancer and circular RNAs from 2002 to 2022, assessing bibliometric characteristics. Specifically, it includes a comprehensive evaluation of publication volume, country/region distribution, institutional sources, publishing journals, citation frequency, H-index, and research hotspots.Results:Through screening the Web of Science database, a total of 787 relevant publications were identified. Over time, the number of articles has shown a growing trend. In the field of breast cancer and circular RNA research, China holds a dominant position, with research institutions in the United States and Iran also being significant contributors. Fang Lin is the most prolific author, followed by Yang Burton B., Tang Hailin, Wang Xuehui, and Xie Xiaoming. "Frontiers in Oncology" and "Molecular Cancer" are the journals with the highest number of publications. The proliferation and migration of tumor cells remain hot research topics. Keywords have gradually shifted from "cell growth," "messenger RNAs," and "biomarker" to "long noncoding RNAs" and "sponge," indicating continuous updates and developments in this field. "Long noncoding RNAs" and "sponge" are likely to become future research hotspots.Conclusion: Bibliometric analysis provides valuable information for research on breast cancer and circular RNAs. In recent years, studies related to the mechanisms of breast cancer and the biological functions of circular RNAs have gained significant attention.
Keywords
Artificial Intelligence, Detection, Microorganisms, Microscopy
Introduction
According to the World Health Organization, in 2020, 2.26 million women worldwide were diagnosed with breast cancer, resulting in 685,000 deaths. By the end of 2020, 7.8 million women had been diagnosed with breast cancer in the past five years, making it the most common cancer type globally. It accounts for approximately 30% of all cancer cases and 15% of female cancer deaths. Despite significant advances in early diagnosis, surgery, chemotherapy, radiotherapy, endocrine therapy, and targeted therapy, breast cancer remains the leading cause of cancer-related deaths among women in over 100 countries [1-5].
Circular RNAs (circRNAs) are single-stranded RNA transcripts without 5' caps or 3' polyadenylated tails, generated through back-splicing from precursor mRNAs, and characterized by a covalently closed loop structure [6]. More than 40 years ago, viroids were first discovered as circular RNA molecules [7]. Subsequently, circular RNAs were observed in the cytoplasm of eukaryotic cell lines through electron microscopy [8]. At that time, these molecules were considered "junk" produced by aberrant splicing events, with only the testis-specific circular RNAs from the Sex-determining Region Y (Sry) gene in mouse testes thought to potentially have a function[9,10].In recent years, with the development of high-throughput sequencing technology (RNA-seq) and specific bioinformatics algorithms, thousands of circular RNAs have been identified in various species, including fungi, protozoa, plants, worms, fish, insects, and mammals [11-15]. Studies have found that certain circular RNAs exhibit cell- or tissue-specific expression in eukaryotes and display high stability and evolutionary conservation [16-18]. Evidence suggests that circular RNAs may play important roles in regulating gene expression through multiple mechanisms, such as acting as miRNA sponges, interacting with RNA-binding proteins, serving as intermediates in alternative splicing, templates for protein translation, and transcriptional regulators [19]. Recent research indicates that circular RNAs are involved in the pathological processes of many human diseases, especially cancer [20-23]. Currently, the mechanisms of circular RNAs in breast cancer are not well understood. However, research on breast cancer and circular RNAs has rapidly developed over the past decade.
Identifying new advancements and trends in the field has become challenging due to the rapid development of research on breast cancer and circular RNAs. Bibliometric analysis holds significant value in describing the current status, publication trends, and scientific output of researchers and institutions, as well as identifying future research hotspots [24]. Additionally, assessing the quality of relevant academic achievements is crucial for researchers, universities, and decision-makers in government departments, helping them to formulate strategies [25]. Bibliometric analysis provides a reliable basis for standardizing academic quality and offering useful information globally [26]. To date, this method has been used to estimate research trends in various medical fields, such as heart transplantation and familial hypercholesterolemia, as well as in non-medical fields [27-29].
Therefore, this study collected quantitative and qualitative data in the field of breast cancer and circular RNAs research, identifying the most productive countries, institutions, authors, and journals based on the number of publications or citations. The research objective is to conduct the first comprehensive bibliometric analysis of breast cancer and circular RNAs research over the past decade. This will provide scholars entering or about to enter the field with new insights into the current status, emerging trends, and future research hotspots in breast cancer and circular RNAs research.
Materials and Methods
Data Source and Search Strategy
On October 14, 2023, a literature search was conducted using the online version of the Web of Science Core Collection (WoSCC, Clarivate Analytics, Philadelphia, PA) database. The WoSCC database is considered one of the most comprehensive, systematic, and authoritative sources worldwide, covering over 12,000 influential journals and encompassing high-quality literature in biomedical, natural, and social sciences. It is widely used for bibliometric analysis and research on scientific literature visualization [30,31].
The search parameters were set as follows: Topic: [circular RNA] and Topic: [breast cancer], Time span: (2002 to 2022); Literature types: (articles and reviews); Language limit: English. All records (including title, author, keywords, source, abstract, and references) were exported in plain text format. The retrieval was conducted in one day (October 14, 2023) to avoid changes due to daily database updates.
Data Extraction and Analysis
All records retrieved from the WoSCC were independently downloaded by the author Sixiazhong and imported into Microsoft Excel 2016 for further data processing. Metrics such as the annual number of publications, countries/regions, source journals, institutions, authors, keywords, and citation frequencies were extracted and statistically analyzed. Additionally, the Impact Factor (IF) and classifications from the Journal Citation Reports (JCR) were used to evaluate the quality of scientific information. The IF represents the average number of citations per article in a journal over the most recent two-year period [32]. JCR assigns each scientific journal to the appropriate IF category and ranks them within specific fields (Q1, Q2, Q3, and Q4) [33]. The IF and JCR classifications used in this study are based on 2022 data. Furthermore, the H-index was applied as another metric. The H-index is defined as the number of h papers published by a researcher that have each been cited at least h times, with a higher H-index indicating greater academic influence. The H-index is often regarded as an important indicator for evaluating a researcher's scientific output and academic standing and is also useful for assessing the academic output capacity of countries, institutions, or journals [34].
Visualized Analysis
Using VOSviewer 1.6.19 (Centre for Science and Technology Studies, Leiden University, The Netherlands), we created a visualization map where nodes represent countries, institutions, authors, or keywords, connected through co-authorship, citation, co-citation, and co-occurrence analysis. The size of each node is determined by the weight of the element, such as the number of publications, citations, or frequency of occurrences. Each node was assigned a color, with the same color indicating the same cluster, representing a collection of items with similar attributes within the network [35]. The links between nodes represent associations between elements, and the thickness of the links indicates the strength of these associations. Total Link Strength (TLS) was used to quantitatively assess the links. Similar analytical methods were employed to create visualization maps of highly cited documents, countries or regions, institutions, journals, and authors across different periods to depict the evolution of these elements. Additionally, VOSviewer was used to categorize keywords with high co-occurrence frequencies into clusters and to create a density visualization map.
CiteSpace V (version 6.2.R4, Drexel University) was used to: (i) generate visual maps of the cited references network from the time zone view, and (ii) detect burst analysis of co-occurring keywords. The parameter settings for CiteSpace were as follows: time slice (2002-2022), with each slice being one year, item source (all selected), node type (one at a time), selection condition (50), pruning (Pathfinder), visualization (cluster view static, show merged networks). The detailed workflow of literature screening and data analysis is illustrated in
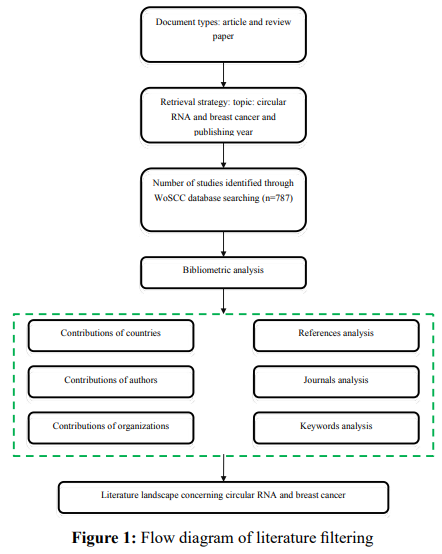
Results
Publication Outputs and Citation Trends
Based on the growth trend shown in Figure 2, the study period was divided into three phases: (i) 2002-2011: Initial Phase, with no more than three publications per year and almost no growth in citation rate; (ii) 2012-2015: Slow Growth Phase, with annual publications ranging between 2-6, maintaining a stable growth rate; and (iii) 2016-2022: Rapid Development Phase.
Figure 2: Global trend of annual publications and citations related to circular RNA and breast cancer research from 2002 to 2022. In 2020, the number of publications on circular RNAs and breast cancer exceeded 100 for the first time. By 2021, the annual number of publications surpassed 200, with a total of 6,807 citations. As of the retrieval date, these publications have been cited 27,966 times, with an average of 35.53 citations per paper. In the Initial Phase, research in this field was relatively limited and underdeveloped, with a low number of citations. However, from 2016 to 2022, the number of citations increased annually. Since 2019, annual citations have exceeded 2,000, showing a steady growth trend. This sustained growth reflects the increasing importance and attention given to circular RNAs and breast cancer research, indicating that this field is gaining more recognition within the academic community.
Contributions of Countries
Among the 787 papers, researchers hail from 52 countries or regions. From 2002 to 2022, the top ten countries in terms of global article output are shown in Figure 3A. The majority of papers originate from China (612 papers, accounting for 77.76%), followed by the United States (51 papers, 6.48%), Iran (48 papers, 6.10%), India (24 papers, 3.05%), and Italy (21 papers, 2.67%) (Table 1).
To ensure the reliability and statistical significance of the analysis, we set the VOSviewer parameters to include countries with at least five publications. A total of 52 countries were included, of which 21 met this threshold. The citation relationships between different countries/regions are shown in the citation network map (Figure 3B). Among the top ten countries in terms of citation count, China ranks highest (citations = 20,650), followed by the United States (citations = 3,975), Canada (citations = 1,283), Singapore (citations = 1,192), and Italy (citations = 1,039), with the latter three having comparable citation counts (Figure 3C). In terms of the number of cited documents, China ranks first (documents = 612), followed by the United States (documents = 51) and Iran (documents = 48) (Figure 3D). Among the top ten countries in terms of total link strength, China ranks highest (Total Link Strength, TLS = 2,207), followed by Iran (TLS = 620) and the United States (TLS = 522) (Figure 3E). China has the strongest connections with Iran and the United States, highlighting the leading positions of China and the United States in circular RNA and breast cancer research.
The international collaboration map shows the close cooperative relationships between countries/regions (Figure 3F). China is at the center of circular RNA and breast cancer research, with the highest total link strength (TLS = 52), a total of 612 publications, and 20,650 citations. China has particularly strong ties with the United States, Iran, and Singapore.
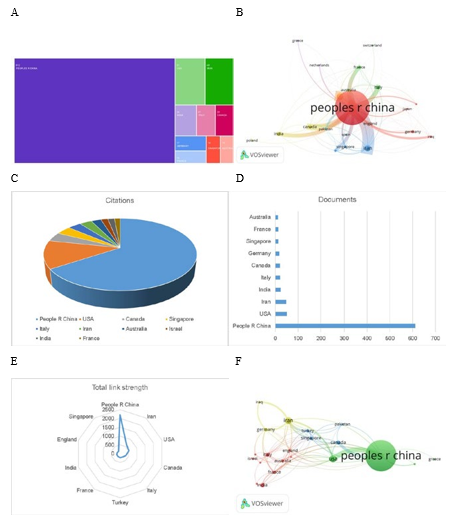
Figure 3: (A) The top 10 countries in terms of article output on circular RNA and breast cancer research from 2002 to 2022. (B) Citation network map showing the relationships between different countries/regions with at least five publications. (C) The top 10 countries by citation count. (D) The top 10 countries by the number of cited documents. (E) The top 10 countries by total link strength of citations. (F) Close cooperation relationships between countries/regions, with line thickness reflecting the frequency of cooperation. Thicker lines indicate stronger cooperation.
|
Countries |
Record count |
% of 787 |
|
people r china |
612 |
77.76% |
|
USA |
51 |
6.48% |
|
Iran |
48 |
6.10% |
|
India |
24 |
3.05% |
|
Italy |
21 |
2.67% |
|
Canada |
20 |
2.54% |
|
Germany |
17 |
2.16% |
|
France |
14 |
1.78% |
|
Singapore |
14 |
1.78% |
|
Australia |
13 |
1.65% |
Table 1: The top 10 countries by global article output in circular RNA and breast cancer research.
Contributions of Organizations
The 787 articles were published by 852 institutions. The top ten high-output institutions, all from China, are shown in Table 2. Nanjing Medical University (TP = 60) is the most prolific institution, followed by Sun Yat-sen University (TP = 42), Zhengzhou University (TP = 23), and Tongji University (TP = 21). Using VOSviewer, we created a network visualization of institutions that published more than five papers. Figure 4 displays the cooperation network among institutions, with 63 projects forming nine different colored clusters. In the ranking of the top ten institutions by total link strength (TLS) (Table 3), four institutions are from China: Nanjing Medical University (TLS = 53), Southeast University (TLS = 24), Sun Yat-sen University (TLS = 24), and Jilin University (TLS = 17). Nanjing Medical University has the highest total link strength (TLS = 53), followed by Southeast University (TLS = 24) and Sun Yat-sen University (TLS = 24), both with equal total link strength. Notably, Southeast University performs best in terms of the average number of citations per publication (CPP = 102.69), indicating its high impact in the field of circular RNA and breast cancer research.
Figure 4: Network visualization of institution collaboration analysis based on VOS viewer.
|
organization |
TP |
TC |
CPP |
TLS |
|
nanjing medical university |
60 |
2519 |
41.98 |
53 |
|
sun yat -sen university |
42 |
2941 |
70.02 |
24 |
|
zhengzhou university |
23 |
857 |
37.26 |
2 |
|
tongji university |
21 |
473 |
22.52 |
12 |
|
qingdao university |
19 |
296 |
15.58 |
14 |
|
jilin university |
18 |
1097 |
60.94 |
17 |
|
shangdong university |
18 |
728 |
40.44 |
9 |
|
huazhong university science and technology |
18 |
480 |
26.67 |
8 |
|
central south university |
18 |
460 |
25.56 |
5 |
|
China medical university |
18 |
230 |
12.78 |
1 |
CPP,Number of citations per publication;CPP = TC/TP;TC,Total number of citations of total publications;TP,Total number of publications;TLS,Total link strength
Table 2: The top 10 most productive institutions in circular RNA and breast cancer research.
|
Organization |
TP |
TC |
CPP |
TLS |
|
nanjing medical university |
60 |
2519 |
41.98 |
53 |
|
southeast university |
13 |
1335 |
102.69 |
24 |
|
sun yat-sen university |
42 |
2941 |
70.02 |
24 |
|
Shahid Beheshti University of Medical Sciences |
14 |
135 |
9.64 |
23 |
|
Islamic Azad University |
9 |
294 |
32.67 |
20 |
|
National University of Singapore |
13 |
1185 |
91.15 |
20 |
|
University of Tehran |
8 |
353 |
44.1 |
19 |
|
Hawler Medical University |
6 |
32 |
5.33 |
17 |
|
jilin University |
18 |
1097 |
60.94 |
17 |
|
University of Toronto |
12 |
1013 |
84.42 |
17 |
CPP,Number of citations per publication;CPP = TC/TP;TC,Total number of citations of total publications;TP,Total number of publications;TLS,Total link strength
Table 3: The top 10 institutions by total collaboration link strength in circular RNA and breast cancer research.
Authors and Co-cited Authors
A total of 4,038 authors contributed to the research output, with 3,454 authors having published only one article. To identify the most influential experts in the field of breast cancer and circular RNAs over the past twenty years, we ranked the top fifteen authors based on total publications (TP) and H-index. As shown in Table 4, Fang Lin from Anhui Agricultural University ranks first in terms of the number of papers (TP = 14, CPP = 17.71, H-index = 56), while Yang, Burton B. from the University of Toronto has the highest average citations per paper (TP = 11, CPP = 91.82, H-index = 58). Among the top fifteen most prolific authors, twelve are from China, two from the United States, and one from Turkey. Chen Bing, Li Yaming, Wang Lijuan, Yang Qifeng, and Zhao Wenjing have the highest total link strength (TLS = 63, TC = 570, TP = 8, CPP = 71.25), indicating a close collaborative relationship.
|
Author |
TP |
TC |
CPP |
TLS |
H-index |
Organization |
|
fang , lin |
14 |
248 |
17.71 |
34 |
56 |
Anhui Agricultural University |
|
yang , burton b. |
11 |
1010 |
91.82 |
48 |
58 |
University of Toronto |
|
tang , hailin |
11 |
940 |
85.45 |
26 |
52 |
Sun Yat Sen University |
|
wang , xuehui |
10 |
229 |
22.9 |
31 |
27 |
Sanya Tropical Fisheries Research Institute |
|
xie , xiaoming |
10 |
892 |
89.2 |
25 |
51 |
Beijing University of Chinese Medicine |
|
du , willian w. |
9 |
763 |
84.78 |
47 |
34 |
University of Toronto |
|
tang , jin-hai |
9 |
324 |
36 |
47 |
51 |
Dalian Medical University |
|
chen , bing |
8 |
570 |
71.25 |
63 |
23 |
Jiangsu Key Lab Mol & Translat Canc Res |
|
li , yaming |
8 |
570 |
71.25 |
63 |
39 |
Hosp China Med Univ 1 |
|
wang , lijuan |
8 |
570 |
71.25 |
63 |
28 |
Shandong University |
|
yang , qifeng |
8 |
570 |
71.25 |
63 |
87 |
Zhejiang University |
|
zhao , wenjing |
8 |
570 |
71.25 |
63 |
19 |
Southern University of Science & Technology |
|
wu , nan |
8 |
492 |
61.5 |
41 |
59 |
Binzhou Medical University |
|
ji , changle |
8 |
96 |
12 |
29 |
11 |
Tongji University |
|
zarrabi , ali |
8 |
384 |
48 |
18 |
52 |
Istinye University |
CPP,Number of citations per publication;CPP = TC/TP;TC,Total number of citations of total publications;TP,Total number of publications;TLS,Total link strength
Table 4: The top 15 most productive authors on circular RNA and breast cancer.
Using VOSviewer, we set the minimum number of published papers threshold to three and found that the most prolific authors also have tight collaboration networks (Figure 5A). Set the minimum citation threshold to 40, creating a co-citation network of authors that includes 126 nodes, 7,737 links, and 4 clusters. (Figure 5B).
Figure 5(A): Network visualization of authors collaboration analysis based on VOSviewer. (B) Network visualization of co-cited authors analysis based on VOSviewer.
Journals and Co-cited Journals
The 787 articles on circular RNAs and breast cancer were published in 284 academic journals. The top ten high-output journals published a total of 191 papers, accounting for approximately 24% of the total output (Table 5). "Frontiers in Oncology" and "Molecular Cancer" lead in total output (TP), but "Molecular Cancer" surpasses "Frontiers in Oncology" in total citations (TC = 2,877), average citations (AC = 106.56), H-index (H-index = 126), and impact factor (IF = 37.3), indicating its significant status in the field of circular RNAs and breast cancer.
|
Rank |
Journal |
TP |
TC |
AC |
H-index |
JCR |
IF2022 |
|
1 |
frontiers in oncology |
27 |
298 |
11.037 |
83 |
Q2 |
4.7 |
|
2 |
molecular cancer |
27 |
2877 |
106.5556 |
126 |
Q1 |
37.3 |
|
3 |
cancer management and research |
20 |
440 |
22 |
40 |
Q3 |
3.3 |
|
4 |
cancer cell international |
17 |
238 |
14 |
56 |
Q2 |
5.8 |
|
5 |
molecular therapy-nucleic acids |
16 |
588 |
36.75 |
59 |
Q1 |
8.8 |
|
6 |
cell death & disease |
15 |
926 |
61.7333 |
111 |
Q1 |
9 |
|
7 |
international journal of molecular sciences |
15 |
1174 |
78.2667 |
162 |
Q1 |
6.2 |
|
8 |
oncotargets and therapy |
15 |
245 |
16.3333 |
60 |
Q2 |
4 |
|
9 |
cancers |
13 |
170 |
13.0769 |
76 |
Q1 |
5.2 |
|
10 |
frontiers in cell and developmental biology |
13 |
252 |
19.3846 |
53 |
Q1 |
5.5 |
IF,Impact Factor; JCR,Journal Citation Reports; TC,Total number of citations of total publications;TP,Total number of publications;AC=TC/TP,Average citations.
Table 5: The top 11 most productive journals on circular RNA and breast cancer.
Based on network analysis, the citation patterns of 61 journals were divided into seven clusters, forming a total of 801 links (Figure 6A). The co-citation relationships between different journals are shown in Figure 6B, which includes 232 items forming five clusters. "Molecular Cancer" has the highest co-citation count at 1,894 times, followed by "Nature" (1,491 times), "Cell" (1,362 times), and "Oncotarget" (1,329 times).
Figure 6: (A) Network visualization of journals analysis based on VOSviewer.(B) Network visualization of co-cited journals analysis based on VOSviewer.
References and Co-cited References
To further explore development trends, we analyzed the ten most-cited documents during the study period (Table 6). These documents include one review and nine research papers, with total citation counts ranging from 103 to 266. The most-cited article is a 2013 study by Thomas B. Hansen (266 citations), followed by a 2013 study by Sebastian Memczak (261 citations). Both articles discuss how circular RNAs mediate protein translation expression by regulating the activity of miR-7, revealing previously unrecognized regulatory potential of coding sequences. Overall, newly published articles have lower citation rates, so the current citation counts may underestimate their actual value.
|
Rank |
Title |
Year |
Journal |
First author |
TC |
|
1 |
Natural RNA circles function as efficient microRNA sponges |
2013 |
Nature |
Hansen, TB |
266 |
|
2 |
Circular RNAs are a large class of animal RNAs with regulatory potency |
2013 |
Nature |
Memczak, S |
261 |
|
3 |
Circular RNAs are abundant, conserved, and associated with ALU repeats |
2013 |
RNA |
Jeck, WR |
176 |
|
4 |
Exon-intron circular RNAs regulate transcription in the nucleus |
2015 |
Nature Structural& Molecular Biology |
Li, Zhaoyong |
126 |
|
5 |
Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries |
2018 |
Ca-Cancer J Clin |
Freddie Bray BSc |
123 |
|
6 |
circRNA Biogenesis Competes with Pre-mRNA Splicing |
2014 |
Molecular Cell |
Ashwal-Fluss, R |
114 |
|
7 |
Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2 |
2016 |
Nucleic Acids Research |
Du, William W. |
112 |
|
8 |
Circular Intronic Long Noncoding RNAs |
2013 |
Molecular Cell |
Zhang, Yang |
106 |
|
9 |
Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs |
2016 |
Nature Communications |
Zheng, Qiupeng |
106 |
|
10 |
Detecting and characterizing circular RNAs |
2014 |
Nature Biotechnology |
Jeck, William R. |
103 |
TC,Total number of citations of total publications
Table 6: The top 10 references on circularRNA and breast cancer with the most co-citations.
Figure 7 presents a co-citation analysis based on CiteSpace, showing 172 documents cited more than five times annually in the field of breast cancer and circular RNAs. Visualizing these documents in a timeline analysis reflects the temporal characteristics of research hotspots in this field. Based on the log-likelihood ratio (LLR) algorithm, nine cluster labels representing hot topics were selected (Table 7) and ranked in Figure 7. Along the dashed lines connecting each label, larger circles indicate higher citation frequencies, and warmer-colored lines indicate later publication dates.
Figure 7: Timeline view of the top 13 most co-cited references on circular RNA and breast cancer using CiteSpace software. For each cluster, the position of each node indicates the publication time of the document, and the node size represents the number of citations.
|
Cluster ID |
Size |
Silhouette |
Mean(Year) |
Label (LLR) |
|
0 |
20 |
0.99 |
2006 |
breast cancer |
|
1 |
17 |
0.992 |
2008 |
mir-326 |
|
2 |
17 |
0.986 |
2017 |
lung cancer |
|
3 |
14 |
1 |
2019 |
cancer therapy |
|
4 |
13 |
0.846 |
2019 |
tumor suppressive |
|
5 |
13 |
0.974 |
2004 |
gene expression |
|
6 |
12 |
1 |
2018 |
abundant |
|
7 |
11 |
0.982 |
2019 |
cell |
|
8 |
11 |
0.857 |
2016 |
noncoding rnas |
LLR: Loglikelihood ratio
Table 7: Main clusters of co-cited references
Cluster 0 (breast cancer) and Cluster 5 (gene expression) appeared the earliest, indicating that early breast cancer research focused on gene expression. Cluster 3 (cancer therapy) and Cluster 4 (tumor suppression) are current research hot topics, reflecting the high current interest in tumor suppressor genes and cancer therapy.
Keywords Analysis of Research Hotspots
The co-occurrence frequency of keywords can indicate scientific hotspots within a specific research theme, serving as an important method for tracking scientific development. This study extracted 2,909 keywords from 787 articles and listed the top 20 keywords based on co-occurrence frequency using VOSviewer software, as shown in Table 8.
First, a density map was generated using 72 keywords with co-occurrence frequencies exceeding 20 times, visually depicting the distribution of popular themes in the field. As shown in Figure 8A, "circular RNA," "breast cancer," "expression," "proliferation," "invasion," and "metastasis" are significant elements of the density map. Next, a cluster analysis was performed on the selected keywords. Each cluster was labeled with index terms extracted from the keywords. After calculation, the 72 high-frequency keywords formed four clusters (Figure 8B), representing the main research directions in this field: Cluster 1 (red circles, 31 keywords): Focuses on various diseases related to breast cancer and circular RNAs, with keywords including "cell lung-cancer," "colorectal-cancer," and "gastric-cancer." Cluster 2 (green circles, 23 keywords): Focuses on the clinical diagnosis of breast cancer and circular RNAs, with keywords including "diagnosis," "resistance," and "prognosis."
|
Keyword |
Occurrences |
TLS |
|
circular rna |
280 |
1480 |
|
breast cancer |
255 |
1240 |
|
breast-cancer |
241 |
1231 |
|
expression |
189 |
1074 |
|
proliferation |
188 |
1095 |
|
circular rnas |
169 |
861 |
|
progression |
147 |
873 |
|
metastasis |
145 |
935 |
|
invasion |
143 |
879 |
|
migration |
117 |
726 |
|
cells |
94 |
484 |
|
circrna |
92 |
587 |
|
biomarker |
89 |
533 |
|
cancer |
78 |
475 |
|
growth |
74 |
452 |
|
promotes |
69 |
438 |
|
hepatocellular-carcinoma |
64 |
391 |
|
circrnas |
62 |
396 |
|
apoptosis |
61 |
340 |
|
colorectal-cancer |
60 |
376 |
TLS:TLS,Total link strength
Table 8: The top 20 keywords in the co-occurrence frequency of circular RNA and breast cancer.
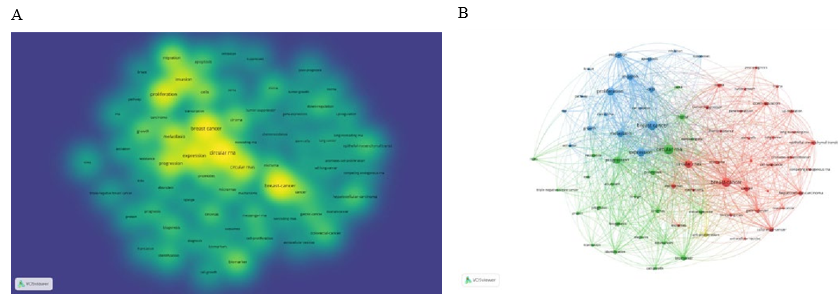
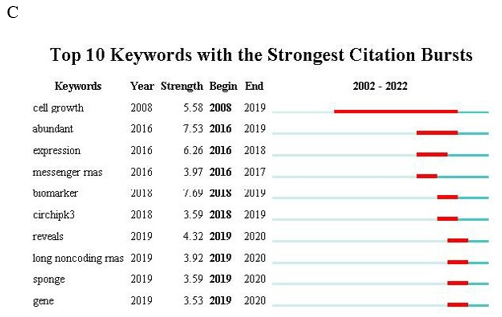
Figure 8: (A) Density map of keywords generated by VOSviewer. (B) Keyword co-occurrence analysis on breast cancer and circular RNA research using VOSviewer. (C) Top 10 keywords with the strongest citation bursts (sorted by the beginning year of the burst). Notes: The blue bars indicate the reference had been published; the red bars indicate citation bursts.
Cluster 3 (blue circles, 17 keywords): Focuses on the mechanisms of action of breast cancer and circular RNAs, with keywords including "activation," "apoptosis," "invasion," and "proliferation." Cluster 4 (yellow circle, 1 keyword): Labeled as "cell-proliferation." Finally, we identified the top 10 potential keywords, which are considered indicators of research frontiers or emerging trends over time. As shown in Figure 8C, since 2018, the keywords with citation bursts include "biomarker" (7.69), "circhipk3" (3.59), "long noncoding RNAs" (3.92), and "sponge" (3.59). These keywords represent the research frontiers in the study of breast cancer and circular RNAs.
Discussions
We conducted a bibliometric analysis of 787 documents related to breast cancer and circular RNAs from the WoSCC database, covering the period from 2002 to 2022. This analysis revealed several key insights. Firstly, this has been an active research field in recent years. From 2002 to 2015, there was almost no growth in the number of research papers on breast cancer and circular RNAs globally. However, since 2016, there has been an explosive increase in the number of research papers in this field. Through bibliometric analysis of countries, institutions, authors, and journals, we observed changes in global research trends.
China has the highest output of publications in this field, followed by the United States and Iran. All top ten high-output institutions are from China, with most collaborative research conducted domestically. European countries have relatively close collaborations, while cooperation among other countries is limited. Notably, Nanjing Medical University, Sun Yat-sen University, and Southeast University have close ties. Strengthening international cooperation, removing academic barriers, and promoting the development of breast cancer and circular RNA research is crucial.
In the co-author analysis, Fang Lin from Anhui Agricultural University has published the most papers (TP=14, CPP=17.71, H-index=56), followed by Yang, Burton B. from the University of Toronto (TP=11, CPP=91.82, H-index=58), and Tang Hailin from Sun Yat-sen University (TP=11, CPP=85.45, H-index=52), who have played significant roles in this field. Additionally, Yang, Burton B. from the University of Toronto stands out in terms of single article citations (TP=11, CPP=91.82, H-index=58), contributing many high-quality articles to this field.
Among the top ten journals, except for "Frontiers in Oncology" (impact factor 4.7), "Cancer Management and Research" (impact factor 3.3), and "OncoTargets and Therapy" (impact factor 4), the remaining journals have relatively high impact factors (greater than 5.0). "Frontiers in Oncology" and "Molecular Cancer" tie for the first place in the number of papers (TP), but "Molecular Cancer" surpasses "Frontiers in Oncology" in total citations (TC=2877), average citations (AC=106.56), H-index (H-index=126), and impact factor (IF=37.3). Furthermore, "Molecular Cancer" has the highest co-citation count at 1,894 times, indicating its significant status in the field of circular RNAs and breast cancer. According to the 2022 JCR standards, among the top ten most active journals, six are classified as Q1, three as Q2, and one as Q3.
The downregulation of circular RNA gene expression promotes the proliferation, invasion, and migration of non-small cell lung cancer (NSCLC) cells by regulating protein expression [36-38]. Circular RNAs also play significant roles in various cancers. In breast cancer, they promote the development of multiple myeloma through the miR-1180/yes-associated protein pathway and facilitate the progression of Mantle Cell Lymphoma via pathways involving three lncRNAs (MALAT1, NEAT1, and XIST), five microRNAs (hsa-miR-129-5p, hsa-miR-3163, hsa-miR-4662a-5p, hsa-miR-101-3p, and hsa-miR-186-5p), and five mRNAs (NOTCH1, FMR1, ABCB1, TWIST1, and VEGFA) [39,40]. Additionally, circular RNAs reduce the proliferation, invasion, and migration of Wilms tumor cells through the miR-145-5p pathway, inhibit the growth and migration of bladder cancer cells through the C-MYC pathway, and suppress the growth and migration of colon cancer cells through pathways involving miR-150-5p, c-Myc, cyclin D1, p53, caspase-3, PARP, PTEN, PI3K, AKT, JAK2, and STAT5 [41-43]. Circular RNAs also inhibit the growth of hepatocellular carcinoma through pathways involving miR-892a and miR-328-3p, HIF1AN, HDGF, PI3K-AKT, the Rapamycin kinase complex 1/β-catenin, NOTCH2, BIRC5, SURVIVIN, and MYC [44].
Circular RNAs regulate carcinogenesis and cancer progression by acting as microRNA sponges, encoding proteins, and interacting with proteins [45]. In breast cancer, researchers detected 3,093 circular RNAs, with 18 exhibiting differential expression between adriamycin (ADM)-resistant and parental MCF-7 cells. Through quantitative real-time polymerase chain reaction (qPCR) analysis, they found that Homo sapiens (hsa)_circ_0006528 was significantly upregulated in ADM-resistant cancer cells. When the expression of hsa_circ_0006528 was inhibited by small interfering RNA, the sensitivity of breast cancer cells to ADM significantly increased. Concurrently, the downregulation of hsa_circ_0006528 was associated with increased expression of miRNA-7-5p. This suggests that the hsa_circ_0006528/miR-7-5p/RAF1 axis plays a regulatory role in ADM-resistant breast cancer [46]. Additionally, circABCB10 has been reported to promote breast cancer proliferation and inhibit apoptosis by acting as a sponge for miRNA-1271 [47]. Studies have also found that the miR-1275-ATG7/ULK1-autophagic axis and miR-92b-3p pathway promote autophagy and proliferation of breast cancer cells, while the miR-190a-3p/TP53INP1 axis pathway induces apoptosis in triple-negative breast cancer cells [22,48,49].
To our knowledge, this is the first bibliometric analysis of the literature on breast cancer and circular RNAs, describing milestones and trends over the past twenty years. The advantages of this analysis lie in its objectivity and its ability to demonstrate development directions in the field while highlighting areas that have not been thoroughly studied. However, several limitations exist. Firstly, our data is solely from the WoSCC database, excluding other databases (such as PubMed), printed publications, and video materials, which may result in the omission of certain articles. Secondly, our selective analysis of the characteristics of the information might overlook some key points and details. Lastly, bibliometric methods have inherent limitations and cannot correct for factors such as population size, overall research output, gross domestic product, and research investment resources. Nevertheless, we have attempted to comprehensively analyze the development and research content of breast cancer and circular RNAs, aiming to summarize the discipline and provide guidance during this period of rapid development in this field.
Conclusion
In summary, research on the mechanisms of breast cancer related to the biological functions of circular RNAs is increasing, with "Molecular Cancer" being the most influential journal in this field. China leads in both output and scientific impact. Among institutions, Nanjing Medical University is the top producer and also has the highest total link strength. Fang Lin from Anhui Agricultural University ranks first in the number of papers, while Yang, Burton B. from the University of Toronto has the highest number of citations per paper. The research hotspots in this field focus on "expression," "proliferation," "invasion," and "metastasis." The research is primarily based on basic studies, concentrating on animal and cellular levels. Future research directions include "biomarker," "circhipk3," "long noncoding RNAs," and "sponge."
References
- Farkas, A. H., &Nattinger, A. B. (2023). Breast cancer screening and prevention. Annals of Internal Medicine, 176(11), ITC161-ITC176.
- Brauer, E. R., & Ganz, P. A. (2022). Moving the translational needle in breast cancer survivorship: connecting intervention research to clinical practice. Journal of Clinical Oncology, 40(19), 2069.
- Miller, K. D., Nogueira, L., Devasia, T., Mariotto, A. B., Yabroff, K. R., Jemal, A., ... & Siegel, R. L. (2022). Cancer treatment and survivorship statistics, 2022. CA: a cancer journal for clinicians, 72(5), 409-436.
- Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R. L., Torre,L. A., & Jemal, A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians, 68(6), 394-424.
- Harbeck, N., &Gnant, M. (2017). Interpretation of the evidence for the efficacy and safety of statin therapy. Lancet, 389(10074), 1134-1150.
- Kristensen, L. S., Andersen, M. S., Stagsted, L. V., Ebbesen,K. K., Hansen, T. B., &Kjems, J. (2019). The biogenesis, biology and characterization of circular RNAs. Nature reviews genetics, 20(11), 675-691.
- Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J., & Kleinschmidt, A. K. (1976). Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proceedings of the National Academy of Sciences, 73(11), 3852-3856.
- Hsu, M. T., & Coca-Prados, M. (1979). Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature, 280(5720), 339-340.
- Cocquerelle, C., Mascrez, B., Hétuin, D., &Bailleul, B. (1993). Mis-splicing yields circular RNA molecules. The FASEB Journal, 7(1), 155-160.
- Capel, B., Swain, A., Nicolis, S., Hacker, A., Walter, M., Koopman, P., ... & Lovell-Badge, R. (1993). Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell, 73(5), 1019-1030.
- Wang, P. L., Bao, Y., Yee, M. C., Barrett, S. P., Hogan, G. J., Olsen, M. N., ... & Salzman, J. (2014). Circular RNA is expressed across the eukaryotic tree of life. PloS one, 9(3), e90859.
- Ivanov, A., Memczak, S., Wyler, E., Torti, F., Porath, H. T., Orejuela, M. R., ... &Rajewsky, N. (2015). Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell reports, 10(2), 170-177.
- Jeck, W. R., Sorrentino, J. A., Wang, K., Slevin, M. K., Burd,C. E., Liu, J., ... & Sharpless, N. E. (2013). Circular RNAs are abundant, conserved, and associated with ALU repeats. Rna, 19(2), 141-157.
- Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L., & Brown,P. O. (2013). Cell-type specific features of circular RNA expression. PLoS genetics, 9(9), e1003777.
- Westholm, J. O., Miura, P., Olson, S., Shenker, S., Joseph, B., Sanfilippo, P., ... & Lai, E. C. (2014). Genome-wide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell reports, 9(5), 1966-1980.
- Maass, P. G., Glazar, P., Memczak, S., Dittmar, G., Hollfinger, I., Schreyer, L., ... &Rajewsky, N. (2017). A map of human circular RNAs in clinically relevant tissues. Journal of molecular medicine, 95, 1179-1189.
- Xia, S., Feng, J., Lei, L., Hu, J., Xia, L., Wang, J., ... & He,C. (2017). Comprehensive characterization of tissue-specific circular RNAs in the human and mouse genomes. Briefings in bioinformatics, 18(6), 984-992.
- Li, X., Yang, L., & Chen, L. L. (2018). The biogenesis, functions, and challenges of circular RNAs. Molecular cell, 71(3), 428-442.
- Wang, S., Zhang, Y., Cai, Q., Ma, M., Jin, L. Y., Weng, M.,... & Quan, Z. (2019). Circular RNA FOXP1 promotes tumor progression and Warburg effect in gallbladder cancer by regulating PKLR expression. Molecular cancer, 18, 1-15.
- Zhang, Y., Liang, W., Zhang, P., Chen, J., Qian, H., Zhang, X., & Xu, W. (2017). Circular RNAs: emerging cancer biomarkers and targets. Journal of Experimental & Clinical Cancer Research, 36, 1-13.
- Zheng, X., Huang, M., Xing, L., Yang, R., Wang, X., Jiang, R., ... & Chen, J. (2020). The circRNA circSEPT9 mediated by E2F1 and EIF4A3 facilitates the carcinogenesis and development of triple-negative breast cancer. Molecular cancer, 19, 1-22.
- Liang, G., Ling, Y., Mehrpour, M., Saw, P. E., Liu, Z., Tan, W., ... & Gong, C. (2020). Autophagy-associated circRNAcircCDYL augments autophagy and promotes breast cancer progression. Molecular cancer, 19, 1-16.
- Sang, Y., Chen, B., Song, X., Li, Y., Liang, Y., Han, D., ... & Yang, Q. (2019). circRNA_0025202 regulates tamoxifen sensitivity and tumor progression via regulating the miR-182-5p/FOXO3a axis in breast cancer. Molecular Therapy, 27(9), 1638-1652.
- King, D. A. (2004). The scientific impact of nations. Nature, 430(6997), 311-316.
- Hirsch, J. E. (2005). An index to quantify an individual's scientific research output. Proceedings of the National academy of Sciences, 102(46), 16569-16572.
- Adnan, S., & Ullah, R. (2018). Top-cited articles in regenerative endodontics: a bibliometric analysis. Journal of Endodontics, 44(11), 1650-1664.
- Du, Y., Duan, C., Yang, Y., Yuan, G., Zhou, Y., Zhu, X., ...& Hu, Y. (2022). Heart transplantation: a bibliometric review from 1990-2021. Current Problems in Cardiology, 47(8),101176.
- Wei, N., Hu, Y., Liu, G., Li, S., Yuan, G., Shou, X., ...&Zhai, H. (2023). A bibliometric analysis of familial hypercholesterolemia from 2011 to 2021. Current problems in cardiology, 48(7), 101151.
- Tseng, M. L., Chang, C. H., Lin, C. W. R., Wu, K. J., Chen,Q., Xia, L., &Xue, B. (2020). Future trends and guidance for the triple bottom line and sustainability: A data driven bibliometric analysis. Environmental Science and Pollution Research, 27, 33543-33567.
- Romero, L., & Portillo-Salido, E. (2019). Trends in sigma-1 receptor research: a 25-year bibliometric analysis. Frontiers in pharmacology, 10, 564.
- Chen, Y., Li, Y., Guo, L., Hong, J., Zhao, W., Hu, X., ... &Xiong,K. (2021). Bibliometric analysis of the inflammasome and pyroptosis in brain. Frontiers in pharmacology, 11, 626502.
- Kamath, P. S., & Bologna, G. (2009). Impact factor: misused and overhyped?. Hepatology, 49(6), 1787-1789.
- Wu, X. F., Fu, Q., & Rousseau, R. (2008). On indexing in the Web of Science and predicting journal impact factor. Journal of Zhejiang University SCIENCE B, 9, 582-590.
- Engqvist, L., &Frommen, J. G. (2008). The h-index and self-citations. Trends in ecology & evolution, 23(5), 250-252.
- Leydesdorff, L., Carley, S., & Rafols, I. (2013). Global maps of science based on the new Web-of-Science categories. Scientometrics, 94, 589-593.
- Chen, D., Zhou, H., Cai, Z., Cai, K., Liu, J., Wang, W., ... & Wen, Z. (2022). CircSCAP interacts with SF3A3 to inhibit the malignance of non-small cell lung cancer by activating p53 signaling. Journal of Experimental & Clinical Cancer Research, 41(1), 120.
- Huang, M. S., Liu, J. Y., Xia, X. B., Liu, Y. Z., Li, X., Yin,J. Y., ... & Liu, Z. Q. (2019). Hsa_circ_0001946 inhibits lung cancer progression and mediates cisplatin sensitivity in non-small cell lung cancer via the nucleotide excision repair signaling pathway. Frontiers in oncology, 9, 508.
- Qin, C., Lu, R., Yuan, M., Zhao, R., Zhou, H., Fan, X.,... &Bian, T. (2021). Circular RNA 0006349 augments glycolysis and malignance of non-small cell lung cancer cells through the microRNA-98/MKP1 axis. Frontiers in Cell and Developmental Biology, 9, 690307.
- Chen, F., Wang, X., Fu, S., Wang, S., Fu, Y., Zhang, J., & Liu, Z. (2020). Circular RNA circ-CDYL sponges miR-1180 to elevate yes-associated protein in multiple myeloma. Experimental Biology and Medicine, 245(11), 925-932.
- Mei, M., Wang, Y., Wang, Q., Liu, Y., Song, W., & Zhang,M. (2019). CircCDYL serves as a new biomarker in mantle cell lymphoma and promotes cell proliferation. Cancer Management and Research, 10215-10221.
- Zhou, R., Jia, W., Gao, X., Deng, F., Fu, K., Zhao, T., ... &Liu, G. (2021). CircCDYL acts as a tumor suppressor in Wilms’ tumor by targeting miR-145-5p. Frontiers in cell and developmental biology, 9, 668947.
- Sun, J., Zhang, H., Tao, D., Xie, F., Liu, F., Gu, C., ... & Xiao,X. (2019). CircCDYL inhibits the expression of C-MYC to suppress cell growth and migration in bladder cancer. ArtificialCells, Nanomedicine, and Biotechnology, 47(1), 1349-1356.
- Cui, W., Dai, J., Ma, J., & Gu, H. (2019). circCDYL/microRNA-150-5p participates in modulating growth and migration of colon cancer cells. General Physiology & Biophysics, 38(6).
- Wei, Y., Chen, X., Liang, C., Ling, Y., Yang, X., Ye, X., ... & Wang, H. (2020). A noncoding regulatory RNAs network driven by Circ-CDYL acts specifically in the early stages hepatocellular carcinoma. Hepatology, 71(1), 130-147.
- Su, Y., Zhong, G., Jiang, N., Huang, M., & Lin, T. (2018). Circular RNA, a novel marker for cancer determination. International Journal of Molecular Medicine, 42(4), 1786-1798.
- Gao, D., Zhang, X., Liu, B., Meng, D., Fang, K., Guo, Z., & Li,L. (2017). Screening circular RNA related to chemotherapeutic resistance in breast cancer. Epigenomics, 9(9), 1175-1188.
- Liang, H. F., Zhang, X. Z., Liu, B. G., Jia, G. T., & Li, W. L.(2017). Circular RNA circ-ABCB10 promotes breast cancer proliferation and progression through sponging miR-1271. American journal of cancer research, 7(7), 1566.
- Liang, G., Ling, Y., Lin, Q., Shi, Y., Luo, Q., Cen, Y., ... & Gong, C. (2021). MiR-92b-3p inhibits proliferation of HER2-positive breast cancer cell by targeting circCDYL. Frontiers in Cell and Developmental Biology, 9, 707049.
- Wang, S., Liu, F., Ma, H., Cui, X., Yang, S., & Qin, R. (2020).circCDYL acts as a tumor suppressor in triple negative breast cancer by sponging miR-190a-3p and upregulating TP53INP1. Clinical Breast Cancer, 20(5), 422-430.