Research Article - (2025) Volume 8, Issue 1
7-Dehydrocholesterol Reductase is a Promising Prognostic and Immune Biomarker in Pan-Cancer
2Department of Laboratory Medicine, The First Afiliated Hospital of USTC, China
3Department of Hematology, The Second Afiliated Hospital, China
4Cancer Institute, Zhejiang University, China
Received Date: Jun 05, 2024 / Accepted Date: Jan 24, 2025 / Published Date: Feb 07, 2025
Copyright: ©© 2025 Dan Lin, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Wu, X., Zheng, W., Wang, L., Zhang, X., Lin, D., et al. (2025). 7-Dehydrocholesterol Reductase is a Promising Prognostic and Immune Biomarker in Pan-Cancer. Adv Hema Onco Res, 8(1), 01-15.
Abstract
Background: 7-Dehydrocholesterol Reductase (DHCR7) is a key enzyme involving the final step of cholesterol synthesis path- way. Abnormalities in the DHCR7 gene can lead to a variety of diseases, such as Smith-Lemli-Opitz syndrome. However, the relationship between DHCR7 and oncogenesis remain unclear.
Methods: We used several bioinformatic databases which the original data from the TCGA and GEO database. Briefly, the gene of DHCR7 expression were explored by the Oncomine, TIMER and GEPIA databases. The effect of DHCR7 on prognosis was analyzed via Kaplan-Meier plotter and GEPIA database. The TISIDB database was used to determine the relationship between DHCR7 expression and pan-cancer stages and the DHCR7 expression in different immune and molecular subtypes of human cancers. The correlations between DHCR7 expression and Immune Checkpoints (ICP), Tumor Mutational Burden (TMB), Mi- crosatellite Instability (MSI), Mutant-Allele Tumor Heterogeneity (MATH), neoantigens and infiltrating immune cells of human cancers and urogenital cancers were analyzed through the SangerBox database (http://vip.sangerbox.com/login.html). The genomic alterations of DHCR7 were analyzed by the c-BioPortal database. The differential expression of DHCR7 in urogenital cancers with different clinical characteristics was analyzed with the UALCAN database. The DHCR7 co-expression genes in BLCA was analyzed through the Linked Omics database. And the association between DHCR7 and related genes and markers of immune cells were analyzed by TIMER database.
Results: The results indicated that DHCR7 was highly expressed in most cancers, except in Cholangio carcinoma, Pheochromo- cytoma and Paraganglioma. Aberrantly expressed DHCR7 was associated with the poor prognosis, advanced tumor stage and metastasis in most tumor types. Additionally, significant strong correlations between DHCR7 expression and tumor immune-in- filtrated cells (TILs), ICP, TMB, MSI, MATH and neoantigens showed in most human cancers, and marker genes of TILs were significantly related to DHCR7 expression in BLCA, KIRC and PRAD. DHCR7 co-expression networks mostly participated in the regulation of immune response regulating signaling pathway, leukocyte differentiation and angiogenesis.
Conclusion: Through pan-cancer analysis, DHCR7 may serve as a potential prognostic and immunological pan-cancer bio- marker, especially in urological tumors.
Keywords
DHCR7, Pan-Cancer, Biomarker, Prognosis, Immunotherapy
List Of Abbreviations
DHCR7: 7-Dehydrocholesterol Reductase
BLCA: Bladder Urothelial Carcinoma
BRCA: Breast Invasive Carcinoma
CESC: Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma
COAD: Colon Adenocarcinoma
ESCA: Esophageal Carcinoma
HNSC: Head and Neck Cancer
KICH: Kidney Chromophobe
LIHC: Liver Hepatocellular Carcinoma
LUAD: Lung Adenocarcinoma
PAAD: Pancreatic Adenocarcinoma
SARC: Sarcoma
LUSC: Lung Squamous Cell Carcinoma
READ: Rectum Adenocarcinoma
STAD: Stomach Adenocarcinoma
THCA: Thyroid Carcinoma
UCEC: Uterine Corpus Endometrial Carcinoma
UVM: Uveal Melanoma
ACC: Adrenocortical Carcinoma
CHOL: Cholangiocarcinoma
KIRC: Kidney Renal Clear Cell Carcinoma
KIRP: Kidney Renal Papillary Carcinoma
TGCT: Testicular Germ Cell Tumors
LGG: Brain Lower Grade Glioma
PCPG: Pheochromocytoma and Paraganglioma
PRAD: Prostate Adenocarcinoma
ICP: Immune Checkpoints
TMB: Tumor Mutational Burden
MSI: Microsatellite Instability
MATH: Mutant-Allele Tumor Heterogeneity
TILS: Tumor-Infiltrating Lymphocytes
TME: Tumor Microenvironment
Introduction
DHCR7 encodes the 7-Dehydrocholesterol Reductase (DHCR7) enzyme, which is the terminal enzyme in the cholesterol biosynthesis pathway [1]. It catalyzes the conversion of 7-dehydrocholesterol (7-DHC) to cholesterol, which is the final step in the biogenesis of cholesterol [2]. Mutations in the DHCR7 gene cause Smith-Lemli- Opitz Syndrome (SLOS), a clinical autosomal recessive genetic disease characterized by multiple abnormalities including growth deficiency, intellectual disability, and frequent infections [3]. DHCR7 has been also found to play roles in cancers. For example, highly expression of DHCR7 was associated with poor prognosis in cervical cancer, gastric cancer and bladder cancer [4-6]. In gastric cancer, researchers found that DHCR7 regulated cell proliferation, migration, and invasion as well as apoptosis resistance via cellular cholesterol metabolic pathway, co-mutation on rs104886035 and rs104886038 of DHCR7 can reduce its stability and then inhibit the proliferation of gastric cancer cells [5]. Additionally, DHCR7 is also considered be a vitamin D-related gene, which may have an influence on calcium homeostasis, bone health, and various cancers [7]. DHCR7 polymorphisms have been shown to increase the risk of thyroid cancer via their effects on circulating vitamin D levels [8].
However, recent two findings revealed a paradox: DHCR7 deficiency is capable of suppressing cancer cell death resulted from the accumulation of its substrate, 7-Dehydrocholesterol (7-DHC) via its antiferroptotic activity [9,10]. In terms of viral infections, DHCR7 inhibitors have been suggested to have potential as viral therapeutics due to their effects on cholesterol metabolism and innate immunity [2]. Furthermore, Hongyan Wang, etc. investigate the role of 7-dehydrocholesterol reductase in the immune response to viral infections, they found that cholesterol, which is synthesized from 7-dehydrocholesterol by DHCR7, can inhibit the activation of Interferon Regulatory Factor 3 (IRF3), a key transcription factor in the antiviral immune response, indicating that inhibition of DHCR7 activity could be a potential therapeutic strategy for enhancing the antiviral immune response [11]. However, there is currently limited evidences to suggest a direct relationship between DHCR7 and tumor immunology. In this study, we comprehensively analyzed the role of DHCR7 in prognosis and immunology in most of human cancers. And we demonstrated the potential association between DHCR7 expression and immune subtypes, molecular subtypes, immune biomarkers and Tumor-Infiltrating Lymphocytes (TILs) in the Tumor Micro Environment (TME) of various cancer types. Furthermore, taking urogenital cancer as an example, we elucidated the connections of DHCR7 expression and urogenital anticancer immunotherapy. Collectively, we summarized the diverse effects of DHCR7 gene in human cancers and explored its roles in tumor immunity of various cancer. Accumulating evidence indicates that DHCR7 might be an optimal biomarker for prognosis and immunotherapy in various cancers.
Materials & Methods
Oncomine Analysis
The Oncomine database (https://www.oncomine.com) is a bioinformatics tool for collecting, standardizing, analyzing and delivering cancer transcriptome data to the biomedical research community. It was used to compare the transcription levels of DHCR7 between cancer specimens and para carcinoma tissue. In Oncomine, Student's t-test is generated for two class differential expression analyses. In the present study, p<0.01 and an absolute fold-change ≥1.5 were selected as the cut off values to analyze the gene expression chart of each DHCR7 family member.
Gene Expression Profiling Interactive Analysis (GEPIA) Database
GEPIA (http://gepia.cancer-pku.cn/) is a web tool that provides fast and customizable functionalities based on data from The Cancer Genome Atlas (TCGA; http://tcga-data.nci.nih.gov/ tcga/) and the Genotype-Tissue Expression project (GTEx; http:// www.gtexportal.org/home/index.html). Differential analysis was performed using one-way ANOVA, using disease state or tumor stage as the variable for calculating differential expression. In our study, GEPIA was used to exhibit the differential expression of DHCR7 between pan cancer and the related para carcinoma tissues, and the association between the expression of DHCR7 and tumor stages in patients.
The Human Protein Atlas Database
The Human Protein Atlas (https://www.proteinatlas.org/) is a database of immunohistochemistry (IHC)-based protein expression profiles in normal tissue, cancer and cell lines. IHC images of DHCR7 protein expression in clinical specimens of patients with LUAD and para carcinoma tissues were obtained from the Human Protein Atlas database.
Kaplan-Meier Plotter
The Kaplan-Meier Plotter tool (www.kmplot.com) includes survival information of 866 patients with LUAD. The prognostic value of DHCR7 expression was assessed by Overall Survival (OS), Progression-Free Survival (PFS) and Post-Progression Survival (PPS), using the Hazard Ratio (HR), 95% Confidence Intervals (CI) and log-rank P-value. In the analysis, patient samples were split into high expression group and low expression group based on the median mRNA levels of the DHCR7. The prognostic value of a gene was assessed by univariate Cox regression analysis. JetSet scores were used to select a single representative probe set for each gene. In the current study, only the probe sets with best JetSet scores for DHCR7 were selected to produce Kaplan-Meier plots. The one-to-one matches between DHCR7 genes and probe sets, identified by Affymetrix IDs, were as follows: DHCR72 and 272107_s_at; DHCR73 and 201555_ at; DHCR74 and 222036_s_at; DHCR75 and 216237_s_at; DHCR76 and 238977_at; DHCR77 and 208795_s_at; DHCR78 and 224320_s_at; and DHCR710 and 223570_at. The relevant concepts are defined as follows: OS, time from diagnosis to death; PFS, time from diagnosis to tumor progression; PPS, time from progression to death; HR>1, worse survival prognosis for the group with high mRNA expression; HR<1, unfavorable survival prognosis in the low mRNA expression group; 95% CI does not cross 1, mRNA expression is associated with survival rate. As not all gene expression levels were available in all patients and only the JetSet probes were included in the study, the sample sizes vary for each survival analysis.
cBioPortal for Cancer Genomics Dataset
cBioPortal (http://cbioportal.org) is based on other authoritative databases, including the Gene Expression Omnibus (GEO; http:// www.ncbi.nlm.nih.gov/geo/) and TCGA database. cBioPortal is a web resource for exploring, visualizing and analyzing multidimensional cancer genomics data. The genomic profile of each gene includes mutations, putative copy-number alterations and mRNA expression z-scores. The z score for each gene is the normalized expression of mRNA using RNA-Seq by expectation maximization count estimates method. The co-expression of each gene pair was performed by Fisher's exact test and the network was constructed according to the correlation.
TISIDB Database for Relationship between DHCR7 and Tumor Immune System
The TISIDB (http://cis.hku.hk/TISIDB/) is a web portal for tumor and immune system interaction, which integrates multiple heterogeneous data types. In this study, we analyzed the associations between DHCR7 expression and immune subtypes and molecular subtype across human cancers. And the relationship between DHCR7 expression and pan-cancer stages also analyzed by TISIDB [12].
Ualcan Database
UALCAN (https://ualcan.path.uab.edu/index.html) is a comprehensive, user-friendly, and interactive web resource for analyzing cancer OMICS data [13]. Here, we analyzed the staining of IHC and HE for DHCR7 in BLCA. And the relationships between the differential DHCR7 expression in bladder cancer and clinical subgroups was analyzed by the UALCAN database.
LinkedOmics Database for DHCR7 Co-Expression Networks
The LinkedOmics database (http://www.linkedomics.org/login. php) is a powerful visual platform that facilitates the exploration of gene expression profiles [14]. In our study, we utilized LinkedOmics to identify the co-expression genes of DHCR7 via Pearson's correlation coefficient, and visualized the results through heat maps and volcano plots. Subsequently, we employed Gene Set Enrichment Analysis (GSEA) to investigate the Gene Ontology Biological Processes (GO_BP) and KEGG pathways associated with DHCR7 and its co-expression genes.
Results
The Expression of Dhcr7 Gene in Pan-Cancers
The Oncomine database demonstrated that mRNA expression of DHCR7 was significantly higher in various human cancers, including colorectal cancer, bladder cancer, breast cancer, head and neck cancer, and ovarian cancer, when compared to corresponding normal tissues. However, DHCR7 expression was found to be significantly lower in brain and CNS cancer, cervical cancer, melanoma, and prostate cancer (Figure 1A). Furthermore, TIMER database revealed that DHCR7 expression was significantly higher in several cancer types, such as BLCA (Bladder Urothelial Carcinoma), BRCA (Breast Invasive Carcinoma), CESC (Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma), COAD (Colon Adenocarcinoma), ESCA (Esophageal Carcinoma), HNSC (Head And Neck Cancer), KICH (Kidney Chromophobe), LIHC (Liver Hepatocellular Carcinoma), LUAD (Lung Adenocarcinoma), LUSC (Lung Squamous Cell Carcinoma), READ (Rectum Adenocarcinoma), STAD (Stomach Adenocarcinoma), THCA (Thyroid Carcinoma), and UCEC (Uterine Corpus Endometrial Carcinoma). Conversely, DHCR7 mRNA expression was low in CHOL (Cholangiocarcinoma), KIRC (Kidney Renal Clear Cell Carcinoma), KIRP (Kidney Renal Papillary Carcinoma), and PCPG (Pheochromocytoma And Paraganglioma) (Figure 1B). Consistently, the GEPIA database displayed significant elevation of DHCR7 mRNA expression across most cancer types (Figure 1C). Additionally, the UALCAN database analysis indicated significant differential expression of DHCR7 between tumors and healthy tissues in various human cancers (Supplementary Figure 1). Overall, these results suggested that DHCR7 is overexpressed in most human cancer tissues.
Figure 1: Dhcr7 Expression Levels in Human Cancers (A) DHCR7 expression in different cancers and paired normal tissue in the Oncomine database. (B) DHCR7 expression levels in different cancer types from the TCGA database analyzed by the TIMER database. (C) DHCR7 expression in several cancers and paired normal tissue in the GEPIA database (*P < 0.05, **P < 0.01, ***P < 0.001).
Highly Expressed DHCR7 is Associated with Poor Prognosis in Human Cancers
Given the high expression of DHCR7 in many cancer cells, it is important to examine its relationship with Overall Survival (OS) or Disease-Free Survival (DFS). We then conducted an analysis using multiple databases. Firstly, from the Kaplan-Meier plotter database, we found that higher DHCR7 expression was associated with poor OS in BLCA (n = 404, HR = 1.66, P = 0.00064) (Figure 2A), CESC (n=304, HR=2.12, P=0.0015) (Figure 2B), HNSC (n=499, HR=1.54, P=0.0027) (Figure 2C), LIHC (n=370, HR=1.59, P=0.01) (Figure 2D), LUAD(n=504, HR=1.54, P=0.0073) (Figure 2E), PAAD (n=177, HR=1.78, P=0.009) (Figure 2F), SARC (n=259, HR=2.11, P=0.00021) (Figure 2G) and UCEC (n=542, HR=1.75, P=0.01) (Figure 2H). Secondly, a higher DHCR7 expression was related to worse OS (n = 76, HR=2.2, P=0.05) (Figure 2I) and diseases free survival (DFS) (n = 76, HR=2.6, P=0.0068) (Figure 2J) in ACC by GEPIA database. We also found that higher DHCR7 expression was related to worse OS in UVM (n = 78, HR=2.9, P=0.019) (Figure 2K). And a higher DHCR7 expression was related to worse DFS in LUSC (n = 482, HR=1.5, P=0.031) (Figure 2L). Similarly, we found that higher DHCR7 expression was related poor prognosis in human cancers from the TISIDB database (Supplementary Figure 2). Collectively, these results indicate that highly expressed DHCR7 is associated with poor prognosis in most human cancers.
Figure 2: Highly DHCR7 Expression is Associated with Poor Overall Survival in Human Cancers. Kaplan-Meier survival curve of human cancers with high and low DHCR7 expression analyzed by the Kaplan-Meier plotter database (A–H) and the GEPAI database (I-L). (A–H) Highly DHCR7 expression was related to worse OS in BLCA (n=404), CESC(n=304), HNSC(n=499), LIHC(n=370), LUAD(n=504), PAAD(n=177), SARC(n=259) and UCEC(n=542) cohorts. (I-J) High DHCR7 expression was related to worse OS and DFS in ACC (n = 76) cohorts. (K) High DHCR7 expression was related to worse OS in UVM (n = 78) cohorts. (L) High DHCR7 expression was related to worse DFS in LUSC cohorts (n = 482). OS, overall survival; DFS, disease free survival.
DHCR7 Expression is Related to Pathological Stages and Metastasis of Human Cancers
Then, we investigate the association between DHCR7 expression and tumor pathological stages as well as metastasis. The TISIDB database showed that the mRNA levels of DHCR7 were significantly different expressed between the tumor stages I to III or IV in BLCA (spearman r=0.13, p=0.00854) (Figure 3A), KICH (spearman r=0.253, p=0.0408) (Figure 3B), KIRP (spearman r=0.249, p=4.75e-05) (Figure 3C), LUAD (spearman r=0.139, p=0.00165) (Figure 3D), LUSC (spearman r=0.114, p=0.0113) (Figure 3E) and TGCT (spearman r=0.353, p=0.00123) (Figure 3F). In addition, we analyzed the differential expression of the DHCR7 in tumor, normal, and metastatic tissues with TNMplot. com [15].
Figure 3: The Relationship between DHCR7 Expression and Pan-Cancer Stages by TISIDB. (A) in BLCA, (B) in KICH, (C) in KIRP,(D) in LUAD, (E) in LUSC, (F) in TGCT
According to the gene chip data from the TNMplot, we can see that DHCR7 are differently expressed in breast invasive carcinoma (p(Kruskal-Wallis)=8.85e-18, n(normal)=242, n(tumor)=7569 and n(metastatic)=82) (Figure 4A), colon cancer (p(Kruskal- Wallis)=6.77e-43,n(normal)=377,n(tumor)=1450andn(metastatic) =99) (Figure 4B), kidney cancer (p(Kruskal-Wallis)=6.34e-17, n(normal)=277, n(tumor)=556 and n(metastatic)=58) (Figure 4C), hepatic carcinoma (p(Kruskal-Wallis)=6.87e-10, n(normal)=379, n(tumor)=806 and n(metastatic)=24) (Figure 4D), lung cancer (p(Kruskal-Wallis)=3.7e-12, n(normal)=391, n(tumor)=1865 and n(metastatic)=8) (Figure 4E), oesophageal carcinoma (p(Kruskal-Wallis)=2.82e-25, n(normal)=88, n(tumor)=440 and n(metastatic)=28) (Figure 4F), oral cancer (p(Kruskal-Wallis)=5.39e-03, n(normal)=18, n(tumor)=38 and n(metastatic)=5) (Figure 4G) and pancreatic cancer(p(Kruskal- Wallis)=1.41e-13, n(normal)=108, n(tumor)=248 and n(metastatic)=17) (Figure 4H).
Based on the above results, elevated expression of DHCR7 was observed in advanced stages and metastatic tissues of human cancers, which means the DHCR7 might be an optimal prognostic, tumor progression, and novel therapeutic biomarker.
Figure 4: The Relationship between Dhcr7 Expression and Pan-Cancer Metastasis. (A) in Breast, (B) in Colon, (C) in Kidney, (D) in Liver, (E) in Lung, (F) in Oesophageal, (G) in Oral cavity, (H) in Pancreas
DHCR7 Expression is Associated with Immune and Molecular Subtypes of Human Cancers
We next explored the DHCR7 expression on immune and molecular subtypes among human cancers by the TISIDB database. Immune subtypes consist six types, which were C1(wound healing), C2(IFN-γ dominant), C3(inflammatory), C4 (lymphocyte depleted), C5 (immunologically quiet) and C6 (TGF-β dominant). The results showed that DHCR7 expression was related to different immune subtypes in ACC, BLCA, BRCA, COAD, HNSC, KIRC, KIRP, LUAD, LUSC, OV, PRAD and UCEC (Figure 5). Particularly, DHCR7 expression varied among different immune subtypes in various tumor types. For instance, DHCR7 exhibited high expression in C5 subtypes of ACC and KIRC. However, in C3 subtypes of LUAD, OV, BRCA, and LUSC, DHCR7 displayed low expression. Regarding molecular subtypes of cancers, our analysis revealed a significant association between DHCR7 expression and ACC, BRCA, ESCA, HNSC, KIRP, LGG, LIHC, LUSC, PCPG, PRAD, STAD, and UCEC (Figure 6). These findings demonstrate that DHCR7 expression varies across immune subtypes and molecular subtypes in various human tumors.
Figure 5: The Relationship between DHCR7 Expression and Pan-Cancer Immune Subtypes. (A) in ACC, (B) in BLCA, (C) in BRCA,(D) in COAD, (E) in HNSC, (F) in KIRC, (G) in KIRP, (H) in LUAD, (I) in LUSC, (J) in OV, (K) in PRAD, (L) in UCEC
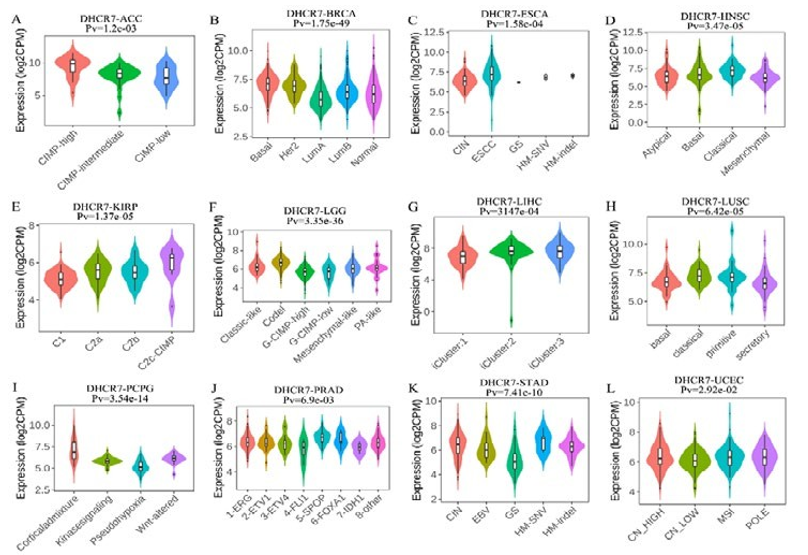
Figure 6: The Relationship between DHCR7 Expression and Pan-Cancer Molecular Subtypes. (A) in ACC, (B) in BRCA, (C) in ESCA,(D) in HNSC, (E) in KIRP, (F) in LGG, (G) in LIHC, (H) in LUSC, (I) in PCPG, (J) in PRAD, (K) in STAD, (L) in UCEC
DHCR7 Expression and Immune Checkpoint (ICP) Genes in Pan Cancers
Nowadays, Immune Checkpoint (ICP)-targeting drugs, including anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4), anti- programmed cell death protein 1 (PD-1), and anti-PD-1 ligand 1 (PD-L1) agents, have demonstrated remarkable efficacy in various cancer immunotherapies, such as lung cancer, melanoma, bladder cancer, and other malignancies. However, despite these achievements, a substantial proportion of patients across different cancer types still exhibit a limited positive response rate to immune checkpoint blockade therapy. This observation underscores the need for further exploration and understanding of the underlying drug resistance mechanisms involved [16,17].
Therefore, we analyzed the relationship between DHCR7 expression and ICP in the tumor microenvironment. Among 60 ICP genes, including 24 immune inhibitors (such as CTLA4, LAG3 or VEGFB) and 36 activators (TNFRSF9, CD28 or TNFRSF9, etc.), we found DHCR7 played roles both in inhibitors and activators of ICP in human cancers (Figure 7). For example, positive correlations between DHCR7 expression and the immune inhibitors CD276 and VEGFA was found in many cancers, such as TGCT, BLCA, HNSC, LUAD, LUSC, STAD, STES, CESC, SARC and PRAD, etc. For immune activators, negative correlations between DHCR7 expression and CD27, GZMA, CCL5, PRF1, ICOS, CD28 or CD40LG was showed in BLCA, HNSC, LUAD, LUSC, KIRC, STAD, STES, CESC, SARC, KIPAN and ACC. These results strongly suggest that DHCR7 could serve as a promising pan-cancer biomarker and a potential optimal target for cancer immunotherapy.
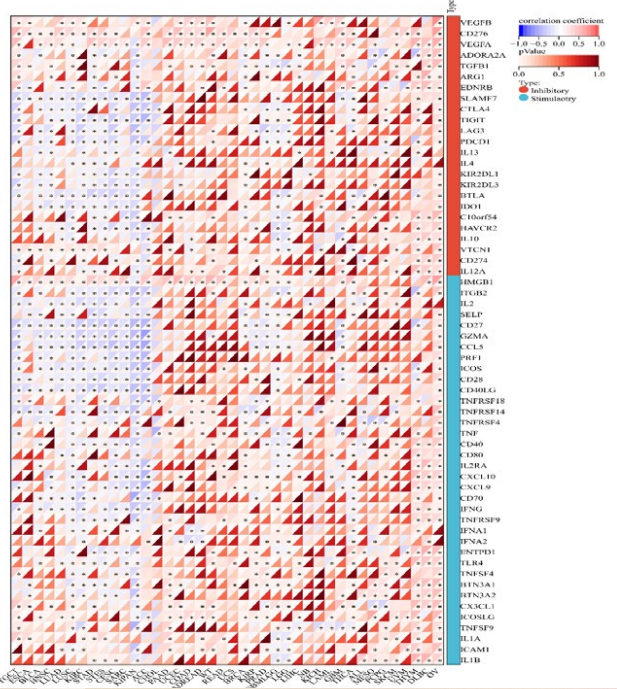
Figure 7: The Relationship between DHCR7 Expression and Pan-Cancer Immune Checkpoint Genes. *P < 0.05; **P < 0.01; ***P < 0.001.
DHCR7 Expression is Related to Microsatellite Instability (MSI), Neoantigen, Tumor Mutational Burden (TMB), and Mutant-Allele Tumor Heterogeneity (MATH)
Multiple studies have demonstrated that patients with microsatellite instability, neoantigen, tumor mutational burden or mutant-allele tumor heterogeneity were significantly associated with immunotherapy-related responses [18-21]. To characterize the roles of DHCR7 in the immune response of the Tumor Micro Environment (TME), we analyzed the correlations between the DHCR7 expression and TMB, MSI, neoantigen and Mutant Allele Tumor Heterogeneity (MATH). We observed that DHCR7 had significant positive correlations with MSI in 4 tumors: CESC (N=302, R=0.127, P=0.028), LUAD (N=511, R=0.122, P=0.006), KIPAN (N=688, R=0.341, P=3.50e-20), TGCT (N=148, R=0.237, P=0.004). And negative relation in 3 tumors: PRAD (N=495, R=- 0.120, P=0.008,) THCA (N=493, R=-0.094, P=0.036), UCS (N=57 R=-0.274, P=0.039) (Figure 8A). For neoantigen, we observed DHCR7 had significant positive correlation in LUAD (N=462, R=0.165, P=0.001) (Figure 8B). Analyzing the relationships between DHCR7 expression and TMB, we found that DHCR7 expression was positively related to TMB in GBM(N=149, R=0.170, P=0.038), LUAD(N=509, R=0.264, P=1.576e-9), BRCA(N=981, R=0.220, P=3.339e-12), STES(N=589, R=0.145, P=0.0004), STAD(N=409, R=0.268, P=3.934e-8), HNSC(N=498, R=0.171, P=0.0001), THYM(N=118, R=0.504, P=5.781e-9), LIHC(N=357, R=0.121, P=0.022), MESO(N=82, R=0.229, P=0.040), PAAD(N=171, R=0.324, P=0.00001), BLCA(N=407, R=0.145, P=0.003), ACC(N=77, R=0.266, P=0.019) and negatively related to TMB in LGG(N=501, R=-0.101, P=0.025), KIPAN(N=679, R=-0.250, P=3.757e-11), THCA(N=489, R=- 0.174, P=0.0001) and SKCM(N=102, R=-0.313, P=0.001) (Figure 8C). We also found that DHCR7 expression was positively related to MATH in LUAD(N=508, R=0.163, P=0.0002), BRCA(N=980, R=0.198, P=4.193e-10), STES(N=589, R=0.281, P=3.618e-12), KIPAN(N=679, R=0.166, P=0.00001), STAD(N=409, R=0.250, P=3.026e-7), HNSC(N=498, R=0.237, P=9.016e-8), LUSC(N=485, R=0.102, P=0.025), THYM(N=118, R=0.185, P=0.044), and BLCA(N=407, R=0.198, P=0.00005) and negatively to MATH in GBMLGG(N=649, R=-0.097, P=0.014), LGG(N=500, R=-0.130, P=0.003) and DLBC(N=37, R=-0.349, P=0.034) (Figure 8D). These results provide further evidence to suggest that the DHCR7 gene may play a critical role in the immunotherapy of human tumors.
Figure 8: The Relationship between DHCR7 Expression (A) MSI, (B) Neoantigen, (C) TMB and (D) MATH in human cancers. MSI: Microsatellite Instability, TMB: Tumor Mutational Burde, MATH: Mutant-Allele Tumor Heterogeneity
Relationship between DHCR7 Expression and Immune Cell Infiltration in Pan Cancers
The presence of multiple immune cell infiltrations in the tumor immune microenvironment can influence tumorigenesis, progression and even metastasis. Therefore, we analyzed the potential connection of DHCR7 expression and immune cells infiltration in TME of human cancers. We obtained 10 categories of immune cell infiltration scores for 10,180 tumor samples in a total of 44 tumor types. We calculated the Pearson's correlation coefficient between gene and immune cell infiltration scores in each tumor using the corr.test function of R package psych (version 2.1.6) to identify significantly correlated immune infiltration scores, and finally we observed that in 40 cancer species (TCGA-GBM (N=152), TCGA-GBMLGG (N=656), TCGA-LGG (N=504), TCGA-UCEC (N=178), TARGET-LAML (N=142), TCGA-BRCA (N= 1077), TCGA-CESC (N=291), TCGA-LUAD (N=500), TCGA-ESCA (N=181), TCGA-STES (N=569), TCGA- SARC (N=258), TCGA-KIRP (N=285), TCGA-KIPAN (N=878), TCGA -COAD (N=282), TCGA-COADREAD (N=373), TCGA- PRAD (N=495), TCGA-STAD (N=388), TCGA-HNSC (N=517), TCGA-KIRC (N=528), TCGA-LUSC (N=491), TCGA-THYM (N =118), TCGA-LIHC (N=363), TARGET-WT (N=80), TCGA- SKCM-P (N=101), TCGA-SKCM (N=452), TCGA-BLCA (N=405), TCGA-SKCM-M (N=351), TCGA-THCA (N=503), TCGA TCGA-OV (N=417), TCGA-UVM (N=79), TCGA-PAAD (N=177), TCGA-TGCT (N=132), TCGA-UCS (N=56), TCGA- LAML (N=214), TARGET-ALL (N=86), TCGA-PCPG (N=177), TCGA-ACC (N=77), TCGA-DLBC (N=46), TCGA-KICH (N=65), TCGA-CHOL (N=36)) in which the gene expression was significantly associated with immune in cells filtration (Figure 9A).
The results demonstrated a robust correlation between DHCR7 expression and various immune cell types in human cancers, including T-cells, CD8 T-cells, Cytotoxic lymphocytes, B-lineage cells, NK-cells, Monocytic lineage cells, Myeloid dendritic cells, Neutrophils, Endothelial cells, and Fibroblasts. These findings were consistent across six urogenital cancers (Figure 9B). Additionally, using the TIMER database, we analyzed the correlation between DHCR7 and immune cell-related genes and markers. Table illustrates that DHCR7 displayed close connections with all included marker genes of CD8+ T cells, monocytes, dendritic cells, regulatory T cells (Tregs), and T cell exhaustion markers in BLCA, KIRC, and PRAD. Notably, there was no correlation between gene markers of M1 macrophages and DHCR7 expression.
However, DHCR7 expression showed moderate to strong correlations with M2 macrophage markers, such as CD163, VSIG4, and MS4A4A in BLCA. Furthermore, DHCR7 expression exhibited positive effects on the expression of Treg and T cell exhaustion markers, including FOXP3, STAT5B, TGFB1, PDCD1, CTLA4, and LAG3 in BLCA, KIRC, and PRAD. Additionally, DHCR7 expression demonstrated significant correlations with the regulation of several markers of T helper cells (Th1, Th2, Tfh, and Th17) in the four bladder urothelial carcinomas. In summary, these findings highlight the crucial roles of DHCR7 in the recruitment and regulation of immune infiltrating cells across multiple cancers. Consequently, DHCR7 may serve as a valuable prognostic biomarker for bladder urothelial carcinomas and other cancer types.
Table: Correlation between DHCR7 and relate genes and markers of immune cells analyzed by TIMER
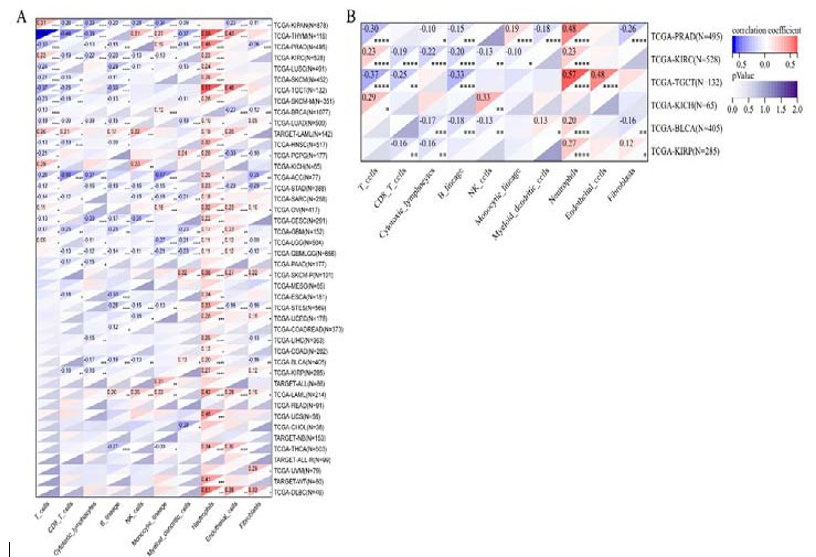
Figure 9: The Relationship between DHCR7 Expression and Infiltrating Immune Cells of Human Cancers and Urogenital Cancers (A) The relationship between DHCR7 expression level and infiltrating levels of B cells, CD4+ T cells, CD8+ T cells, macrophages, neutrophils, dendritic cell in human cancers. (B) The relationship between DHCR7 expression level and infiltrating levels of B cell lineages, CD8+ T cells, cytotoxic lymphocytes, endothelial cells, fibroblasts, monocytic cell lineages, myeloid dendritic cells, neutrophils, natural killer cells, T cells in six urogenital cancers. *P < 0.05; **P < 0.01; ***P < 0.001.
Genetic Alterations and Expression of DHCR7 in Urogenital Cancers
Next, we conducted an analysis of genomic alterations of DHCR7 in urogenital cancers using the cBioPortal website. The results revealed that DHCR7 genomic alterations were present in 1.5% of patients. These alterations encompassed different types, such as missense mutations, truncating mutations, amplifications, and deep deletions (Figure 10A). Interestingly, urogenital cancers exhibited varying levels of gene expression depending on the specific type of DHCR7 gene alteration (Figure 10B). Copy Number Variations (CNVs) were predominantly observed in BLCA and PRAD, while no CNVs were found in kidney chromophobe and kidney renal papillary cell carcinoma (Figure 10C). To assess DHCR7 expression at the protein level, we obtained immunohistochemical images from the HPA database. Taking BLCA as an example, we observed significantly higher protein expression of DHCR7 in BLCA tissues compared to normal tissues (Figure 11A and B). Additionally, we explored DHCR7 expression in BLCA with different clinical characteristics using the UALCAN database. The data demonstrated significant differential expression of DHCR7 across various factors, including cancer stages, histological subtypes, molecular subtypes, nodal metastasis status, and TP53 mutation status in BLCA (Figure 11C–H). Collectively, these findings indicate that DHCR7 experiences genomic alterations and differential expression in diverse tumor types, underscoring its pivotal role in cancer onset and progression.
Figure 10: DHCR7 Genomic Alterations in Six Urogenital Cancers Analyzed by the cBioPortal database (A–C) (A) OncoPrint of DHCR7 gene alterations in cancer cohort. (Different colors means different types of genetic alterations and amplification accounts for the largest proportion). (B) Main type of DHCR7 gene alterations in cancer groups. (C) Details of DHCR7 gene alteration types in cancer cohorts
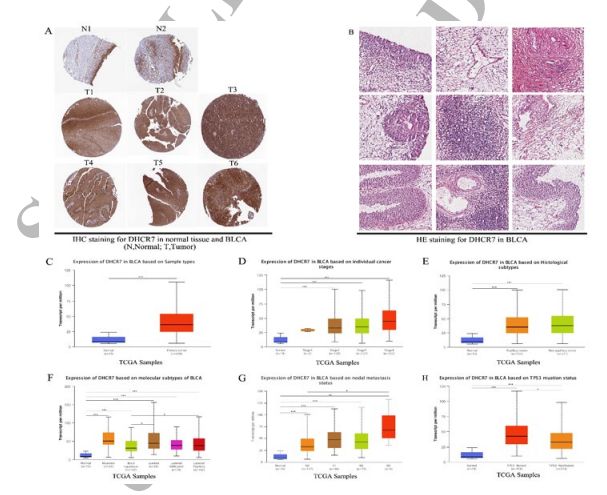
Figure 11: The Staining of IHC and HE for DHCR7 in BLCA (A-B) and DHCR7 Differential Expression in Bladder Cancer with Different Clinical Subgroups (C–H) Analyzed by the UALCAN Database. (A) IHC staining for DHCR7 in normal tissue and BLCA patient tissue analyzed by HPA database (B) HE staining for DHCR7 in BLCA patient tissue analyzed by HPA database; (C–H) DHCR7 expression between normal(n=19) and BLCA primary tumor(n=408) (C) DHCR7 differential expression in BLCA with individual cancer stages (n = 400) (D) Histological Subtypes (n = 403) (E) Molecular Subtypes (n = 408) (F) Nodal Metastasis Status (n = 366) (G) TP53 Mutation Status (n = 408) (H) (*P < 0.05, **P < 0.01, ***P < 0.001)
Gene Set Enrichment Analysis of DHCR7 in BLCA
The above results revealed a significant association between DHCR7 and the prognosis and immunology of cancers. To verify the potential function of DHCR7 in tumor tissue, we explored DHCR7 co-expression networks using the Linked Omics database. In BLCA, a set of genes (represented by dark red dots) displayed a significant positive correlation with DHCR7, while another set of genes (represented by dark green dots) showed a negative correlation (False Discovery Rate (FDR) < 0.01) (Figure 12 A). A heat map was generated to show the top 50 genes that positively and negatively correlated with DHCR7 (Figure 12B, C). Furthermore, From the Supplementary Table, we can see that SQLE (squalene epoxidase) (r = 0.598), SCD (stearoyl-CoA desaturase) (r = 0.556), and FASN (fatty acid synthase) (r = 0.524) were three genes which exhibited the strongest association with DHCR7 expression in BLCA (p=7.653e-41, 1.885e-34, and 4.095e-30, respectively). To determine the main gene ontology (GO) terms of DHCR7 co-expression genes, we employed Gene Set Enrichment Analysis (GSEA). Analysis of the biological process categories of GO revealed that DHCR7 and its co-expression genes primarily participate in the regulation of the immune system process, leukocyte differentiation, virus response, and angiogenesis (Figure 12D). Subsequently, we conducted Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, which demonstrated that the co-expressed genes were enriched in cell cycle, metabolic pathways, protein processing in the endoplasmic reticulum, and the chemokine signaling pathway (Figure 12E). These findings suggest that DHCR7 expression may play essential roles in human cancers by regulating the immune response of the tumor microenvironment.
Figure 12: DHCR7 Co-Expression Genes in BLCAAanalyzed through the Linked Omics Database. (A) Highly correlated genes of DHCR7 tested by Pearson test in BLCA cohort. (B, C) Top 50 positive co-expression genes (B) Negative co-expression genes (C) of DHCR7 in heat map in BLCA (D) Directed acyclic graph of DHCR7 GO analysis (Biological Process) in BLCA cohort. (E) Volcano plot of DHCR7 KEGG pathways in BLCA cohort
Discussion
DHCR7 gene was knew largely because of its links to the developmental disorder Smith-LemliOpitz syndrome, which is caused by mutations in the DHCR7 gene [22]. Recently, some studies showed DHCR7 plays key roles in several malignant tumors. However, the underlying effects on oncogenesis and tumor immunology are poorly defined.
In this work, we systematically examined the expression and prognostic values of DHCR7 in various types of cancer by several datasets. We found that DHCR7 was highly expressed in most of the cancers, and elevated DHCR7 expression was associated with poor prognosis. Intriguingly, our results also showed that DHCR7 involve immune cells infiltration and immune checkpoint markers, which indicated that DHCR7 plays a pivotal role in tumor immunology.
Firstly, we analyze the expression of DHCR7 in cancers and normal tissues by the Oncomine, TIMER, GEPIA, and the UALCAN database. Our results showed that DHCR7 was overexpressed in the majority of cancers other than the brain/ CNS cancer, cervical cancer, melanoma, and prostate cancer. These results were consistent with previous studies in bladder cancer and cervical cancer [4,6].
Next, we explored the relationships between the expression of DHCR7 and overall survival, tumor stages, and metastasis. The results indicated that highly DHCR7 expression was related to worse OS in BLCA, CESC, HNSC, LIHC, LUAD, PAAD, SARC, UCEC, ACC, UVM, and LUSC. As we know the stage of a cancer describes the size of a tumor and how far it has spread from where it originated. And a higher grade usually indicates a shorter survival time for patients. Our results showed that there is a positive correlation between DHCR7 expression and tumor stages or cancer metastasis in pan-cancer, such as BLCA, KICH, KIRP, LUAD, LUSC and TGCT. These results proved that DHCR7 was a potential prognostic pan-cancer biomarker and involves in tumorigenesis. To explored the potential mechanism, we analyzed the DHCR7 expression in immune subtypes and molecular subtypes of human cancers. The results showed that different expression level of DHCR7 was occurred in different immune subtypes and molecular subtypes in most cancersï¼?which suggests DHCR7 may be involved in tumor immunomodulation. Furthermore, clinical practice reveals that individual patient shows different response to immune checkpoint inhibitors. There are multiple factors that may be responsible for resistance Of Immune Checkpoint Inhibitors (ICIs) such as antigen presentation, TME, Tumor Associated Macrophages (TAM), immunosuppressive metabolites, genetic factors and biomarker response [23].
In this study, our data showed that strong correlations between the DHCR7 expression and ICP genes, TMB, MSI, neoantigen and Mutant Allele Tumor Heterogeneity (MATH) were found in most human cancers. Additionally, we also found that DHCR7 had a significant association with immune cells infiltration in TME of human cancers. For example, DHCR7 expression exhibited positive effects on the expression of Treg and T cell exhaustion markers, including FOXP3, STAT5B, TGFB1, PDCD1, CTLA4, and LAG3 in BLCA, KIRC, and PRAD. The above results suggest that the higher the DHCR7 expression, the more insensitive in the tumor immunotherapy, which implying the possibility of combining DHCR7 inhibitors of tumor therapeutic strategy in the future.
Conclusively, highly DHCR7 expression was correlated with advanced tumor stage, tumor metastasis, and poor prognosis for the most human cancers. However, further experimental researches are warranted to confirm the role of DHCR7 gene in oncogenesis and immunotherapy.
Funding
This study was supported in part by the National Natural Science Foundation of China (grant no.82200163), the Natural Science Foundation of Zhejiang Province of China (grant no. LQ22H080007) and the Medical Health Science and Technology Project of Hangzhou Municipal Health Commission (grant no. B20230646).
Availability of Data and Materials
The datasets analyzed during the study are publicly available from the following online databases: Oncomine database (https:// www.oncomine.com/); GEPIA database (http://gepia.cancer- pku.cn/); Kaplan-Meier Plotter (www.kmplot.com); The Human Protein Atlas (https://www.proteinatlas.org/); cBioPortal (http:// cbioportal.org); GEO (http://www.ncbi.nlm.nih.gov/geo/); TCGA (https://tcga-data.nci.nih.gov/tcga/);TIMER database(http://timer. cistrome.org/); TISIDB(http://cis.hku.hk/TISIDB/); UALCAN (https://ualcan.path.uab.edu/). LinkedOmics (https://linkedomics. org/login.php).
Ethical Approval and Consent to Participate
This study was performed in accordance with standard guidelines and was approved by the Ethics Committee of the Second Affiliated Hospital, Zhejiang University School of Medicine.
Authors Contribution
Xianghua Wu, Weiwei Zheng and Li Wang contributed equally to this work. Dan Lin and Zhaoxing Wu conceived, initiated, designed, and supervised the study. Xianghua Wu, Weiwei Zheng, Li Wang and Xuzhao Zhang acquired, analyzed and interpreted the data. Zhaoxing Wu and Weiwei Zheng drafted the manuscript, Dan Lin and Zhaoxing Wu revised it. All authors gave final approval of the version to be published, and agreed to be accountable for all aspects of the work.
References
- Kim, J. H., Lee, J. N., & Paik, Y. K. (2001). Cholesterol biosynthesis from lanosterol: a concerted role for Sp1 and NF-Y-binding sites for sterol-mediated regulation of rat 7-dehydrocholesterol reductase gene expression. Journal of Biological Chemistry, 276(21), 18153-18160.
- Xiao, J., Li, W., Zheng, X., Qi, L., Wang, H., Zhang, C., ... & Wang, H. (2020). Targeting 7-dehydrocholesterol reductase integrates cholesterol metabolism and IRF3 activation to eliminate infection. Immunity, 52(1), 109-122.
- Waye, J. S., Nakamura, L. M., Eng, B., Hunnisett, L., Chitayat, D., Costa, T., & Nowaczyk, M. J. M. (2002). Smith-Lemli- Opitz syndrome: carrier frequency and spectrum of DHCR7 mutations in Canada. Journal of medical genetics, 39(6), e31-e31.
- Zou, J., Liu, S., Long, J., & Yan, B. (2022). High DHCR7 expression predicts poor prognosis for cervical cancer. Computational and mathematical methods in medicine, 2022(1), 8383885.
- Chen, Y., Yan, W., Yang, K., Qian, Y., Chen, Y., Wang, R.,... & Chen, W. (2023). Integrated multi-dimensional analysis highlights DHCR7 mutations involving in cholesterol biosynthesis and contributing therapy of gastric cancer. Journal of Experimental & Clinical Cancer Research, 42(1), 36.
- Li, Y., Zhou, Y., Huang, M., Wang, Z., Liu, D., Liu, J., ... & Zhang, X. (2023). DHCR7 promotes tumorigenesis via activating PI3K/AKT/mTOR signalling pathway in bladder cancer. Cellular signalling, 102, 110553.
- Wen, J., Li, J., Liang, X., & Wang, A. (2021). Association of Polymorphisms in Vitamin Dâ?ÂÂMetabolizing Enzymes DHCR7 and CYP2R1 with Cancer Susceptibility: A Systematic Review and Metaâ?ÂÂAnalysis. Disease Markers, 2021(1), 6615001.
- Maciejewski,A., & Lacka, K. (2022). Vitamin D-Related Genes and Thyroid Cancer—A Systematic Review. International Journal of Molecular Sciences, 23(21), 13661.
- Li, Y., Ran, Q., Duan, Q., Jin, J., Wang, Y., Yu, L., ... &Wang, P. (2024). 7-Dehydrocholesterol dictates ferroptosis sensitivity. Nature, 626(7998), 411-418.
- Freitas, F. P., Alborzinia, H., Dos Santos, A. F., Nepachalovich, P., Pedrera, L., Zilka, O., ... & Friedmann Angeli, J. P. (2024). 7-Dehydrocholesterol is an endogenous suppressor of ferroptosis. Nature, 626(7998), 401-410.
- Korade, Z., Tallman, K. A., Kim, H. Y. H., Balog, M.,Genaro-Mattos, T. C., Pattnaik, A., ... & Porter, N. A. (2022). Dose–Response Effects of 7-Dehydrocholesterol Reductase Inhibitors on Sterol Profiles and Vesicular Stomatitis Virus Replication. ACS Pharmacology & Translational Science, 5(11), 1086-1096.
- Ru, B., Wong, C. N., Tong, Y., Zhong, J. Y., Zhong, S.S. W., Wu, W. C., ... & Zhang, J. (2019). TISIDB: anintegrated repository portal for tumor–immune system interactions. Bioinformatics, 35(20), 4200-4202.
- Chandrashekar, D. S., Karthikeyan, S. K., Korla, P. K., Patel, H., Shovon, A. R., Athar, M., ... & Varambally, S. (2022). UALCAN: An update to the integrated cancer data analysis platform. Neoplasia, 25, 18-27.
- Vasaikar, S. V., Straub, P., Wang, J., & Zhang, B. (2018). LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic acids research, 46(D1), D956-D963.
- Bartha, Á., & GyÃÂ??rffy, B. (2021). TNMplot. com: a web tool for the comparison of gene expression in normal, tumor and metastatic tissues. International journal of molecular sciences, 22(5), 2622.
- He, X., & Xu, C. (2020). Immune checkpoint signaling and cancer immunotherapy. Cell research, 30(8), 660-669.
- Tang, T., Huang, X., Zhang, G., Hong, Z., Bai, X., & Liang,T. (2021). Advantages of targeting the tumor immunemicroenvironment over blocking immune checkpoint in cancer immunotherapy. Signal transduction and targeted therapy, 6(1), 72.
- Li, K., Luo, H., Huang, L., Luo, H., & Zhu, X. (2020).Microsatellite instability: a review of what the oncologist should know. Cancer cell international, 20, 1-13.
- Zhang, Z., Lu, M., Qin, Y., Gao, W., Tao, L., Su, W., &Zhong, J. (2021). Neoantigen: A new breakthrough in tumor immunotherapy. Frontiers in Immunology, 12, 672356.
- Jardim, D. L., Goodman, A., de Melo Gagliato, D., & Kurzrock, R. (2021). The challenges of tumor mutational burden as an immunotherapy biomarker. Cancer cell, 39(2), 154-173.
- Gao, Y., Yang, C., He, N., Zhao, G., Wang, J., & Yang, Y. (2020). Integration of the tumor mutational burden and tumor heterogeneity identify an immunological subtype of melanoma with favorable survival. Frontiers in oncology, 10, 571545.
- Kandutsch, A. A. (1962). Enzymatic reduction of the Δ7 bond of 7-dehydrocholesterol. Journal of Biological Chemistry, 237(2), 358-362.
- Dobosz, P., Stepien, M., Golke, A., & Dzieciatkowski, T. (2022). Challenges of the immunotherapy: perspectives and limitations of the immune checkpoint inhibitor treatment. International journal of molecular sciences, 23(5), 2847.