Research Article - (2025) Volume 4, Issue 6
The Voxel-Based Three-Dimensional Conjecture of Double Helical DNA Structure
Received Date: May 02, 2025 / Accepted Date: May 25, 2025 / Published Date: Jun 04, 2025
Copyright: ©©2025 Jer-Fong Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Chen., J. F. (2025). The Voxel-Based Three-Dimensional Conjecture of Double Helical DNA Structure. J Math Techniques Comput Math, 4(6), 01-12.
Abstract
In molecular biology, the double helical structure of DNA has long captivated researchers due to its elegance and functionality. However, no single study has definitively proven that this configuration is the most efficient means for genetic information storage and transmission. We acknowledge the existence of this structure, yet the underlying reasons for its form remain elusive. This paper hypothesizes the existence of three-dimensional voxels and employs the voxel- based chemical element cube, constructed akin to a three-dimensional Othello board, to re-examine the DNA structure within this virtual framework and compare it to the DNA structure observed in the real world.
Keywords
DNA Structure, Voxel-Based, Virtual System
Css concepts
• Applied computing • Life and medical sciences • Computational biology • Molecular sequence analysis
• Computing methodologies • Modeling and simulation • Simulation types and techniques
• Computing methodologies • Artificial intelligence • Computer vision • Image and video acquisition • 3D imaging
Introduction
The discovery of the DNA double helical structure marks a pivotal moment in the history of molecular biology. In 1952, Rosalind Franklin and Raymond Gosling captured an X-ray diffraction image "Photo 51", which provided critical experimental evidence for the helical nature of DNA [1]. This image, though not immediately published, was instrumental in informing the model later proposed by James Watson and Francis Crick.
In April 1953, Nature published three landmark papers: Watson and Crick's structural model of DNA, supported by base-pairing rules; Wilkins and colleagues' crystallographic data; and Franklin and Gosling's detailed fiber diffraction analysis [2-4]. Together, these works established the now-canonical B-form double helix structure of DNA. This paper proposes a novel type of voxel, designed based on a chemical element cube constructed from a three-dimensional Othello board framework [5]. The foundational definitions introduced enable the cube to undergo three- dimensional rotations, facilitating the bonding of elemental cubes at specific orientations.
Justification for Applying the 3d Voxel-Based Framework to Cubic Elemental Rotations
According to the paper "Conjecture of the Marquee on Cube Surface of the Virtual Atomic Orbital and its Implementation of the Chemical Reactions System", molecules such as H2, O2, N2, H2O, CO2, H2PO2, and C2H2O2 can be constructed using cubic chemical elements, with each molecular structure exhibiting right- angled configurations, as illustrated in Figures 1 through 8 [5].
Unlike glucose, deoxyribose cannot be entirely constructed using non-rotated cubic elements. To address this, the present study proposes the three-dimensional voxel-based solution to achieve deoxyribose formation with minimal rotational adjustments. The proposed voxel unit consists of four cubic elements (Figures 9–12), and their assembled structure is shown in Figure 13. By replacing the second and third carbon atoms of deoxyribose with the voxel unit, and calculating the shortest spatial connection between them, the optimal solution involves rotating the second carbon element by 45 degrees along the z-axis, and the third carbon element by 45 degrees along the x-axis. This configuration enables the compound to be formed using single-bond-connected cubic chemical elements, as illustrated in Figures 14 and 15.
Figure 1: Hâ?? (hydrogen)
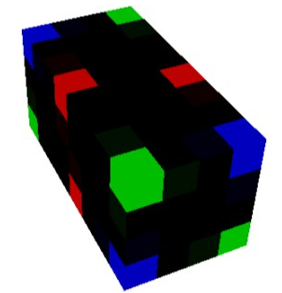
Figure 2: O2 (oxygen)

Figure 3: N2 (nitrogen)
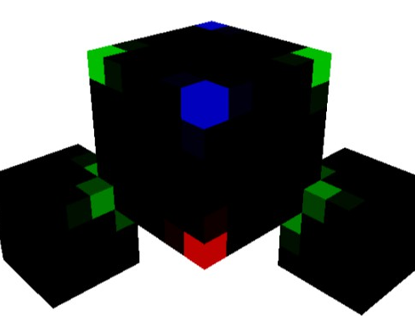

Figure 4: H2O (water)
Figure 5: CO2 (carbon oxide)
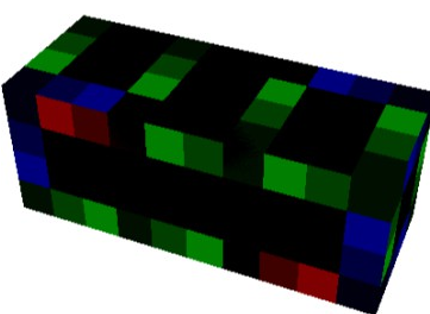

Figure 6: H2PO2 (phosphoric acid)

Figure 7: C2H2O2 (glucose front view)
Figure 8: C6H12O6 (glucose side view
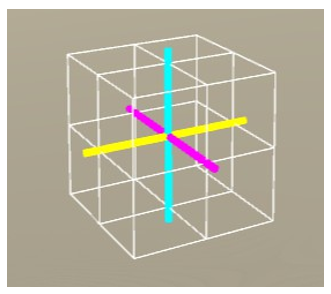
Figure 9: The unrotated voxel. (x-axis: yellow, y-axis: cyan, z-axis: magenta)
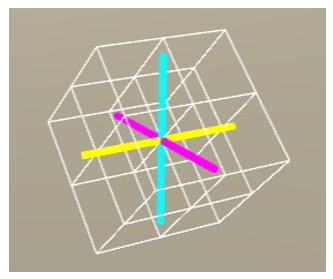
Figure 10: Rotate the voxel by 45° about the x-axis.
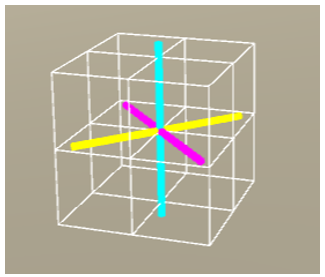
Figure 11: Rotate the voxel by 45° about the y-axis.
Figure 12: Rotate the voxel by 45° about the z-axis
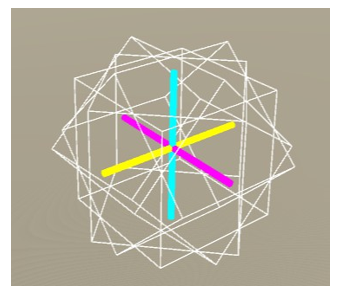
Figure 13: The voxel composed of four-unit cubes.

Figure 14: C5H10O4 (deoxyribose front view)
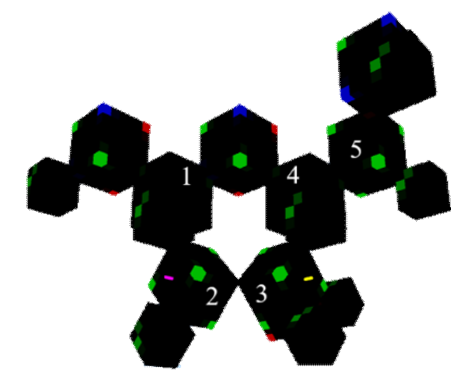
Figure 15: C5H10O4 (deoxyribose back view)
Subsequently, explore the construction of various DNA bases using cubic chemical elements. As illustrated in Figure 16, cytosine can be assembled without necessitating any voxel rotation. In the case of thymine, substituting the non-bonding nitrogen atom with the three-dimensional voxel and applying a 45-degree rotation along the x-axis resolves the connectivity issue, as depicted in Figure 17. Guanine presents a more complex scenario; it requires replacing both a carbon and a nitrogen atom—whose proximity hinders bonding—with two separate three-dimensional voxels; by rotating each voxel 45 degrees along the x-axis, we alleviate the excessive covalent bonding that would otherwise lead to electron density surpassing the permissible maximum on the cube surfaces, as shown in Figure 18 [2]. Furthermore, like guanine, adenine requires rotational adjustments to alleviate steric hindrance between closely positioned carbon and nitrogen atoms. Additionally, a specific nitrogen atom must be rotated 45 degrees about the y-axis to facilitate the formation of both a double bond and a single bond with two adjacent carbon atoms. This configuration enables the assembly of adenine using voxel-based cubic elements, as illustrated in Figure 19.
Figure 16: Cytosine (unrotated elements)
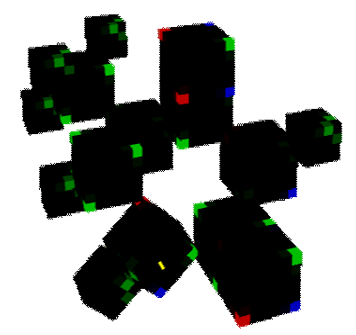

Figure 17: Thymine (a rotated non-hydrogen element exists)

Figure 18: Guanine (two rotated non-hydrogen elements exist)
Figure 19: Adenine (three rotated non-hydrogen elements exist)
Combining DNA
Nucleotides
To comprehend DNA, it is essential first to understand nucleotides. DNA is composed of four nucleotides: cytidine triphosphate (CTP), thymidine triphosphate (TTP), guanosine triphosphate (GTP), and adenosine triphosphate (ATP), as depicted in Figures 20 through 23. In these figures, the regions where the nitrogenous base connects to the deoxyribose sugar are highlighted in purple. These “connection points” serve as pivot axes, allowing the bases to rotate 45 degrees along the x-, or y-, or z-axis. Such rotations facilitate the formation of hydrogen bonds between complementary bases, leading to the canonical base pairings of cytosine with guanine (C-G) and adenine with thymine (A-T), as illustrated in Figure 24.
Figure 20: Cytidine triphosphate (CTP)
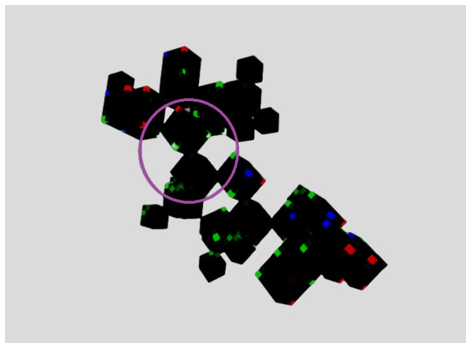
Figure 21: Thymidine triphosphate (TTP)
Figure 22: Guanosine triphosphate (GTP)

Figure 23: Adenosine triphosphate (ATP)
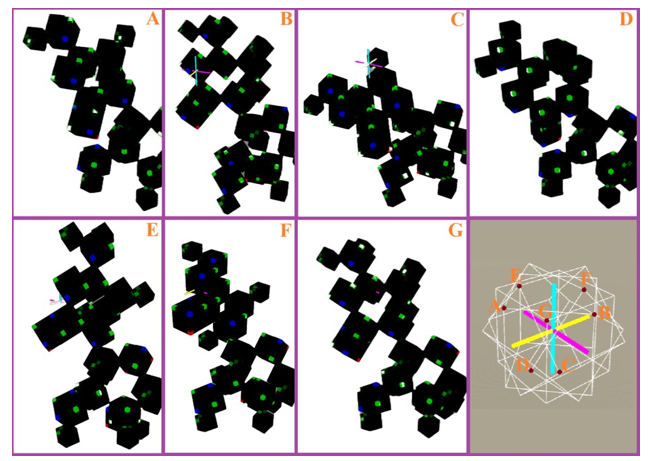
Figure 24: Assuming that point G serves as a non-rotated chemical single-bond connection point, applying 45-degree rotations about the X, Y, and Z axes respectively can generate six additional chemical single-bond connection points.
A-Form & B-Form
It is well-established that DNA can adopt multiple conformations, notably the A-form and B-form. In the voxel-based virtual modeling framework, it is feasible to represent these conformations through specific rotational configurations. Given that each voxel rotation is constrained to 45-degree increments, the orientation of the phosphate group's double-bonded oxygen atom cycles back to its original position after eight such rotations. In this model, when the double-bonded oxygen atoms of the phosphate groups—each connected via deoxyribose to complementary bases—are aligned in the same direction, the structure corresponds to the A-form DNA, as illustrated in Figure 25. Conversely, when these oxygen atoms are oriented oppositely, the structure represents the B-form DNA, as depicted in Figure 26.
Figure 25(a): The A-form DNA in the voxel-based virtual modeling framework. (abstract concept)

Figure 25(b): The A-form DNA in the voxel-based virtual modeling framework. (Cube composition, only 135° double helical, not 360°)
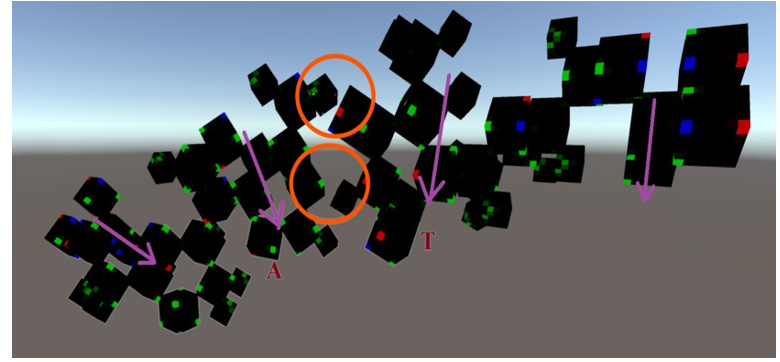
Figure 25(c): The A-form DNA in the voxel-based virtual modeling framework. (A-T pair)
Figure 25(d): The A-form DNA in the voxel-based virtual modeling framework. (C-T pair)
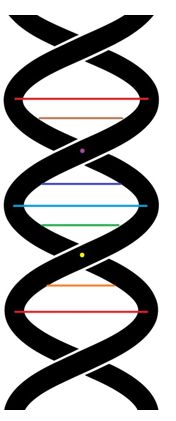
Figure 26(a): The B-form DNA in the voxel-based virtual modeling framework. (abstract concept)
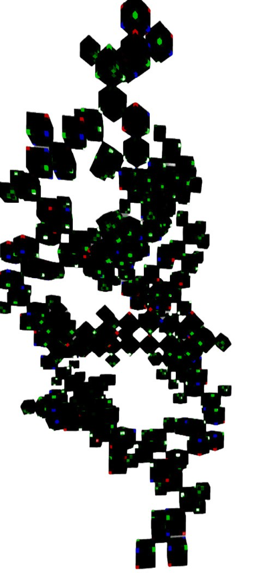

Figure 26(b): The B-form DNA in the voxel-based virtual modeling framework. (cube composition, only 135° double helical, not 360°)
Figure 26(c): The B-form DNA in the voxel-based virtual modeling framework. (A-T pair)
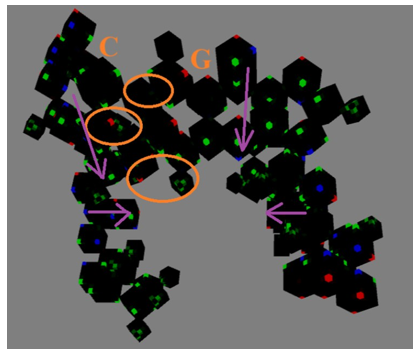
Figure 26(d): The B-form DNA in the voxel-based virtual modeling framework. (C-G pair)
From these illustrations, the A-form DNA resembles an elongated spring, whereas the B-form DNA appears akin to a compressed spring.
Implementation
The Voxel-Based Three-Dimensional Conjecture of Double Helical DNA Structure implementation URL is: https://www. cpapeijer.com/home_autolearn3.html . It is written in C# and uses Unity.
Results
In the voxel-based virtual environment, the emergence of the double helical DNA structure parallels that of the real world, primarily arising from the combined influence of the puckered conformation of the pentose sugar ring and the phosphate groups. However, within this virtual framework, the differentiation between A-form and B-form DNA is not attributed to variations in sugar puckering specifically, the C2'-endo and C3'-endo conformations observed in reality but rather to the orientation of the hydrogen-bonded base pair planes relative to the direction of the phosphate group's double-bonded oxygen atoms[6]. Comparative analyses between the virtual and actual DNA structures are presented in Tables 1 and 2.
|
Structural Parameters |
Virtual A-form DNA |
Real A-form DNA |
|
Helical direction |
Can be left-handed or right-handed |
right-handed |
|
Rise per base pair |
Height of 5 carbon atom cubes |
about 2.6 Å (0.26 nm) |
|
Helix pitch (height per turn) |
Height of 40 carbon atom cubes |
about 28.6 Å (2.86 nm) |
|
Number of base pairs per turn |
8 |
about 11 base pairs |
|
Helix diameter |
Width of 18 carbon atom cubes |
about 23 Å (2.3 nm) |
|
Base pair tilt angle |
90° in abstract conceptual diagram; about 80° in cube model |
about 20° |
Table 1: Comparison between virtual and real (A-form).[7]
|
Structural Parameters |
Virtual B-form DNA |
Real B-form DNA |
|
Helical direction |
Can be left-handed or right-handed |
right-handed |
|
Rise per base pair |
Height of 5 carbon atom cubes |
about 3.4 Å(0.34 nm) |
|
Helix pitch (height per turn) |
Height of 40 carbon atom cubes |
about 35.7 Å(3.57) |
|
Number of base pairs per turn |
8 |
about 10.5 base pairs |
|
Helix diameter |
Width of 11 carbon atom cubes |
about 20 Å(2.0 nm) |
|
Base pair tilt angle |
0° in abstract conceptual diagram; about 5° in cube model |
about 6° |
Table 2: Comparison between virtual and real (B-form).[7]
Conclusion
This paper introduces the voxel-based modeling framework to reinterpret the structural characteristics of DNA, particularly focusing on the distinctions between the A-form and B-form conformations. By employing a three-dimensional voxel hypothesis, we constructed chemical element cubes capable of specific rotational behaviors, enabling the assembly of complex molecular structures such as deoxyribose and various nucleobases. Our findings suggest that, within this virtual voxel environment, the differentiation between A-form and B-form DNA is not governed by traditional sugar puckering conformations (C2'-endo and C3'- endo) as observed in the real world. Instead, it is determined by the orientation of hydrogen-bonded base pair planes relative to the direction of the phosphate group's double-bonded oxygen atoms. This alternative perspective offers a novel approach to understanding DNA's structural dynamics and may provide new insights into the principles governing genetic information storage and transmission.
References
- Glynn, J. (2008). Rosalind Franklin: 50 years on. Notes and Records of the Royal Society, 62(2), 253-255.
- Watson, J. D., & Crick, F. H. (1953). Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature, 171(4356), 737-738.
- Wilkins, M. H. F., Stokes, A. R., & Wilson, H. R. (2003). Molecular structure of deoxypentose nucleic acids. Nature, 421(6921).
- Franklin, R. E., & Gosling, R. G. (1953). Molecular configuration in sodium thymonucleate. Nature, 171(4356), 740-741.
- J.F. Chen. 2025. Conjecture of the Marquee on Cube Surface of the Virtual Atomic Orbital and its Implementation of the Chemical Reactions System. Journal of Mathematical Techniques and Computational Mathematics, 4(1), 01-06.
- Saenger, W. (1984). Principles of Nucleic Acid Structure.
- Richmond, T. J., & Davey, C. A. (2003). The structure of DNAin the nucleosome core. Nature, 423(6936), 145-150.



