Mini Review Article - (2021) Volume 5, Issue 1
The Five-Steps Rule and Its Impacts on Molecular Biology
Received Date: Apr 23, 2021 / Accepted Date: Apr 27, 2021 / Published Date: Apr 30, 2021
Copyright: ©Copyright: �?©2021 Kuo-Chen Chou. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Kuo-Chen Chou (2021) The Five-Steps Rule and Its Impacts on Molecular Biology, Stem Cell Res Int 5(1): 01-02
Abstract
In 2011 the 5-steps rule [1] was proposed. Ever since then, it has been stimulating the development in many areas of molecular biology (see e.g, [1-46]), indicating the “5-steps rule” is extremely important. It can be schematically expressed in Figure 1.
Introduction
In 2011 the 5-steps rule [1] was proposed. Ever since then, it has been stimulating the development in many areas of molecular bi- ology (see e.g, [1-46]), indicating the “5-steps rule” is extremely important. It can be schematically expressed in Figure 1.
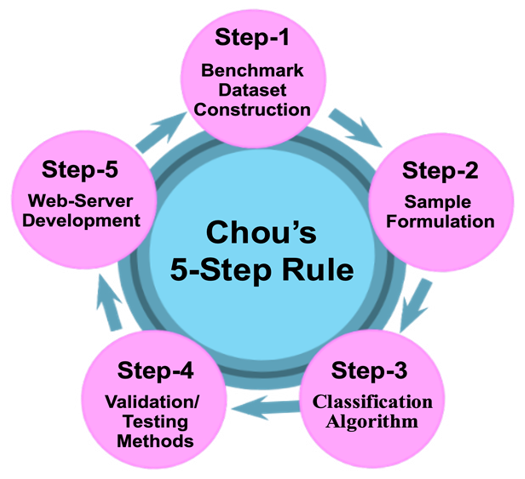
Figure 1: Adapted from International Academic & Research Con- sortium with permission.
References
- KC Chou (2011) Some remarks on protein attribute prediction and pseudo amino acid composition (50th Anniversary Year Review, 5-steps rule), J. Theor. Biol. 273: 236-247.
- O Barukab, YD Khan, SA Khan, KC Chou (2019) iSulfo- Tyr-PseAAC: Identify tyrosine sulfation sites by incorporat- ing statistical moments via Chou’s 5-steps rule and pseudo components Current Genomics, 20: 306-320.
- Y Chen, X Fan (2019) Use Chou’s 5-Steps Rule to Reveal Ac- tive Compound and Mechanism of Shuangsheng Pingfei San on Idiopathic Pulmonary Fibrosis, Current Molecular Medi-cine. 19: 511-563.
- KC Chou (2019) Recent progresses in predicting protein sub- cellular localization with artificial intelligence tools devel- oped via the 5-steps rule, Medicinal Chemistry.
- KC Chou (2019) Impacts of pseudo amino acid components and 5-steps rule to proteomics and proteome analysis, Current Topics in Medicinak Chemistry (CTMC) (Special Issue ed. G.P Zhou), 19: 2283-2300.
- KC Chou (2019) Artificial intelligence (AI) tools constructed via the 5-steps rule for predicting post-translational modifica- tions, Trends in Artificial Inttelengence (TIA). 3: 60-74.
- KC Chou (2019) Recent Progresses in Predicting Protein Sub- cellular Localization with Artificial Intelligence (AI) Tools Developed Via the 5-Steps Rule, Japanese Journal of Gastro- enterology and Hepatology. 2: 1-4.
- KC Chou (2019) An Insightful 10-year Recollection Since the Emergence of the 5-steps Rule, Current Pharmaceutical De- sign, 25: 4223-4234.
- X Du, Y Diao, H Liu, S Li (2019) MsDBP: Exploring DNA-binding Proteins by Integrating Multi-scale Sequence Information via Chou’s 5-steps Rule, Journal of Proteome Re- search. 18: 3119-3132.
- A Dutta, A Dalmia, A R, KK Singh, A Anand (2019) Using the Chou’s 5-steps rule to predict splice junctions with interpreta- ble bidirectional long short-term memory networks, Comput Biol Med. 116: 103558.
- W Hussain, SD Khan, N Rasool, SA Khan, KC Chou et al (2019) SPalmitoylC-PseAAC: A sequence-based model de- veloped via Chou’s 5-steps rule and general PseAAC for iden- tifying S-palmitoylation sites in proteins, Anal. Biochem. 568: 14-23.
- W Hussain, YD Khan, N Rasool, SA Khan, KC Chou et al (2019) SPrenylC-PseAAC: A sequence-based model devel- oped via Chou’s 5-steps rule and general PseAAC for identi- fying S-prenylation sites in proteins, J. Theor. Biol. 468: 1-11.
- S Ilyas, W Hussain, A Ashraf, YD Khan, KC Chou et al (2019) iMethylK-PseAAC: Improving accuracy for lysine methyla- tion sites identification by incorporating statistical moments and position relative features into general PseAAC via Chou’s 5-steps rule, Current Genomics.
- Z Jun, SY Wang (2019) Identify Lysine Neddylation Sites Us- ing Bi-profile Bayes Feature Extraction via the Chou’s 5-steps Rule and General Pseudo Components, Current Genomics, 20: 592-601.
- S Khan, M Khan, N Iqbal, T Hussain,KC Chou et al (2019) A Two-Level Computation Model Based on Deep Learning Al- gorithm for Identification of piRNA and Their Functions via Chou’s 5-Steps Rule Human Genetics. 19: 756-799.
- J Lan, J Liu, C Liao, DJ Merkler, Q Han (2019) A Study for Therapeutic Treatment against Parkinson’s Disease via Chou’s 5-steps Rule, Current Topics in Medicinal Chemistry. 19: 2318-2333.
- R Liang, J Xie, C Zhang, M Zhang, H Huang et al (2019) Identifying Cancer Targets Based on Machine Learning Meth- ods via Chou’s 5-steps Rule and General Pseudo Components, Current Topics in Medicnal Chemistry. 19: 2301-2317.
- Y Liang, S Zhang (2019) Identifying DNase I hypersensitive sites using multi-features fusion and F-score features selec- tion via Chou’s 5-steps rule, Biophys Chem. 253:106227.
- A Wiktorowicz, A Wit, A Dziewierz, L Rzeszutko, D Dudek et al (2019) Calcium Pattern Assessment in Patients with Se- vere Aortic Stenosis Via the Chou’s 5-Steps Rule, Current Pharmaceutical Design. 25: 6-31.
- L Yang, Y Lv, S Wang, Q Zhang, Y Pan et al (2019) Identi- fying FL11 subtype by characterizing tumor immune micro- environment in prostate adenocarcinoma via Chou’s 5-steps rule, Genomics. 112: 1500-1515.
- MA Akmal, W Hussain, N Rasool, YD Khan, KC Chou et al (2020) Using Chou’s 5-steps rule to predict O-linked ser- ine glycosylation sites by blending position relative features and statistical moment, IEEE/ACM Trans Comput Biol Bio- inform, PP.
- H Bouziane, A Chouarfia (2020) Use of Chou’s 5-steps rule to predict the subcellular localization of gram-negative and gram-positive bacterial proteins by multi-label learning based on gene ontology annotation and profile alignment, J Integr Bioinform.
- P Charoenkwan, N Schaduangrat, C Nantasenamat, T Pia- cham, W Shoombuatong (2020) iQSP: A Sequence-Based Tool for the Prediction and Analysis of Quorum Sensing Pep- tides via Chou’s 5-Steps Rule and Informative Physicochemi- cal Properties, Int. J. Mol. Sci. 21: 75.
- Y Chen, X Fan (2020) Use of Chou’s 5-Steps Rule to Reveal Active Compound and Mechanism of Shuangshen Pingfei San on Idiopathic Pulmonary Fibrosis, Curr Mol Med. 20: 220-230.
- KC Chou (2020) Other Mountain Stones Can Attack Jade: The 5-Steps Rule, Natural Science. 12: 59-64.
- KC Chou, Proposing 5-Steps Rule Is a Notable Milestone for Studying Molecular Biology, Natural Science. 12: 74-79.
- KC Chou (2020) The Significant and Profound Impacts of Chou’s 5-Steps Rule, Natural Science. 12: 633-637.
- KC Chou (2020) Analyze the Role of the “5-Steps Rule” Guidelines in Stimulating the Drug Development (Short Communication), Scholarly Journal of Food and Nutrition (SJFN). 3: 385-386.
- L Du, Q Meng, H Jiang, Y Li (2020) Using Evolutionary In- formation and Multi-Label Linear Discriminant Analysis to Predict the Subcellular Location of Multi-Site Bacterial Pro-teins via Chou’s 5-Steps Rule, IEEE Access. 8: 56452-56461.
- Z Ju, SY Wang (2020) Prediction of lysine formylation sites using the composition of k-spaced amino acid pairs via Chou’s 5-steps rule and general pseudo components, Genomics. 112: 859-866.
- M Kabir, S Ahmad, M Iqbal, M Hayat (2020) iNR-2L: A two-level sequence-based predictor developed via Chou’s 5-steps rule and general PseAAC for identifying nuclear re- ceptors and their families, Genomics. 112: 276-285.
- C KoyunoÃÂ??lu (2020) Use Chou’s 5-Steps Rule to Reveal Why SARS+ MERS= COVID-19, J Biochem Analyt Stud.
- W Lin, X Xiao, W Qiu, KC Chou (2020) Use Chou’s 5-Steps Rule to Predict Remote Homology Proteins by Merging Grey Incidence Analysis and Domain Similarity Analysis, Natural Science 12: 181-198.
- D Nguyen, T Ho-Quang, L Nguyen Quoc Khanh, V Dinh- Phan, YY Ou (2020) Use Chou’s 5-steps rule with different word embedding types to boost performance of electron trans- port protein prediction model, IEEE/ACM Trans Comput Biol Bioinform, PP.
- RP Pandey, S Kumar, S Ahmad, A Vibhuti, VS Raj et al (2020) Use Chou’s 5-steps rule to evaluate protective efficacy in- duced by antigenic proteins of Mycobacterium tuberculosis encapsulated in chitosan nanoparticles, Life Sci. 256: 117961.
- T Roy, P Bhattacharjee (2020) A LabVIEW-based real-time modeling approach via Chou’s 5-steps rule for detection of abnormalities in cancer cells, Gene Reports.100788.
- H Vundavilli, A Datta, C Sima, J Hua, R Lopes et al (2020) Using Chou’s 5-steps rule to Model Feedback in Lung Cancer IEEE Journal of Biomedical and Health Informatics, 21: 1-24.
- L Yang, Y Lv, S Wang, Q Zhang, Y Pan et al (2020) Identi- fying FL11 subtype by characterizing tumor immune micro- environment in prostate adenocarcinoma via Chou’s 5-steps rule, Genomics. 112:1500-1515.
- S Zhang, T Xue (2020) Use Chou’s 5-steps rule to identify DNase I hypersensitive sites via dinucleotide property matrix and extreme gradient boosting, Molecular Genetics and Ge- nomics. 295.
- Z Zhang, L Wang (2020) Using Chou’s 5-steps rule to identi- fy N(6)-methyladenine sites by ensemble learning combined with multiple feature extraction methods, J. Biomol. Struct. Dyn. 1-11.
- XF Zhao, Z Min, X Wei, Y Ju (2020) Using the Chou’s 5-steps rule, Transient Overexpression Technique, Subcellular Loca- tion, and Bioinformatic Analysis to verify the Function of Vitis vinifera O-methyltranferase 3 (VvOMT3) Protein, Plant Physiology and Biochemistry 151: 621-629.
- MKM Asma Ehsan 1, Yaser Daanial Khan3, Yu-Ming Chu4’5, Kuo-Chen Chou6 (2021) PredISCs: Using Chou’s 5-Steps Rules to Predict Iron-Sulfur (2Fe-2S) Cluster Binding Sites in Protein. 1-15.
- H Wang, Y Ding, J Tang, Q Zou, F Guo (2021) Identify RNA-associated subcellular localizations based on multi-la- bel learning using Chou’s 5-steps rule, BMC Genomics, 22: 22-56.


