Review Article - (2022) Volume 7, Issue 1
The Evolving Role of Microrna Subtypes 21, 155 and 10b as Oncogenes and Biomarkers in Breast Cancer in a Decade: A Systematic Review Article
2Institute of Endemic diseases, University of Khartoum, Sudan
Received Date: Jan 06, 2022 / Accepted Date: Jan 12, 2022 / Published Date: Jan 18, 2022
Copyright: ©Mubarak Elnour, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Mubarak Elnour, Nagham Eltayeb, Arwa Esam, Shiraz Faisal, Nuha M Elhassan. (2022). The Evolving Role of Microrna Subtypes 21, 155 and 10b as Oncogenes and Biomarkers in Breast Cancer in a Decade: A Systematic Review Article. Int J Women's Health Care, 7(1), 01-05.
Abstract
Breast cancer is a malignant disease with a high incidence worldwide in women- and to less extent in men-, thus remains a health burden. miRNA plays a key role in the development and prognosis of breast cancer; as a single miRNA has the ability to regulate multiple gene targets and adversely alter their expression. In this review, we attempt to systematically analyze the rapidly accumulating body of research regarding the role of miRNAs subtypes particularly as oncogenes and biomarkers in breast cancer, and their clinical implications. A total of 11 pertinent articles in the past 10 years were extracted from high-rank reviews, analyzed, and summarized. Amongst many (eight miRNA subtypes), three subtypes (miR21, 155, and 10b) were selected in order to evaluate the validity of their oncogenicity as well as their diagnostic value as biomarkers.
Keywords
MicroRNA (miRNA), Breast cancer (BC), Oncomir
Introduction
Breast Cancer
According to the American Society, breast cancer is the most common cancer among women [1]. It is a highly heterogeneous complex neoplastic disease with diverse intertumoral and intra-tumoral non-uniformity that occurs due to abnormal proliferation of cells or tissues lining the mammary glands and ducts. Various factors play a role in the development and promotion of breast cancer by changes in the expression of certain genes, amongst which the noncoding RNAs (miRNAs) are worth mentioning [f2]. Breast cancer can be classified based on its unique phenotype, tu¬mor grade, and molecular characterization [f1]. It has been reported that the differential expression of miRNAs in breast cancer. Conse¬quently, an increasing number of studies have suggested that miR-NAs are closely associated with BC occurrence and development. Undoubtedly, TNM staging is of great prognostic value. Howev-er, considering all the limitations of the currently available prog-nostic strategies, it is generally recognized that new, affordable, more accurate methods indicative of molecular characteristics of tumors are needed to achieve personalized treatment. It remains difficult to achieve these goals, because of the absence of refined (sensitive and specific) biomarkers for disease monitoring and for addressing breast cancer on an individual basis [3]. MicroRNAs are small single-stranded, non-coding RNA molecules that span 19 to 25 nucleotides in length [4]. The first discovery of microR- NAs (miRNA), lin-4, in 1993 by the Ambros and Ruvkun groups in Caenorhabditis elegans, has developed the field of molecular biology [5]. miRNAs are transcribed from DNA sequences into primary miRNAs (primiRNAs) and processed into precursor miR-NAs (pre-miRNAs) and mature miRNAs. In most cases, miRNAs interact with the 3′ UTR of target mRNAs to interrupt gene expres¬sion [5]. miRNAs are critical for post-transcriptional gene regula¬tors of various biological functions. They regulate ample cellular and signaling pathways, including cell development and differen¬tiation, cell proliferation and apoptosis as well as cell migration [6]. It is believed that miRNA genes constitute about 1–2% of the known genes in eukaryotes. miRNAs also downregulate gene ex¬pression by binding to their specific target mRNAs 3′-untranslat-ed regions (3′UTRs) leading to either mRNA degradation, trans¬lational repression, mRNA decay and deadenylation depending on the degree of complementarity with target mRNA sequences. Many genuine studies on miRNA have highlighted their role in the etiopathogenesis of a variety of human diseases, such as neurode¬generative disorders, diabetes, cardiac hypertrophy, and respirato¬ry diseases, and there is a long history of regulatory relationships between miRNAs and cancer hallmarks, particularly breast cancer. miRNAs have been reported to play many important roles as onco¬genes (oncomiRNAs), tumor suppressors and biomarkers. MiRNA abnormal expressions are also considered to be possible biomark¬ers of BC because they are stably found in tumor tissues as well as in patients’ body fluids such as blood, plasma, and saliva. miRNAs in body fluids, also called circulating miRNAs, are remarkably stable, packaged into extracellular microparticles or bound with lipoproteins, which protect them from RNase digestion. In this re-view, we attempt to scientifically analyze the rapidly accumulating research regarding the emerging role of miRNAs subtypes, par¬ticularly as oncogenes and biomarkers in breast cancer, and their clinical implications [6].
Biogenesis and Maturation
Winter et al viewed that the biogenesis of mature miRNA in-volves a series of biological processes. Firstly, the primary miR-NA (pri-miRNA) is transcribed by RNA polymerase II or III in the nucleus, forming an elongated RNA hairpin structure that is subsequently cleaved by Drosha into a small stem-loop structure of~70 nt, (pre-miRNA). The pre-miRNA is exported from the nu-cleus into the cytoplasm by exportin-5 and the loop is cleaved after the pre-miRNA is loaded onto Dicer, producing a double-stranded structure of miRNA and antisense miRNA. The latter is normal¬ly degraded, however the long (~22 nt) mature miRNA strand is integrated into the miRNA-induced silencing complex (mRISC), resulting in mRNA degradation or translational repression. Mature miRNA levels are regulated via binding to ceRNAs such as circu-lar (c) RNAs, pseudogenes, and lncRNAs, which act as sponges to prevent miRNA binding to target mRNA [7]. See (Figure1).
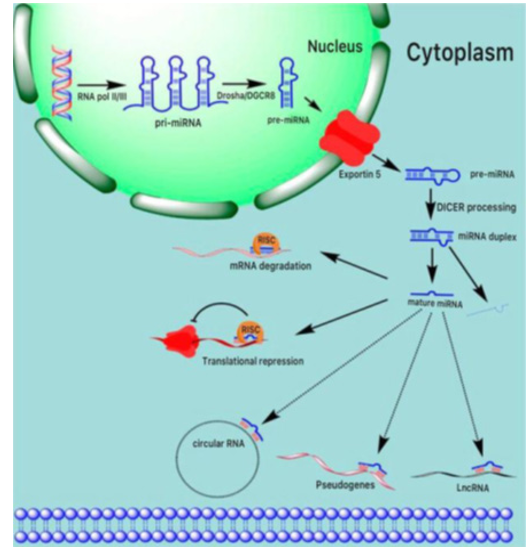
Figure 1: Schematic representation of miRNA biogenesis and function. The primary miRNA transcript (pri-miRNA) is transcribed in the nucleus by RNA polymerase II/III, creating an elongated hairpin structure of RNA that is then split into a small stem-loop structure of ~70 nt by Drosha (pre-miRNA). Pre-miRNA is exported by exportin-5 from the nucleus into the cytoplasm and the loop is cleaved after loading the pre-miRNA onto Dicer, creating a double-stranded miRNA and antisense miRNA* structure. The latter is usually degraded, while in the miRNA-induced silencing complex (mRISC), the long (~22 nt) mature miRNA strand is in¬tegrated, leading to mRNA degradation or translational repression. Via binding to ceRNAs such as circular (c) RNAs, pseudogenes, and lncRNAs, which function as sponge, mature miRNA levels, are regulated to prevent miRNA binding to target mRNAs.
miRNAs subtypes as oncogenes
The first proof of the link between miRNA and cancer derived from studies on B cell chronic Lymphocytic leukemia (CLL), par¬ticularly in an attempt to identify a translocation induced deletion on chromosome 13q14 that is frequently detected in CLL [8]. The mechanisms of oncomiRNA can be explained by a certain number of proteins. Some of these proteins are integral components of the RNA induced silencing complex (RISC) that delivers those small RNAs to complementary sites within mRNA. miRNAs exert their silencing function usually through interactions with the 3’ untrans-lated region (3’ UTR) of the target gene through imperfect base pairing. Moreover, miRNAs can function as master gene regula¬tors, impacting a number of cellular pathways important for nor¬mal cellular functions as well as cancer development and progres¬sion. These alterations culminate in an increased expression of the oncomiR such as miR-21, miR-155 and miR-10b and decreased tumor suppressors such as Let-7a (f2). MiR-10b was shown to be upregulated in breast cancer cells. Overexpression of miR10b is a target for the transcription factor Twist, which causes interrup¬tion of homeobox D10 (HOXD10) mRNA translation. This leads to increased expression of Ras homolog gene family member C (RhoC), which promotes cell invasion and metastasis [9]. Similar¬ly, miR-21 plays an important role in many carcinomas, especially breast cancer development and progression. Through increased cell growth in vitro, tumor growth in vivo and inhibited expression of multi-tumor suppressor proteins such as TIMP3, PDCD4 and tropomyosin 1 (alpha). In addition, it has a major role in invasion and metastasis by targeting multiple anti-metastatic genes. Finally, miR-155 acts as an oncogene/upregulate factor in breast cancer cells, resulting in suppressing the expression of protein Suppres¬sor of Cytokine Expression 1 (SOCS1) both in vitro and in vivo. Pro-tumorigenesis is inversely associated to SOCS1 inhibition. The miR-155 also represses tumor suppressor for head box O3a expression in breast cancer [10].
miRNAs as Biomarkers
Lawrie et al. conducted the first study of miRNA quantification in serum. Who found that serum levels of miR-21were associat¬ed with relapse-free survival in patients with diffuse large B-cell lymphoma. After that, several studies have assessed the potential biomarker use of serum or plasma miRNAs in breast cancer and other different types of cancer. Current challenges in the manage¬ment of breast cancer include a continuing search for sensitive, minimally invasive markers that can be exploited to detect early neoplastic changes, thus facilitating the detection of breast cancer at an early stage, as well as monitoring the progress of patients with breast cancer and their response to treatments. MicroRNAs (miRNAs or miRs) have been proposed as promising biomarkers in breast cancer because they can be easily detected in tumor bi-opsies (non-circulating miRNAs) as well as blood, plasma, serum, and saliva (circulating miRNAs). Furthermore, circulating miR-NAs bind to lipoproteins such as HDL, are linked to the Argonaute 2 (Ago2) protein, or are bundled into exosome-like microparticles, microvesicles, and apoptotic bodies.There is also a great need for the identification of sensitive, reliable, and acceptable markers of response to neoadjuvant and adjuvant therapies [11].
Method
A search of PubMed Medline has yielded thousands of papers. The search databases were filtered to identify articles published from 2010 to 2020 using the following search strategy: (microRNA OR miRNA) AND (oncogenes and biomarkers) AND (breast cancer). This systematic review was conducted in accordance with the PRISMA guidelines. The Real Time RT-PCR technique was used to identify the miRNA expression level of selected miRNAs. In addition, the statistical analysis of miRNA expression profiles of the BC and its stages was performed using the program SPSS. For the inferential analysis of data, parametric data from the T-test and non-parametric data from the Mann-Whitney method were used. A P value of less than 0.05 is considered as a significant value.
Results
The search strategy retrieved 48 articles. 16 of these articles were duplicates, 6 were not freely available, 2 were not in English, 11 were irrelevant, 2 were not pertinent, 1 was earlier than 2010, and 11 were eligible. The above-mentioned steps concerning the se-lection of studies are illustrated in detail in (Figure2). Therefore, a total of 11 articles were included in this systematic review, and the role of 3 miRNAs molecules as oncogenes & biomarkers is described in (Table1). According to the results presented in Table1, 81.8% of published studies have not taken into account statisti¬cal analysis to assess the role of miRNAs in breast cancer. Since (45.4%) of the studies focused on oncogenic miRNAs, and their associated genes in BC, 27.3% of their main goal is breast can¬cer, its classification, and tumor grades, 9.1% of the articles ad¬dressed miRNAs as biomarkers, and the rest studied the processes of miRNAs biogenesis), the remaining 18.2% were quantitatively analyzed for the role of miRNAs as oncogenes and biomarkers in breast cancer.
Expression levels of miR-10b were correlated with breast cancer tumor stages. Generally, the miR-10b expression levels are sig¬nificantly elevated when the tumor grade is increased (p-value ≤ 0.05). There were no statistically significant changes observed be¬tween Stage II and III (p-value ≥ 0.05). Stage III miR-10b expres¬sion level is significantly lower than Stage IV (p-value ≤ 0.05). Significant expression level changes were also observed between stages I-II and stages II-IV (p-value ≤ 0.05).
The expression levels of circulating miR-21 in breast cancer pa¬tients were significantly higher (p-value ≤ 0.05). There was no significant difference in the histopathological level of miR-21 expression in patients with breast cancer Grade II and Grade III differentiation, nor Grade III and Grade IV (p-value ≥ 0.05). In addition, there was no significant difference in the expression of miR-21 according to tumor sizes and breast cancer subtypes. Cir¬culating miR-155 was significantly higher in patients with tumors larger than 5 cm (p-value ≤ 0.05). MiR-155 expression levels were significantly different according to various tumor grades, and clini¬cal stages (p-value ≤ 0.05). No significant difference was observed between any of the studied miRNAs regarding lymph node metas¬tasis. Mir-21 was significantly over-expressed in ER-/PR- cases. Specific miRNAs (miR-10b, miR-21, and miR-155) are associated with tumor metastasis and other clinical characteristics for BC, fa¬cilitating the identification of individuals who are at risk.
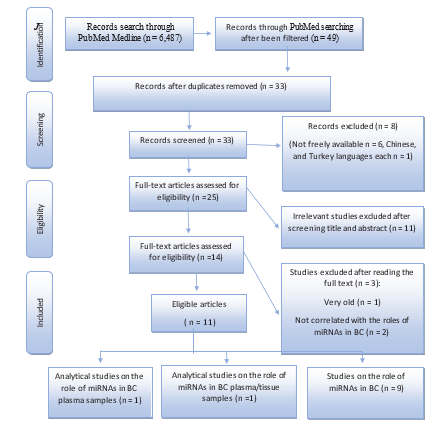
Figure 2: Flow Diagram Represents the Methodology of this Study and its Process
Table 1: List of role of miRNAs in Breast Cancer
|
microRNA subtype |
Receptor |
Detection method |
Role |
Target gene |
Biological sample, and Tumor stage |
P- value |
References |
|
miR-10b |
ER+,PR+, and Her- |
qRT-PCR |
Oncogenes, and biomarker |
HOXD10 |
Plasma, Tissue Stage II, III Stage III, IV |
0.5969
0.3689
0.7400
0.0040 |
[12]
[12]
[13]
[13] |
|
miR-21 |
ER+,PR+, and Her- |
qRT-PCR |
Oncogenes, and biomarker |
MASPIN, TPM1, PDCD4 |
Plasma, Tissue Stage II, III Stage III, IV |
0.7684
0.7516
0.0900
0.5400 |
[12]
[12]
[13]
[13] |
|
miR-155 |
ER+,PR+, and Her- |
qRT-PCR |
Oncogenes, and biomarker |
SOCS1 |
Plasma, Tissue Stage II, III |
0.8804
0.2644
0.0078 |
[12]
[12]
[13] |
Discussion
The expression levels of miR-21, miR-10b, and miR-155 were de-rived from patients’ tissue (breast cancer biopsy) and plasma sam¬ples of BC patients. The results showed a comparable expression pattern among the different samples, with no significant discrepan¬cy. Matamala, et al. represent an exception to this rule [14]. Nev¬ertheless, Svoronos et al. concluded a matched expression level of the miRNAs subtypes between breast tumor tissues and serum. In our analysis, miR-21, and miR-155 were significantly over ex¬pressed in tumor specimens in comparison to the control group. [15]. In addition, the expression of miR-21, miR-155, and some other types of miRNAs was powerfully correlated with the clinico-pathologic features of breast cancer, such as tumor grades and sex hormone receptor expression [15,16]. We also analyzed the results of the previous miRs expression in the plasma to see whether there was a correlation between the expression level of the miRs and the various clinic-pathologic features or not. For example, these studies found the expression of the estrogen receptor (ER) and progesterone receptor (PR) is associated with miR-21 and miR- 155. The ER is recognized as a key factor in motivating mammary cell proliferation, which can lead to up-regulation during the early stages of tumorigenesis and tumor development [17]. Likewise, Al-Khanbashi et al. observed the expression patterns of the miRs related to Her2/neu/ER/PR in breast tumors. In this context, two other independent studies have also shown the expression of miR-21 in breast cancer patients compared to healthy ones is increased.
In addition, the discovered plasma levels of miR-10b and miR-155 between breast cancer patients and the healthy, as well as the serum level of miR-155 in PR+ tumors have exposed a significant dissimilarity in comparison to PR- cases [18]. Wang et al. (2010) explain the clincopathological features of breast tumors such as histological grades and the sex hormone receptor were connected with the expression of miR-21 and miR-155 [19]. Additionally, we have stratified the patients to look at the relationship between the plasma levels of the selected miRs and the stage II/III of the disease based on the TNM staging. In 30 cases of breast cancer, miR-21, miR-10b, and miR-155 did prove a significant difference in the staging compromised control. Furthermore, there was a sig-nificant difference in miR-155 expression in the plasma between stages II and III. Wang et al. (2010) in a comparative between the higher malignancy grades and control breast tissues showed that miR-21, miR-106a, and miR-155 were significantly up regulated in malignancy grades [19]. The comparative expression of MiR-155 expression was not changed in the benign tumors but was in¬creased two fold and five fold in grade II and III. The miR-21 was not changed in the benign tumors but increased 2 fold in the grade II tumors and 4.5 fold in the grade III tumors. Similarly, in both grades, miR-21, miR-126, miR-155, miR-199a, and miR-335 were strongly linked to ER or PR. Additional studies have also shown that miR-21 can up-expression of the oncogene proteins and down expression of the tumor suppressor proteins. Likewise, the miR-21 in breast tumor tissues has been shown to increase expression to be directly linked to the incidence of the disease, tumor size, and staging.
Conclusion
In summary, the expression of miR-21, miR-155, and miR-10b are downregulated in healthy individuals and upregulated in breast cancer. As well, the results showed a similar expression between the tissue and the plasma samples and noted that there were no significant changes. Furthermore, after operations, it was found that the chemotherapy and radiotherapy led to more reductions in the oncomiRs and increased the tumor suppressor genes during treatment. This can be considered as a good diagnostic tool for the process of perfection and suitable response to standard treatment. Generally, the difference between the expressions of the miRs could be due to differences in the sample sources, age, stage of tumor cancer, different analytical methods, and/or different plat¬forms in the studies. Therefore, it seems that an increasing or de¬creasing expression of miRs can be related for various reasons.
References
- Eroles, P., Bosch, A., Pérez-Fidalgo, J. A., & Lluch, A. (2012). Molecular biology in breast cancer: intrinsic subtypes and signaling pathways. Cancer treatment reviews, 38(6), 698-707.
- Singh, R., & Mo, Y. Y. (2013). Role of microRNAs in breast cancer. Cancer biology & therapy, 14(3), 201-212.
- Zografos, E., Zagouri, F., Kalapanida, D., Zakopoulou, R., Kyriazoglou, A., et al. (2019). Prognostic role of microRNAs in breast cancer: A systematic review. Oncotarget, 10(67), 7156.
- Ha, M., & Kim, V. N. (2014). Regulation of microRNA biogenesis. Nature reviews Molecular cell biology, 15(8), 509-524.
- O’Brien, J., Hayder, H., Zayed, Y., & Peng, C. (2018). Overview of microRNA biogenesis, mechanisms of actions, and circulation. Frontiers in endocrinology, 9, 402.
- Yang, Z., & Liu, Z. (2020). The emerging role of microRNAs in breast cancer. Journal of Oncology, 2020.
- Hamam, R., Hamam, D., Alsaleh, K. A., Kassem, M., Zaher, W., et al. (2017). Circulating microRNAs in breast cancer: novel diagnostic and prognostic biomarkers. Cell death & disease, 8(9), e3045-e3045.
- Sonoki, T., Iwanaga, E., Mitsuya, H., & Asou, N. (2005). Insertion of microRNA-125b-1, a human homologue of lin-4, into a rearranged immunoglobulin heavy chain gene locus ina patient with precursor B-cell acute lymphoblastic leukemia.Leukemia, 19(11), 2009-2010.
- Mo, M. H., Chen, L., Fu, Y., Wang, W., & Fu, S. W. (2012).Cell-free circulating miRNA biomarkers in cancer. Journal ofCancer, 3, 432.
- Shah, N. R., & Chen, H. (2014). MicroRNAs in pathogenesis of breast cancer: Implications in diagnosis and treatment. World journal of clinical oncology, 5(2), 48.
- Heneghan, H. M., Miller, N., Lowery, A. J., Sweeney, K. J., & Kerin, M. J. (2009). MicroRNAs as novel biomarkers for breast cancer. Journal of oncology, 2010.
- Khalighfard, S., Alizadeh, A. M., Irani, S., & Omranipour, R. (2018). Plasma miR-21, miR-155, miR-10b, and Let-7a as the potential biomarkers for the monitoring of breast cancer patients. Scientific reports, 8(1), 1-11.
- Zhang, J., Jiang, C., Shi, X., Yu, H., Lin, H., et al. (2016). Diagnostic value of circulating miR-155, miR-21, and miR-10b as promising biomarkers in human breast cancer. Int J Clin Exp Med, 9, 10258-10265.
- Matamala, N., Vargas, M. T., Gonzalez-Campora, R., Mi-nambres, R., Arias, J. I., et al. (2015). Tumor microRNA expression profiling identifies circulating microRNAs for early breast cancer detection. Clinical chemistry, 61(8), 1098-1106.
- Svoronos, A. A., Engelman, D. M., & Slack, F. J. (2016). On-comiR or tumor suppressor? The duplicity of microRNAs in cancer. Cancer research, 76(13), 3666-3670.
- Isanejad, A., Alizadeh, A. M., Shalamzari, S. A., Khodayari, H., Khodayari, S., et al. (2016). MicroRNA-206, let-7a and microRNA-21 pathways involved in the anti-angiogenesis effects of the interval exercise training and hormone therapy in breast cancer. Life sciences, 151, 30-40.
- Han, J. G., Jiang, Y. D., Zhang, C. H., Yang, Y. M., Pang, D., et al. (2017). A novel panel of serum miR-21/miR-155/miR-365 as a potential diagnostic biomarker for breast cancer. Annals of surgical treatment and research, 92(2), 55-66.
- Al-Khanbashi, M., Caramuta, S., Alajmi, A. M., Al-Haddabi, I., Al-Riyami, et al. (2016). Tissue and serum miRNA profile in locally advanced breast cancer (LABC) in response to neo-adjuvant chemotherapy (NAC) treatment. PLoS One, 11(4), e0152032.
- Wang, F., Zheng, Z., Guo, J., & Ding, X. (2010). Correlation and quantitation of microRNA aberrant expression in tissues and sera from patients with breast tumor. Gynecologic oncology, 119(3), 586-593.


