Review Article - (2021) Volume 2, Issue 1
Review of RNA sequence of pX330-U6
Received Date: Nov 27, 2020 / Accepted Date: Dec 26, 2020 / Published Date: Jan 25, 2021
Copyright: ©Copyright: ©2021 Manu Mitra. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Manu Mitra. (2021). Review of RNA sequence of pX330-U6. In j Fore Res, 2(1), 49-56
Abstract
RNA sequencing can be used for studying various factors such as mRNA including single cell gene expression. For more than 10 years RNA sequencing has become vital means for wide analysis of gene expression and differential splicing of mRNAs. Gene manipulation has become more important and sophisticated to study the gene structure of RNA especially when it comes to deadly viruses. In this review paper RNA sequence of pX330-U6 is elaborated based on plasmid structure, its RNA sequence is illustrated from 10 to 8500 bases. Index Terms — pX330-U6, RNA, RNA sequence.
Introduction
Plasmids contain two expression containers, a human codon-optimized SpCas9 and the single guide RNA. Vector can be processed using Bbsl and a pair of annealed oligos can be cloned scarlessly into the vector before the sgRNA scaffold. The oligos are calculated based on the target site sequence (20bp) and essential to be trailed on 3’ end by a 3bp NGG PAM arrangement. One of the applications are cloning grade DNA is applicable for PCR, cloning reactions or transformation into E. coli. Important aspects of these are Genome Editing, expression vectors for cancer modeling, expression vectors for genome editing in the brain [1-3].
RNA SEQUENCE OF PX330-U6
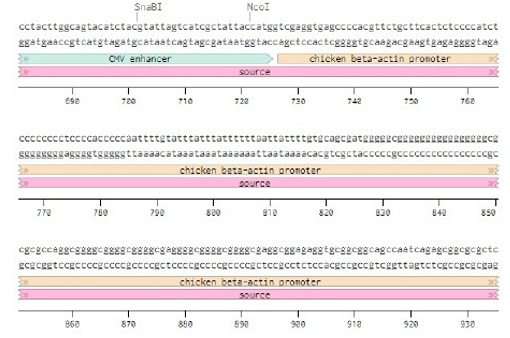
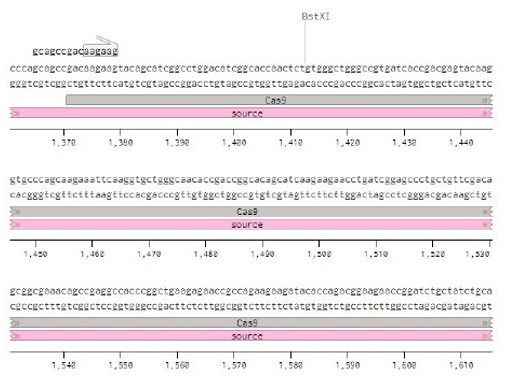
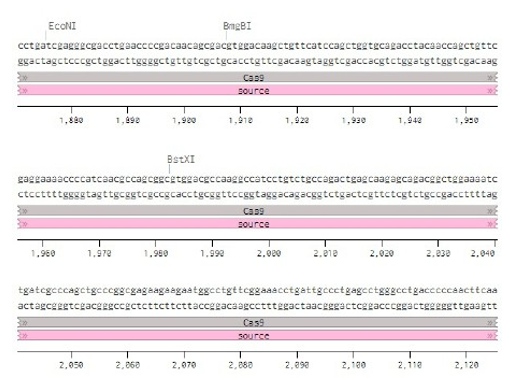
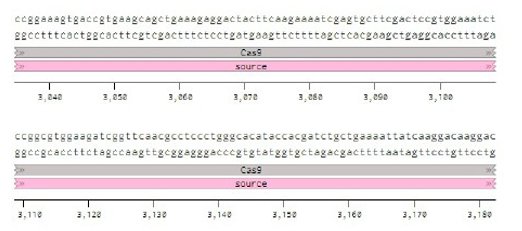
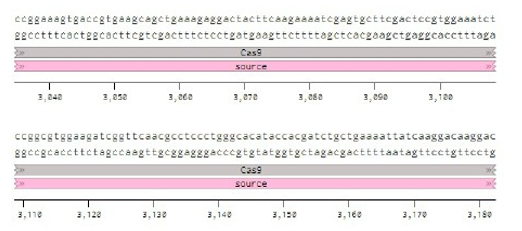
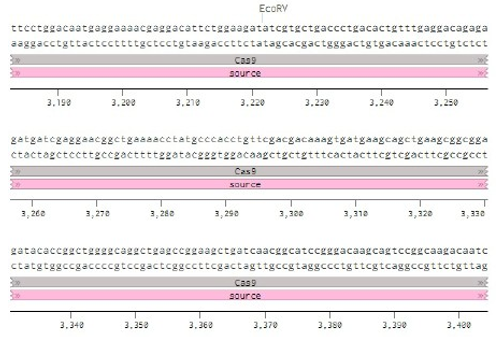
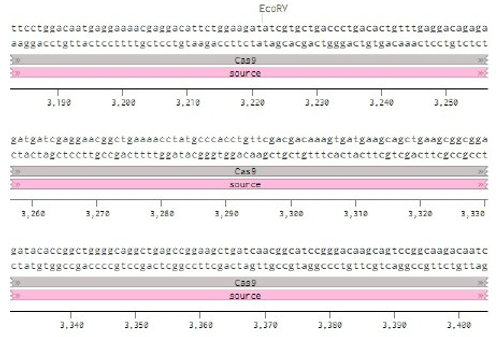
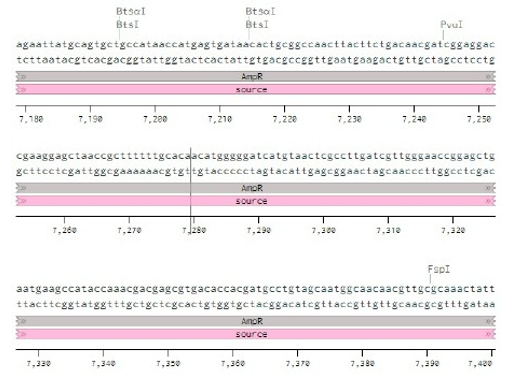
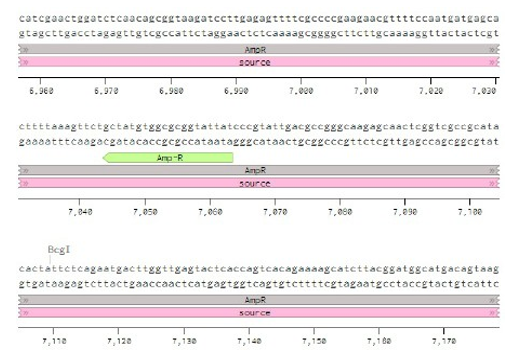
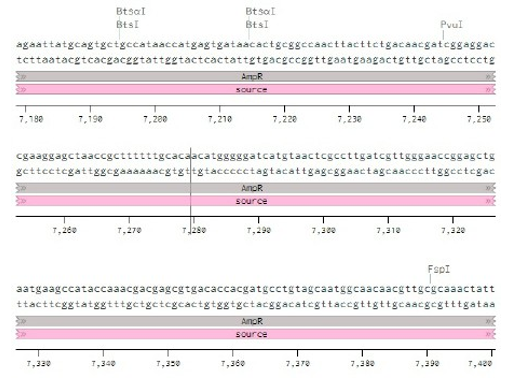
Below is the RNA sequence of pX330-U6 for comprehension of how RNA sequence is distributed [2].
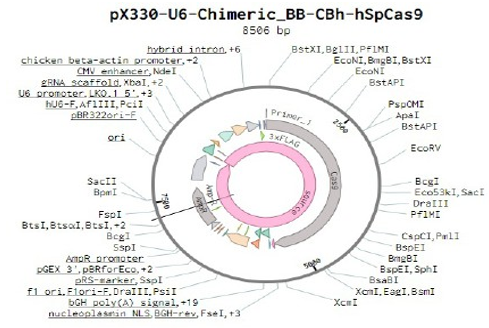

Figure 2: U6 Promotor (10 - 250)




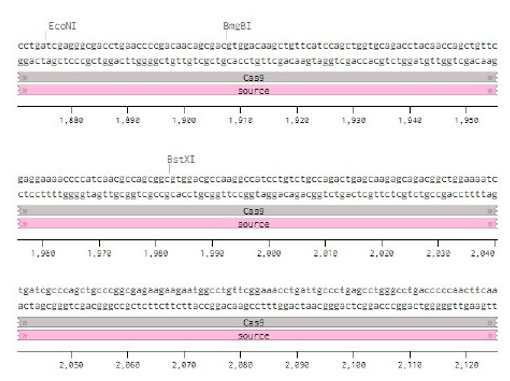
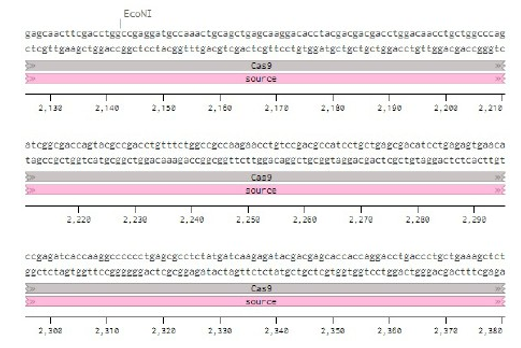
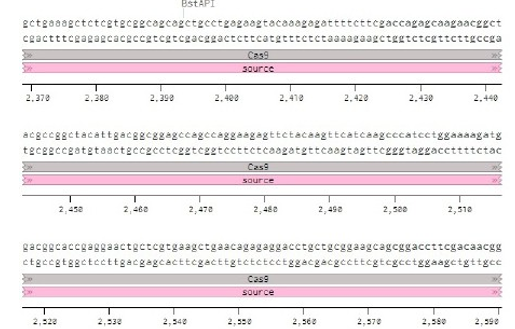
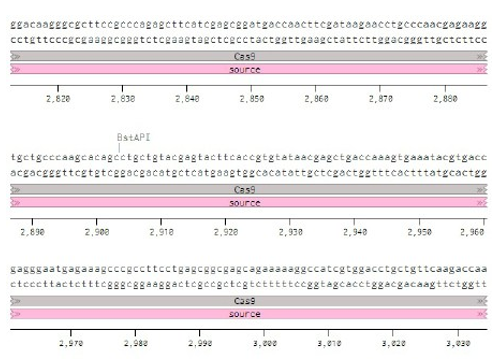
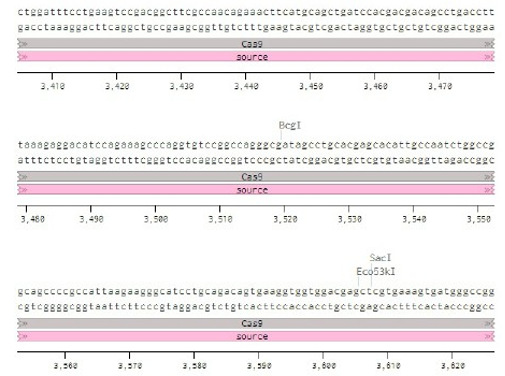
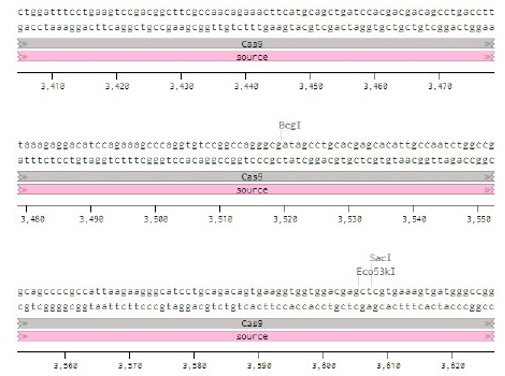
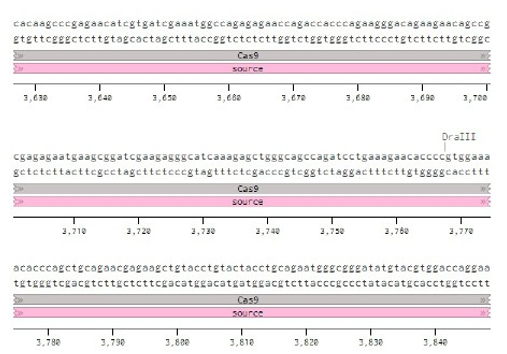
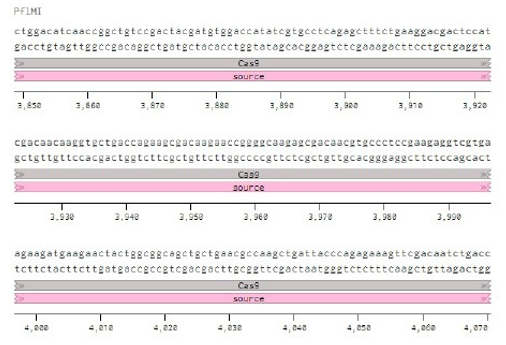
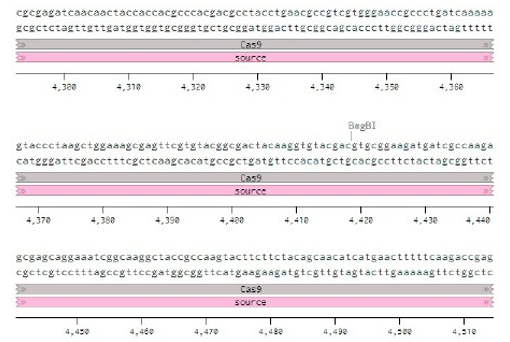
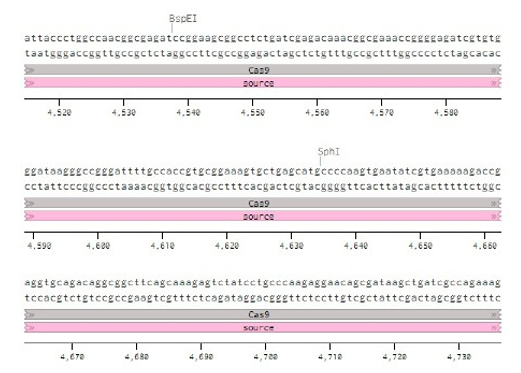
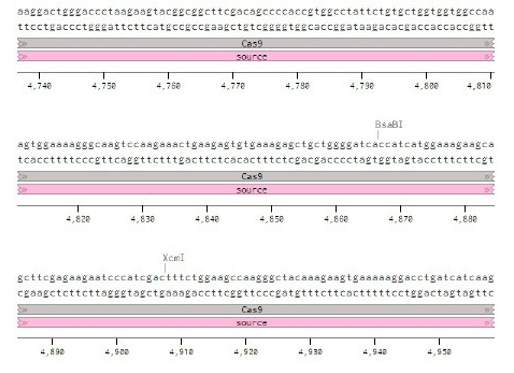

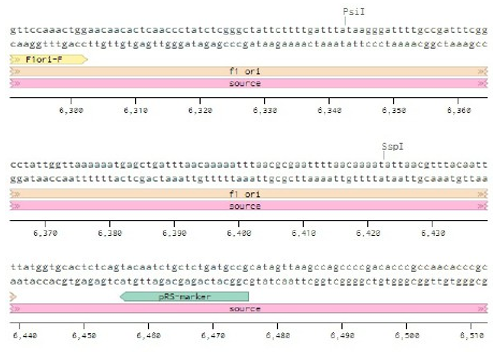

Conflicts of Interest
There is no conflict of interest as per Author’s point of view.
Acknowledgments
Author would like to thank Prof. Navarun Gupta for their academic support. Author also thanks anonymous reviewers for their comments.
Conclusion
Below is the sequence discussed on high level of
1. BstXI, BglII, Pf1MI
2. EcoNI, BmgBI, BstXI
3. EcoNI
4. BstAPI
5. PspOMI
6. ApaI
7. BstAPI
8. EcoRV
9. BcgI
10. Eco53kI, SacI 11. DraIII
12. Pf1MI
13. CspCI, Pm1I
14. BspEI
15. BmgEI
16. BspEI, SphI
17. BsaBI
18. XcmI, EagI, BsmI
19. XcmI
20. nucleoplasmin NLS, BGH-rev, FseI, +3
21. bGH poly(A) signal, +19
22. fl ori, F1ori-F, DraIII, PsiI
23. pRS-marker, SspI
24. pGEX 3’, pBRforEco, +2
25. AmpR promotor
26. SspI
27. BcgI 2
8. BtsI, BtsαI, BtsI, +2 29. FspI
30. BpmI
31. SacII
32. ori
33. pBR322ori-F
34. hU6-F, Af1III, PciI
35. U6 promotor, LK0.15’, +3
36. gRNA scaffold, XbaI, +2
37. CMV enhancer, NdeI 38. chicken beta-actin promotor, +2
39. hybrid intron, +6
References
- Addgene (2020) Addgene: Zhang lab CRISPR. https://www. addgene.org/crispr/zhang/
- Benchling (2019) https://benchling.com/
- pX330-U6-Chimeric_BB-CBh-hSpCas9 was a gift from Feng Zhang (Addgene plasmid # 42230; http://n2t.net/addgene: 42230; RRID: Addgene_42230)


