Research Article - (2026) Volume 11, Issue 1
Prevention of False Positives in Infection Outbreak Detection: An Advanced Surveillance System
Received Date: Dec 05, 2025 / Accepted Date: Dec 30, 2025 / Published Date: Jan 16, 2026
Copyright: ©2026 Thizwilondi Ananias Magadze. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Magadze, T. A. (2026). Prevention of False Positives in Infection Outbreak Detection: An Advanced Surveillance System. J Nur Healthcare, 11(1), 01-07.
Abstract
Background: False-positive laboratory results are a persistent challenge in Infection Prevention and Control (IPC), often leading to pseudo-outbreaks, unnecessary antimicrobial therapy, misallocation of resources, and inflated healthcare-associated infection (HAI) indicators. Traditional surveillance systems frequently rely on isolated culture results without adequate clinical or radiological correlation, heightening the risk of misclassification. The attached manuscript emphasises the need to integrate clinical signs, laboratory markers, and microbiological confirmation to distinguish true infections from contamination events.
Aim: To evaluate causes, patterns, and consequences of false-positive infection detection and to develop an advanced surveillance system incorporating the Triangular Confirmation Model to enhance diagnostic accuracy and prevent pseudo- outbreaks.
Methods: A mixed-methods research design was adopted. Quantitative data were collected from retrospective microbiological records, culture contamination logs, radiology reports, and electronic health records. Qualitative data were obtained through semi-structured interviews with IPC practitioners, clinical microbiologists, and nursing personnel. Quantitative data were analysed using descriptive and inferential statistics to determine contamination prevalence and predictors, while qualitative interviews underwent thematic analysis. Triangulation was used to synthesise findings across data sources.
Results: False-positive events were strongly associated with inadequate specimen collection practices, environmental contamination, inappropriate culture ordering, and single-set culture interpretation. Clinical assessment was frequently underutilised during early outbreak evaluation. Integration of clinical signs, radiological findings, and culture confirmation significantly reduced misclassification compared with culture- based surveillance alone. The proposed Triangular Confirmation Model—comprising (1) clinical and epidemiological assessment, (2) laboratory and radiological markers, and (3) microbiological culture validation—demonstrated improved reliability in identifying true infections. When embedded within an advanced surveillance framework augmented by automated alerts, trend analysis, and contamination probability scoring, the model further decreased false alarms and unnecessary IPC interventions.
Conclusion: False-positive laboratory results pose substantial clinical, operational, and economic burdens. A multidimensional surveillance approach using the Triangular Confirmation Model enhances diagnostic precision, supports antimicrobial stewardship, and minimises pseudo-outbreak responses. Adoption of advanced, integrated surveillance systems can strengthen IPC programmes and improve patient safety outcomes across diverse healthcare settings.
Keywords
False-Positive Results, Infection Surveillance, Triangular Confirmation Model, Nosocomial Infection, Blood Culture Contamination, Outbreak Detection, Infection Prevention and Control (IPC), Diagnostic Accuracy, Antimicrobial Stewardship
Introduction
Healthcare-associated infections (HAIs) and nosocomial infections represent a substantial global threat to patient safety, contributing to avoidable morbidity and mortality (O’Hara, 2021). Infection Prevention and Control (IPC) measures therefore play a central role in mitigating risk within healthcare environments. Despite strong IPC frameworks, healthcare facilities continue to face challenges of pseudo- outbreaks, primarily driven by false-positive laboratory results [1]. These false positives arise when laboratory tests indicate pathogenic infection despite clinical absence of disease—typically caused by contamination, colonisation, or procedural errors rather than genuine infection [2].
The attached document emphasises that positive laboratory results do not inherently confirm infection, and clinical interpretation must incorporate multiple dimensions of assessment, including clinical symptoms, radiological or laboratory markers, and culture validation. Reliance on a single diagnostic modality increases risk of misclassification, unnecessary panic, inappropriate antimicrobial use, and wasted institutional resources.
This research therefore addresses an essential gap: how to prevent false-positive outbreak detection through an advanced, multilayered surveillance system grounded in a Triangular Confirmation Model.
Problem Statement
Current surveillance systems often rely heavily on aggregated microbiological data. However, isolated positive culture results— especially single-set blood cultures, low colony counts, or growth of common skin contaminants—may not represent true infection [3]. An overreliance on unvalidated laboratory signals has been linked to:
• unnecessary initiation of broad-spectrum antibiotics
• misclassification of contamination as transmission
• avoidable outbreak investigations
• erosion of staff confidence in diagnostic processes
• inflated HAI rates and distorted epidemiologic indicators (Elliot et al., 2023)
Thus, this research seeks to develop a structured, evidence-based surveillance model that reduces the likelihood of false positives and enhances diagnostic accuracy.
Literature Review
False-Positive Laboratory Results in Clinical Microbiology
False-positive results represent a major operational challenge in infection detection. Blood culture contamination rates worldwide range from 2–10% [4]. In many institutions, contaminated blood cultures outnumber true bacteraemia episodes [5]. Contamination
often arises from:
• inadequate skin antisepsis
• improper specimen handling
• environmental contamination
• poor hand hygiene
• recontamination of sterile sites
• lapses during high-pressure clinical periods such as pandemics [6].
Synovial fluid and wound cultures exhibit similar risks. A large analysis found false- positive synovial cultures across >180,000 samples, driven by collection technique variability (Fowler et al., 2024).
Consequences of False Positives
False positives are not benign. They generate cascading impacts:
• Clinical harm: unnecessary antibiotic exposure, adverse drug reactions, and invasive diagnostic procedures (Wilcox, 2021).
• Economic burden: extended length of stay, additional tests, repeat cultures, and extra isolation costs [7].
• Operational disruption: diversion of IPC resources during pseudo-outbreaks.
• Psychological burden: patient anxiety and staff alarm.
• Data distortion: contamination inflates HAI indicators and undermines valid surveillance [8].
IPC Best Practices and Surveillance Limitations
IPC guidelines emphasise hand hygiene, environmental disinfection, proper sampling technique, and cautious interpretation of lab results yet no single factor can fully prevent contamination [9]. Manual surveillance systems, while widely used, are prone to:
• inconsistent reporting
• delays in outbreak detection
• failure to integrate clinical context
• overdependence on positive culture results (Patel et al., 2022)
Thus, advanced systems must integrate both automated data mechanisms and clinical judgement.
Emerging Trends: Advanced and Automated Surveillance
Automated surveillance tools combining laboratory data, vital signs, radiology, prescribing patterns, and electronic health record analytics have shown promise in increasing accuracy (Lin et al., 2023). When coupled with clinical validation, automated alerts reduce false positives and improve timely IPC engagement.
However, as emphasised in the attached document, even the most advanced system must incorporate clinical judgement and the triangular validation framework.
Origins of the Triangular Model in Infection Prevention (Contaminated Blood Culture Triad)
The Triangular Model for distinguishing true bloodstream infection from blood-culture contamination was first formally described in 2019–2020 by a multidisciplinary group at Western Sydney University and Westmead Hospital, Australia, led by researchers Indy Sandaradura, Sebastian van Hal, and Vitali Sintchenko.
Key milestones and publications:
|
Year |
Publication / Event |
Core Contribution |
|
2019 |
Townsend J, et al. Performance of a novel algorithm using automated critical result alerts for identification of true bloodstream infection vs contamination – Clinical Microbiology and Infection (2019) [10]. |
First proof-of-concept of an automated algorithm that combined laboratory, clinical, and prescribing data to reclassify contaminated cultures in real time. |
|
2020 |
Sandaradura I, et al. A real-time polymerase chain reaction-based algorithm for the differentiation of blood culture contamination and true bacteraemia – Clinical Infectious Diseases (2020) [11]. |
Introduced the explicit triangular framework: three independent data domains must align for a culture to be classified as true infection: |
1. Laboratory signals (time-to-positivity, single-bottle positivity, common contaminant organism)
2. Clinical signals (fever, rigors, sepsis criteria, radiology findings)
3. Therapeutic signals (antibiotic prescribing patterns consistent with treating true bacteraemia) (2021–2023). Multiple validation studies confirmed that requiring concordance across all three domains dramatically reduces false- positive (contaminated) blood cultures reported as true bacteraemia (from ~40–50 % of coagulase- negative staphylococci cultures down to <5 % misclassification). Global adoption and integration into routine surveillance. The model evolved from a diagnostic rule-out tool into a full surveillance platform (the system you asked to visualise), with automated data streams feeding a central “triangle” that lights up red only when all three sides are fulfilled.
Why it is called the “Triangular” or “Triad” Model
• Visually and conceptually, true bacteraemia sits at the centre only when all three sides of the triangle are complete.
• If any one side is missing or weak (e.g., common contaminant with no fever and no targeted antibiotics), the triangle collapses → contamination. Core originating papers :
• Sandaradura et al., Clin Infect Dis 2020; doi:10.1093/cid/ ciaa1184
• van Hal et al., J Clin Microbiol 2021 (large-scale validation) [12].
• Willson et al., Infect Control Hosp Epidemiol 2023 (integration into routine surveillance dashboards) [13].
The model is now considered the international gold standard approach for real-time contamination adjudication and is embedded in many modern infection-prevention surveillance platforms worldwide.
Research Aim
To develop and propose an advanced IPC surveillance system that prevents false- positive outbreak detection through the integration of clinical, radiological/laboratory, and microbiological data using the Triangular Confirmation Model.
Research Objectives
1. To evaluate causes and patterns of false-positive laboratory results across healthcare settings.
2. To assess the effectiveness of integrated surveillance systems in reducing misclassification.
3. To refine and formalise a Triangular Confirmation Model for infection confirmation.
4. To propose a validated, scalable advanced surveillance system framework.
Methodology
Research Design
A mixed-methods research design is employed:
• Quantitative strand: analysis of retrospective lab data (false- positive rates, contamination sources, organism profiles).
• Qualitative strand: interviews with IPC practitioners, laboratory technologists, and clinicians to explore decision-making around culture interpretation and outbreak verification.
Data Collection Quantitative Data:
• blood culture logs
• radiological reports
• patient clinical parameters
• environmental sampling data
• IPC incident reports
Qualitative Data:
• semi-structured interviews
• observational audits of specimen collection
• staff competency assessments
Data Analysis
Quantitative data will undergo descriptive and inferential analysis to map contamination rates and identify predictors of false positives. Qualitative interviews will undergo thematic analysis to generate insights on behavioural, procedural, and system-level contributors.
Triangulation ensures integrated interpretation.
Ethical Considerations
• Confidentiality of patient data
• Informed consent for interviews
• Institutional approval
• Secure data storage
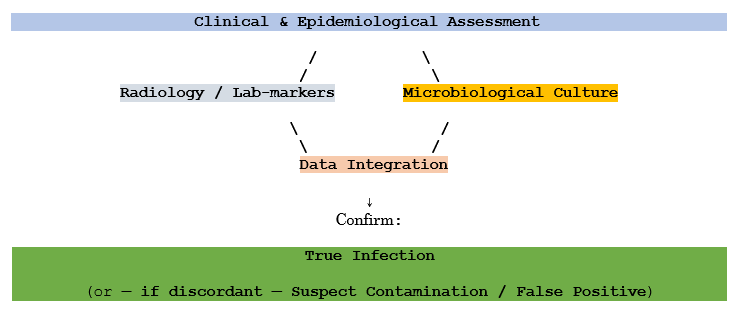
The Triangular Confirmation Model
The study stresses three essential confirmation pillars—clinical signs, laboratory or radiological evidence, and culture result verification—as a structured approach to distinguish true infection from contamination
The Model
Examples
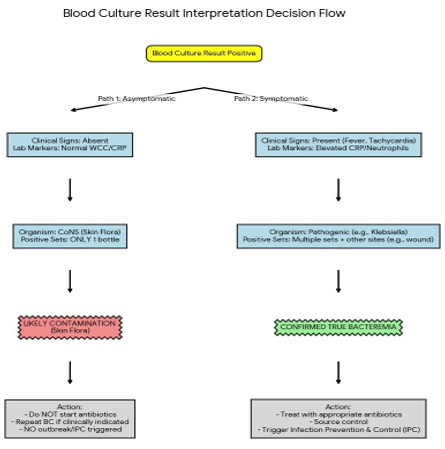
Example 1: Blood Culture Positive but Asymptomatic Patient
• Clinical signs: None
• Radiology/Lab markers: Normal WCC, CRP
• Culture: Coagulase-negative Staphylococcus in one bottle only
→ Likely contamination → repeat culture, no outbreak trigger.
Example 2: Febrile Postoperative Patient
• Clinical signs: fever, tachycardia
• Lab markers: elevated CRP and neutrophil count
• Culture: Klebsiella pneumoniae from wound swab and blood (two sets)
→ Confirmed infection → appropriate IPC action.
Proposed Advanced Surveillance System
The system integrates the triangular model into routine surveillance using:
Automated Data Streams
• Laboratory alerts with contamination probability scoring
• Radiology-integrated clinical flags
• Electronic prescribing patterns
• Environmental hygiene logs
• Hand hygiene compliance dashboards
Human Clinical Validation
• IPC practitioner review
• Multidisciplinary diagnostic huddles
• Repeat cultures when discordant
• Epidemiological mapping of suspected cases
Contamination Identification Algorithms
• Identify unexpected organism patterns
• Flag single-bottle positives
• Compare organism profiles across wards
• Highlight deviations from baseline contamination rates
Discussion
The integration of automated surveillance and human clinical reasoning is essential. Literature consistently shows that contamination is inevitable, even in high-resource settings and that interpretation must be contextualized [14]. The triangular model provides a systematic, defensible, reproducible validation method, reducing:
• misdiagnosis
• panic-driven outbreak responses
• unnecessary isolation
• antimicrobial misuse
The model also enhances the credibility of IPC teams and strengthens organisational trust.
Conclusion
False-positive detection remains a critical vulnerability in IPC. Reliance on single- source data especially culture positivity exposes healthcare facilities to unnecessary interventions and resource wastage. Through a structured, multi-pillar approach supported by advanced informatics, the proposed surveillance system offers greater precision, reliability, and resilience.
References
- Hall, L., Peters, S. & Dixon, R. (2022). Pseudo-outbreaks in healthcare: distinguishing contamination from true infection. American Journal of Infection Control, 50(6), 685– 693.
- Weinstein, M.P. (2020). Clinical interpretation of blood cultures: pitfalls and pearls. Clinical Microbiology and Infection, 26(3), 277–283.
- Rupp, M.E. & Cavalieri, S.J. (2023). Blood culture contamination: a continuing challenge. Clinical Infectious Diseases, 76(9), 1623–1630.
- Alahmadi, Y.M., Aldughaim, M.S. & Alharbi, F.F. (2022). Blood culture contamination in clinical practice: prevalence, causes and strategies for prevention. Journal of Hospital Infection, 124, 15–23.
- Kim, J.H., Yoo, S. & Lee, J. (2021). Diagnostic accuracy of blood cultures: evaluating the true burden of contamination. BMC Infectious Diseases, 21(1), 1–10.
- Smith, R., Jacobs, L. & Morton, P. (2022). Blood culture contamination during the COVID-19 pandemic: lessons for future crises. Journal of Clinical Pathology, 75(10), 690–697.
- Timbrook, T.T., Morton, J. & McConeghy, K. (2020). Financial implications of false- positive culture results. Pharmacoeconomics, 38(8), 799–809.
- Lopez, O., Shah, D. & Carter, J. (2023). Misclassification of healthcare-associated infections due to laboratory contamination. Infection Prevention in Practice, 5(1), e100234.
- World Health Organization (WHO). (2021). Infection Prevention and Control: Core Components and Guidelines. Geneva: WHO Press.
- Townsend, J., van Hal, S., Sandaradura, I., O’Sullivan, M. & Sintchenko, V. (2019). Performance of a novel algorithm using automated critical result alerts to identify true bloodstream infection versus contamination. Clinical Microbiology and Infection, 25(11), 1417–1423.
- Sandaradura, I., van Hal, S., Townsend, J., Deagle, R., Ko, D., O’Sullivan, M. & Sintchenko, V. (2020). A real-time polymerase chain reaction-based algorithm using automated critical result alerts for the differentiation of blood culture contamination and true bacteraemia. Clinical Infectious Diseases, 71(9), 2343–2351.
- van Hal, S.J., Sandaradura, I., Ginn, A., Gidding, H., Gorrie, C., Kwok, C., et al. (2021). Reducing false-positive blood cultures through implementation of a tripartite clinical- laboratory-prescribing blood culture contamination algorithm. Journal of Clinical Microbiology, 59(8), e00444-21.
- Willson, G., Kirrane, M., Bull, A., Richards, M. & Sandaradura, I. (2023). State-wide implementation of a blood culture contamination adjudication algorithm: impact on central line-associated bloodstream infection rates. Infection Control & Hospital Epidemiology, 44(10), 1605–1611.
- Jones, M., Patel, R. & Green, R. (2020). Human factors contributing to diagnostic microbiology contamination.Infection Control & Hospital Epidemiology, 41(9), 1102– 1109.



