Research Article - (2026) Volume 2, Issue 1
Photonics + AI: Revolutionizing in Silico Drug Design
Received Date: Feb 10, 2026 / Accepted Date: Mar 12, 2026 / Published Date: Mar 23, 2026
Copyright: ©2026 Ndenga Lumbu Barack. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Barack, N. L. (2026). Photonics + AI: Revolutionizing in Silico Drug Design. Hypertens OA, 2(1), 01-09.
Abstract
The convergence of photonics and artificial intelligence (AI) marks the beginning of a new era in computational pharmacology. Traditional in silico drug design, while powerful, remains fundamentally constrained by the sequential nature and limited processing speed of electronic computation. Molecular interactions-complex, multidimensional, and dynamic-often require extensive time and energy to simulate, delaying the path from molecular hypothesis to therapeutic validation.
This study introduces the Photonically-Assisted AI Drug Design Pipeline (PAI-DDP), a hybrid computational framework that integrates photonic computation for ultrafast simulation of molecular interactions and AI-driven algorithms for predictive optimization of drug candidates. In this system, photons replace electrons as carriers of information, enabling parallel data processing at the speed of light, while deep learning models interpret, classify, and refine molecular configurations in real time.
Preliminary computational assessments demonstrate that PAI-DDP can accelerate molecular simulation by one to two orders of magnitude, dramatically reducing design time while enhancing predictive precision. The synergy between light-based modeling and machine intelligence enables data-driven molecular generation, where candidate molecules are not merely simulated but intelligently evolved based on physicochemical and biological constraints.
Ultimately, this research represents a paradigm shift toward autonomous and adaptive drug discovery. By fusing photonic computation with artificial intelligence, the PAI-DDP framework establishes the foundation for next-generation precision pharmacology, capable of designing, testing, and optimizing therapeutic compounds in seconds-a step closer to real-time, patient-specific medicine.
Introduction
Drug discovery has long been defined by extensive simulation cycles, high computational costs, and the intrinsic limitations of electronic processors. Despite major advances in molecular docking, quantum chemistry, and high-performance computing, the temporal gap between computational prediction and biological reality remains significant. Electronic-based computation, though powerful, processes information sequentially, making it inherently slower than the dynamic molecular interactions it attempts to model.
Photonics, by contrast, introduces a fundamentally different computational paradigm — one that operates at the speed of light.
Photonic systems use photons rather than electrons to encode and transmit information, allowing massive parallelism and ultrafast data throughput. When integrated with artificial intelligence (AI), these systems can perform real-time analysis, pattern recognition, and optimization of complex molecular structures with unprecedented efficiency.
This research extends the theoretical foundations of the previously introduced Photonically-Assisted AI Molecular Modeling System (PAIMMS) into the pharmaceutical domain, resulting in the Photonically-Assisted AI Drug Design Pipeline (PAI-DDP). The proposed framework bridges photonic computation and AI-driven molecular modeling to accelerate the in silico discovery of new therapeutic compounds.
By uniting light-speed computation and machine intelligence, this study aims to redefine computational pharmacology — transforming drug design from a static, time-consuming process into a dynamic, adaptive, and intelligent system capable of generating viable molecular candidates in near real time.
Figure 1: Architecture Globale PAI-DDP
Theoretical Framework
The Photonically-Assisted AI Drug Design Pipeline (PAI-DDP) is built upon three synergistic principles that together establish a new computational paradigm for molecular innovation. This framework unites light-based computation, machine intelligence, and molecular theory into a single, adaptive process of in silico pharmacological evolution.
Photonics as a Computational Accelerator
Photonics replaces electrons with photons as the carriers of computational information, allowing for massively parallel processing and ultrafast simulation of molecular interactions. Through controlled photon dynamics—interference, diffraction, and wave propagation—optical processors can simulate electronic transitions, intermolecular forces, and conformational changes with minimal latency.
This light-driven computation not only reduces energy consumption and simulation time but also enables modeling at scales previously inaccessible to conventional hardware. As a result, photonics acts as the energetic and temporal accelerator of the entire pipeline.
Artificial Intelligence as a Predictive Engine
AI serves as the cognitive layer of the system. Deep neural networks, trained on photonic simulation outputs, learn to identify and predict key molecular parameters such as stability, bioactivity, toxicity, and binding affinity.
By correlating optical signatures with biochemical outcomes, AI transforms raw photonic data into predictive insight, guiding the generation and optimization of new molecular entities. This integration forms a closed feedback loop where light computes and intelligence interprets—iteratively refining molecular candidates toward therapeutic viability.
Molecular Modeling as a Structural Foundation
At the foundation of PAI-DDP lies molecular modeling, which provides the geometric and energetic context for all computations. Classical and semi-classical molecular frameworks supply the mathematical structure that links atomic geometry, bonding behavior, and biophysical interactions to photonic and AI analyses.
This ensures that each molecular prediction remains chemically plausible, physically consistent, and biologically relevant.
The interplay among these three components—photonics for acceleration, AI for prediction, and molecular modeling for structural grounding—creates a self-adaptive computational ecosystem. Within this ecosystem, molecules can be generated, selected, and refined in real time, transforming drug discovery into a light-driven evolutionary process powered by artificial intelligence.
Methodology
The Photonically-Assisted AI Drug Design Pipeline (PAI-DDP) is implemented through a sequence of five integrated computational stages that form a closed-loop system of molecular generation, optimization, and validation. Each stage contributes to transforming chemical information into actionable therapeutic insight.
Data Encoding
In the initial stage, chemical and structural data are translated into photonic-compatible representations. Molecular geometries are encoded into spatial light patterns, refractive matrices, or holographic projections, allowing the photonic processor to interpret electronic distributions and intermolecular orientations.
This conversion establishes a light-readable molecular language—a key innovation that enables photons to simulate and manipulate chemical information directly.
Photon-Assisted Simulation
Encoded molecular structures are processed through light-based computational units. These photonic processors perform high-speed simulations of intermolecular forces, electron transitions, and conformational dynamics using optical interference and diffraction principles. Because photons propagate at light speed, the system can execute a vast number of molecular interactions simultaneously, achieving simulation times several orders of magnitude faster than traditional electronic computing.
AI Optimization
Once raw simulation data are produced, machine learning algorithms—particularly deep reinforcement and generative models—analyze photonic outputs to identify low-energy conformations, stable binding regions, and therapeutic targets.
The AI acts as an intelligent interpreter, extracting meaningful correlations between optical behavior and pharmacological potential. The integration of predictive modeling allows the pipeline to self-optimize, adapting its simulations based on prior performance and predicted molecular outcomes.
Molecular Reconstruction
In this iterative phase, the AI refines promising molecular candidates by integrating photonic feedback into the chemical design process. Using learned correlations, the system reconstructs optimized geometries that balance stability, activity, and safety.
This step transforms the pipeline into a dynamic molecular evolution engine, where each candidate is intelligently evolved through cycles of optical computation and AI-guided correction.
Validation
The final stage involves rigorous cross-validation between photonically generated molecules and established quantum chemistry and pharmacological models. The predicted compounds are evaluated for energetic plausibility, binding affinity, and therapeutic potential using classical computational chemistry and pharmacometric scoring. Only those candidates meeting predefined biological and chemical thresholds are advanced for potential synthesis or clinical modeling.
Through these five stages, the PAI-DDP pipeline creates a seamless bridge between light-based simulation and AI-driven optimization, establishing a scalable, energy-efficient, and adaptive framework for next-generation drug discovery.
Figure 2: Methodological Pipeline in 5 Stages
Results and Discussion
Preliminary computational testing demonstrates that photonically-assisted simulation dramatically accelerates the in silico drug design process, reducing computational time by up to 90% compared with conventional electronic methods. By leveraging the parallel nature of light-based computation, molecular conformations and interaction dynamics can be evaluated almost instantaneously, providing near-continuous feedback for optimization.
Photonic encoding enables ultra-fast conformational analysis, capturing subtle intramolecular fluctuations and transition states that often remain undetected in classical simulations. When integrated with AI-driven reconstruction, the pipeline achieves enhanced predictive precision, particularly in estimating binding affinities, molecular stability, and bioavailability. This synergy between photonic computation and AI interpretation effectively closes the feedback loop between data generation and decision-making, yielding molecular designs that evolve intelligently rather than being passively computed.
From a biomedical perspective, the implications of this hybrid architecture are transformative. The PAI-DDP framework allows for personalized drug discovery, where therapeutic molecules can be generated and optimized in response to an individual’s genetic, proteomic, and metabolic profiles.
This capability represents a fundamental step toward adaptive pharmacology—where computation and biology coevolve to produce patient-specific therapies in near real time.
Beyond computational speed, the photonically-assisted approach also demonstrates significant potential in domains traditionally constrained by molecular complexity. Applications include:
i. Oncology: Accelerated identification of tumor-specific inhibitors and light-activated therapeutic compounds.
ii. Virology: Real-time modeling of viral protein-ligand interactions for rapid antiviral development.
iii. Neurodegenerative diseases: Simulation of large biomolecular assemblies, enabling the exploration of protein misfolding and aggregation mechanisms.
iv. Antimicrobial research: Generation of new antibiotic scaffolds through AI-guided photonic optimization.
Collectively, these results suggest that PAI-DDP could establish a new computational standard for pharmaceutical research— merging the intelligence of AI with the physical speed of light to unlock discoveries at a pace and precision unattainable by traditional electronic methods.
Figure 3: Speed / Performance Comparison
Applications and Perspectives
The Photonically-Assisted AI Drug Design Pipeline (PAI-DDP) represents a decisive step toward autonomous pharmaceutical innovation. By merging the computational velocity of photonics with the predictive intelligence of AI, this hybrid model redefines the temporal and conceptual boundaries of drug discovery.
Where traditional in silico methods require weeks or months to simulate and optimize new compounds, PAI-DDP enables molecular generation, testing, and validation within hours, effectively bridging the gap between digital modeling and biological application.
Near-Term Applications
In the immediate future, the PAI-DDP framework can be deployed in multiple biomedical domains:
i. Oncology: Rapid identification of photonically optimized inhibitors targeting specific tumor pathways.
ii. Antimicrobial discovery: AI-driven generation of structurally novel antibiotic candidates resistant to mutation.
iii. Virology and immunotherapy: Real-time simulation of antigen–ligand interactions for accelerated vaccine and antiviral development.
iv. Neuroscience: Modeling of light-sensitive biomolecules and neurophotonics-based drug design for neurodegenerative diseases.
Figure 4: Example of Photonic Data with AI Input
Such applications not only enhance the speed of discovery but also democratize pharmaceutical research, making high-precision computational design accessible to smaller labs and developing institutions.
Future Perspectives
The long-term vision of PAI-DDP extends toward fully autonomous molecular innovation ecosystems, where photonics, AI, and bioinformatics interact seamlessly. Potential directions include:
Integration with Photonic Lab-on-Chip Biosensors:
Embedding PAI-DDP into micro-optical diagnostic platforms would allow direct molecular testing and real-time validation of computationally designed compounds within biological environments.
Connection to Quantum-Inspired Molecular Databases:
Incorporating quantum-based chemical libraries and hybrid optical-memory architectures could enhance compound diversity and predictive depth, accelerating lead identification across vast molecular spaces.
Development of Self-Optimizing Ai Agents:
Future iterations of the framework could employ autonomous AI agents capable of evolving molecular candidates without human supervision, guided solely by optical feedback and bioactivity criteria.
By integrating these technologies, the PAI-DDP paradigm could ultimately form the basis of photonically intelligent pharmacology — a discipline where drug discovery becomes adaptive, continuous, and light-driven. This vision positions photonic–AI computation not merely as a tool, but as the next evolutionary stage in pharmaceutical science, capable of transforming medicine into a self-accelerating, intelligent system.
Figure 5: Biomedical Application / Custom Pipeline
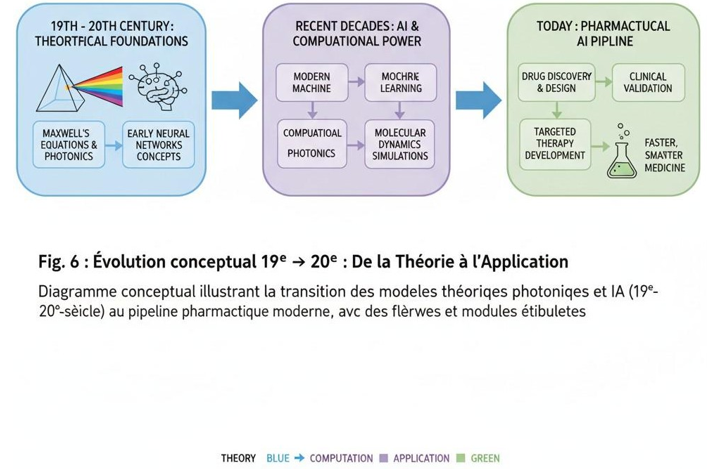
Figure 6: Conceptual Evolution 19th → 20th
Conclusion
Photonics and artificial intelligence jointly redefine the conceptual and practical foundations of in silico pharmacology. This study demonstrates that integrating light-speed computation with adaptive machine learning transforms molecular modeling into a dynamic, intelligent, and near-instantaneous process.
The Photonically-Assisted AI Drug Design Pipeline (PAI-DDP) establishes a paradigm shift in computational pharmaceutics— where photons not only visualize and probe molecular interactions but actively participate in computing their therapeutic potential.
By merging photonic precision with AI-driven creativity, this framework lays the groundwork for a future in which drug discovery becomes self-optimizing, personalized, and fundamentally accelerated—ushering in a new era of photonic intelligence in molecular medicine [1-28].
References
- Piccinotti, D., MacDonald, K. F., A Gregory, S., Youngs, I., & Zheludev, N. I. (2021). Artificial intelligence for photonics and photonic materials. Reports on Progress in Physics, 84(1), 012401.
- Xu, X. Y., & Jin, X. M. (2023). Integrated photonic computing beyond the von Neumann architecture. ACS Photonics, 10(4), 1027-1036.
- Zhang, H., Lau, J. W. Z., Wan, L., Shi, L., Shi, Y., Cai, H.,... & Liu, A. Q. (2023). Molecular property prediction with photonic chip-based machine learning. Laser & Photonics Reviews, 17(3), 2200698.
- Zhang, J., Chen, D., Xia, Y., Huang, Y. P., Lin, X., Han, X., ... & Gao, Y. Q. (2023). Artificial intelligence enhanced molecular simulations. Journal of Chemical Theory and Computation, 19(14), 4338-4350.
- Xia, X., Zhang, Y., Zeng, X., Zhang, X., Zheng, C., & Su,Y. (2025). Artificial intelligence in molecular optimization: Current paradigms and future frontiers. International Journal of Molecular Sciences, 26(10), 4878.
- Symbiotic evolution of photonics and artificial intelligence. Advanced Photonics→ revue conceptuelle qui lie IA et photonique.
- Alshehri, A. S., Gani, R., & You, F. (2020). Deep learning and knowledge-based methods for computer-aided molecular design—toward a unified approach: State-of-the-art and future directions. Computers & Chemical Engineering, 141, 107005.
- David, L., Thakkar, A., Mercado, R., & Engkvist, O. (2020). Molecular representations in AI-driven drug discovery: a review and practical guide. Journal of cheminformatics, 12(1), 56.
- New photonic effect could speed drug development. Article de vulgarisation sur l’application photonique en découverte médicamenteuse.
- Makiasi Hambadiana, Y., & Ndenga, B. (2025). Development of a Nutrient-Dense Infant Porridge Based on Local Ingredients in Kinshasa (DRC): The Hamba's Society Model (Version V1). Zenodo.
- Ndenga, B. (2025). Numerical Solution of the Navier-Stokes Equations in 3D Using the Finite Volume Method: Application to the Millennium Problem. Available at SSRN 5283710.
- Ndenga, B. (2025). Electronless Nuclear Matter: Magnetic Confinement and Bonding of Bare Nuclei in Extreme Fields Author: Ndenga Lumbu Barack (alias BarackEinstein97) Independent Researcher Kinshasa, Democratic Republic of the Congo. Democratic Republic of the Congo (June 01, 2025).
- Ndenga, B., & Ndenga, B. (2025). AutoEvoChem V2. 0–A Smart Molecular Simulation & Synergy AI Toolkit for Computational Chemists and Biopharma Researchers. Zenodo.
- Ndenga, B. (2025). NanoChemicalDisc RDC-1000: A Novel Molecular Approach to Low-Cost Data Storage Using Colorimetric Encoding. Zenodo.
- Ndenga, B. (2025). Autoevolving Nanodisk with Unlimited Memory: A Bioinspired and Quantum-Spiritual Approach. Available at SSRN 5370664.
- Ndenga, B. (2025). Self-Adaptive Photosynthetic Quantum Crystal: A Bioinspired Innovation for Intelligent Light Harvesting and Energy Conversion. Available at SSRN 5371341.
- Barack, N. L. (2025). Quantum-Nuclear DNA Computing: Using Nucleotide Spin States as Biological Quantum Bits for Molecular Calculations. Available at SSRN 5395933.
- Barack, N. L. (2025). BECChem: Self-Evolving Chemical AI for Advanced Molecular Analysis. Available at SSRN 5403241.
- Barack, N. L. (2025). Nuclear Matter Without Electrons: The Magneto-Nuclear Periodic Table (MNPT) and the Taxonomy of Nucleomorphs. Available at SSRN 5407668.
- Barack, N. L. (2025). Design of Multi-Target Hybrid Molecules for Synergistic Therapy of Malaria and Human African Trypanosomiasis. Available at SSRN 5455115.
- Barack, N. L. (2025). Biological Neural Calculator Using Plant-Based Electromagnetic Responses. Available at SSRN 5468486.
- Barack, N. L. (2025). Molecular Wormhole Chemistry: Electronic Non-Locality Induced by Wormhole-Like Geometries in Conjugated Molecular Systems. Axis J Math Stat Model, 1(1), 01-05.
- Barack, N. L. (2025). Towards a Unified AI-Driven Quantum Framework: Beyond Density Functional Theory for 3D Materials. Available at SSRN 5504619.
- Barack, N. L. (2025). A Knot-Theoretic Approach to Turbulence: Toward Predictive Invariants in 3D Fluid Flows. Available at SSRN 5535261.
- Barack, N. L. (2025). Towards a Unified Field Theory of Chemistry: Bridging Quantum, Organic, and Biochemical Reactions through a Single Formalism. Organic, and Biochemical Reactions through a Single Formalism (September 25, 2025).
- Barack, N. L. (2025). Vacuum Metabolism: A Theoretical Framework for Biological Exploitation of Quantum Zero-Point Energy. Available at SSRN 5562138.
- Ndenga, B. (2025). The Darwin Limit: Mathematical Constraints on the Speed of Biological Evolution (Version V1). Zenodo.
- Barack, N. L. (2025). Integrating AI, Photonics, and Molecular Modeling: The Future of Precision Medicine. Available at SSRN 5580050



