Research Article - (2026) Volume 2, Issue 1
Personalized Prediction of Lymph Node Involvement in Head and Neck Squamous Cell Carcinomas Using Mixture Hidden Markov Models Incorporating Tumor
Received Date: Jan 27, 2026 / Accepted Date: Feb 26, 2026 / Published Date: Mar 05, 2026
Copyright: ©2026 Latha Kiran Krishna Rajendran. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Rajendran, L. K. K. (2026). Personalized Prediction of Lymph Node Involvement in Head and Neck Squamous Cell Carcinomas Using Mixture Hidden Markov Models Incorporating Tumor. J Demo Res, 2(1), 01-07.
Abstract
Head and neck squamous cell carcinomas (HNSCC) frequently metastasize to regional lymph nodes, making accurate prediction of lymphatic spread crucial for treatment planning. Current diagnostic imaging techniques fall short in detecting microscopic lymph node metastases, often leading to broad and non-personalized irradiation strategies. In this study, we present an advanced model for predicting the risk of occult nodal disease by integrating the primary tumor location into a hidden Markov model (HMM) framework. We focus on tumors in the oropharynx and oral cavity, incorporating detailed subsites as defined by ICD- 10 codes. By leveraging multi-centric data from over 1,200 patients, we developed a mixture of HMMs that account for the distinct patterns of lymphatic spread observed in different tumor subsites. Our approach enhances the precision of lymph node involvement predictions, potentially allowing for more personalized and targeted radiation therapy. The results indicate that our mixture model outperforms traditional models, especially in cases where lymph node involvement patterns vary significantly across subsites. This work represents a significant step towards personalized treatment planning in HNSCC, with the potential to reduce treatment-related side effects while maintaining therapeutic efficacy.
Keywords
Lymph Node Involvement, Personalized Prediction, Statistical Modeling, Risk Stratification, Bayesian Inference, Predictive Modeling, Lymph Node Metastasis, Head and Neck Squamous Cell Carcinoma
Introduction
Head and neck squamous cell carcinomas (HNSCCs) are a mixed and assorted category of malignancies that originate from the mucosal linings of the upper aerodigestive tract. These cancers exhibit a robust tendency for initial metastasis to local lymph nodes, making the detection and description of nodal participation a foundation in their clinical management [1]. The presence or lack of lymph node metastases substantially influences prognosis, treatment planning, and overall survival. One of the important and significant clinical problems in managing HNSCC lies in correctly identifying patients with occult lymph node metastases; those not detected through traditional imaging or clinical examination.
Despite developments in radiographic techniques such as PET/ CT and MRI, these methods have limitations in sensitivity and specificity, especially for microscopic disease [2,3]. Consequently, several patients undergo prophylactic irradiation of big and substantial neck regions, which can result in overtreatment and increased morbidity. The standard clinical approach to nodal irradiation commonly follows generalized guidelines derived from aggregated population data, such as those developed by collaborative oncology groups or specialist panels [4].
While these guidelines are helpful and beneficial in standardizing care, they frequently fail to consider individual patient attributes that may affect lymphatic spread patterns. As a result, patients may be subjected to unnecessarily broad radiation fields, rising the risk of toxicities like xerostomia, mucositis, and long-term swallowing dysfunction [5]. To address this, researchers have turned to predictive modeling techniques that incorporate patient-specific variables to approximate the risk of nodal metastasis. Among these, Hidden Markov Models (HMMs) have emerged as powerful instruments for modeling probabilistic progression of disease through the lymphatic system [6]. HMMs are especially wellsuited for representing sequential and partly detectable phenomena, such as cancer metastasis, by permitting for the incorporation of both noted and hidden states in a mathematically thorough and meticulous framework.
Prior work has displayed that HMMs can be trained on multi-institutional datasets to approximate the likelihood of lymph node participation in individual patients established on their clinical presentation. These models have demonstrated enhanced predictive accuracy over customary heuristic procedures, notably when used to evaluate clinically negative lymph node levels [7]. However, one important and significant limitation of previous HMM-based models is their incapacity to account for the heterogeneity of tumor subsites within the head and neck region.
Different anatomical subsites; such as the tonsil, foundation of tongue, floor of mouth, and palate; are known to exhibit distinct patterns of lymphatic drainage and metastasis [8].
For instance, tumors originating in the oral cavity frequently follow a dissimilar route of spread compared to those emerging in the oropharynx. These anatomical and pathological differences are not adequately captured when employing a single, generalized HMM for all tumor locations. Recognizing this limitation, new efforts have concentrated on stratifying patients by tumor subsite. However, producing and maintaining separate models for each individual ICD-10 tumor code is not practical and workable due to the need for big and substantial, well-annotated datasets for each category. Moreover, tumor subsites that lie anatomically at the interface between regions; such as tumors of the palate that border both the oral cavity and oropharynx; may share metastatic attributes with numerous and manifold regions, further complicating the modeling effort.
To overcome these problems, we propose a innovative approach that combines tumor subsite information into a mixture of Hidden Markov Models. Instead of making discrete models for each subsite, we utilize a probabilistic framework that denotes each patient as a mixture of numerous and manifold HMM components. This enables us to model continuous variation across tumor subsites and account for shared patterns of lymphatic spread in anatomically neighboring regions. The goal of this study is to enhance the precision of nodal participation prediction by leveraging this mixture-HMM architecture. We incorporate ICD-10 tumor subsite information and train our model employing a huge and massive, multicentric dataset of over 1,200 patients with HNSCC.
By capturing the unique metastatic behavior associated with explicit and defined anatomical regions, our procedure provides the potential to reduce redundant and superfluous radiation exposure while maintaining therapeutic effectiveness. In doing so, this work contributes toward a more personalized approach to head and neck cancer treatment. Our model has the potential to notify clinical decision-making by identifying patients who may safely undergo more cautious and traditional radiation methods established on individualized risk evaluations. Such precision-guided interventions could improve quality of life while maintaining oncologic results.
Background and Related Work
Predicting lymphatic spread in head and neck squamous cell carcinomas (HNSCC) has long presented a notable difficulty to oncologists and radiation planners. The conventional understanding of lymphatic distribution has been derived from big and substantial retrospective surgical series and autopsy studies that mapped frequent and prevalent nodal routes established on tumor site [1,8]. While these studies continue essential, their broad and universal nature frequently lacks detail for individualized risk prediction. The introduction of cross-sectional imaging methods like computed tomography (CT), magnetic resonance imaging (MRI), and positron emission tomography (PET) marked a leap forward in non-invasive nodal staging. However, the diagnostic accuracy of these instruments is still restricted, especially in detecting microscopic or initial metastatic spread [2].
The inherent limitations of imaging make it essential and required to explore computational approaches that can probabilistically infer occult metastases. Recent years have witnessed an rising shift towards the application of machine learning and probabilistic graphical models in oncology. Among these, Hidden Markov Models (HMMs) have proven valuable in modeling temporally developing and partly visible and noticeable biological systems. HMMs are especially well-suited for modeling the spread of disease through interconnected networks; such as the lymphatic system; where changes between states (for instance, healthy to metastatic) are affected by prior conditions and neighboring nodes [6].
Early implementations of predictive models for lymph node participation in HNSCC frequently counted on logistic regression or decision trees applying tumor features, imaging data, and anatomical landmarks as inputs. While educational and enlightening, these models normally treated each lymph node level (LNL) as an independent binary classification problem, therefore disregarding the spatial and anatomical dependencies between nodes. HMMs address this limitation by treating lymphatic progression as a sequential process, permitting the modeling of probabilistic shifts between neighboring lymph node levels. This makes it possible to approximate the likelihood of participation at a downstream node conditioned on the upstream node being beneficial.
The result is a more biologically reasonable and convincing model of tumor dispersed that aligns well with clinical and anatomical knowledge [6]. Notably, Ludwig et al. proposed a data-driven HMM that leveraged multi-center patient datasets to quantify the probability of occult participation in clinically negative lymph node levels. Their work demonstrated that HMMs could surpass traditional models in forecasting true pathological status and guiding optional radiation fields [7]. However, their model applied the identical parameterization across all tumor subsites, implicitly supposing homogeneity in metastatic behavior. This assumption overlooks well-documented clinical evidence showing that tumor subsites within the head and neck exhibit varied and distinct lymphatic drainage patterns. For instance, oral cavity tumors more often include level I and II nodes, while oropharyngeal tumors frequently spread to level II and III [4].
Ignoring these patterns can compromise the specificity of predictive models and restrict their clinical utility. To address this subsite-level variability, some studies have proposed stratifying models by tumor location, training separate models for the oral cavity, oropharynx, hypopharynx, etc. While effective in theory, this plan is frequently unrealistic and unworkable in practice due to the limited sample sizes available for rare subsites, and it fails to model the continuum between anatomical boundaries. A more flexible solution includes employing mixture models, which allow a tumor to be probabilistically represented as a combination of various canonical patterns of lymphatic spread. This approach is especially helpful for tumors positioned near anatomical borders (for instance, the palate), which may exhibit attributes of more than one region.
Mixture HMMs offer a tractable strategy to encode such intricacy and are gaining attention for their potential to bridge anatomical detail with statistical power. In summary, the literature highlights both the commitment and the problems of personalized prediction in HNSCC. HMMs have already proven effective in capturing the dynamics of lymphatic spread, but their complete and entire potential is realized exclusively when integrated with anatomical subsite information. Our work builds upon these advancements by introducing a mixture-HMM framework trained on a huge and massive, varied and distinct patient cohort, permitting individualized prediction that is both anatomically enlightened and aware and statistically strong and durable.
Methodology
A. Modeling Lymphatic Spread with Hidden Markov Models
We adopt a Hidden Markov Model (HMM) framework to capture the probabilistic progression of metastasis across lymph node levels (LNLs) in head and neck squamous cell carcinoma (HNSCC). Each patient is represented by a binary vector X = (X1,X2,...,XV) where Xv ∈ {0,1} indicates the involvement (1) or absence (0) of metastasis at LNL v. The model assumes metastasis evolves over discrete time steps, governed by a transition matrix A and an initial state distribution π, as described in previous work by Ludwig et al. [6].
B. Integrating Tumor Subsite Information with Mixture Models
To address the heterogeneity in lymphatic spread between different anatomical tumor locations, we introduce a mixture model architecture. Rather than training separate HMMs for each ICD-10 subsite, we define a set of M canonical HMMs {H1,...,HM}, each parameterized by a distinct transition matrix θm. For a tumor at subsite s, the model assumes a mixture probability πsm of being generated by HMM component m. This enables representation of tumors in transitional anatomical regions (e.g., the palate) using weighted combinations of canonical spread patterns [9].
C. Parameter Estimation via EM and MCMC
Training the model involves estimating the transition dynamics θm and the subsite-specific mixing coefficients πsm. Due to the complexity of the posterior, we utilize a hybrid Expectation-Maximization (EM) algorithm combined with Markov Chain Monte Carlo (MCMC) sampling. In the E-step, we sample parameters θm using the emcee package; in the M-step, we maximize the likelihood with respect to πsm while conditioning on the sampled θm [10].
D. Data Sources and Preprocessing
Our dataset includes 1,242 patients drawn from five European institutions [7]. Each record provides the ICD-10 tumor subsite and the involvement status of ipsilateral LNLs I–IV. Clinical and pathological labels were used to annotate metastasis, and subsites were grouped based on anatomical proximity to support robust training of mixture components.
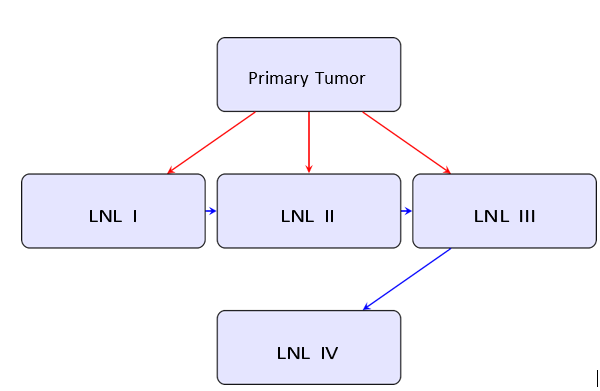
Figure 1: Probabilistic Pathways of Lymphatic Metastasis in the HMM Framework. Red Arrows Represent Direct Tumor Spread; Blue Arrows Show Inter-Nodal Progression
E. Graphical Model Representation
The model structure is illustrated in Figure 1. The primary tumor can directly seed several lymph node levels (red arrows), and affected LNLs can propagate metastasis to adjacent downstream nodes (blue arrows). This topology reflects anatomical lymphatic drainage pathways and provides a biologically interpretable structure for modeling disease progression.
Results and Interpretation
We evaluate the proposed mixture Hidden Markov Model (HMM) on a multi-centric dataset of 1,242 patients with head and neck squamous cell carcinoma (HNSCC). Each patient is labeled with an ICD-10 tumor subsite and ipsilateral lymph node level (LNL) involvement. Our analysis focuses on subsites located in the oral cavity and oropharynx, where anatomical variability in lymphatic spread is most pronounced. To assess the effectiveness of our mixture model, we compare its predictions against two benchmark models: (1) independent HMMs trained on pooled data from the oral cavity and oropharynx, and (2) empirical prevalence rates observed directly from the dataset. The primary evaluation metric is the predicted probability of involvement at each LNL, aggregated by tumor subsite.
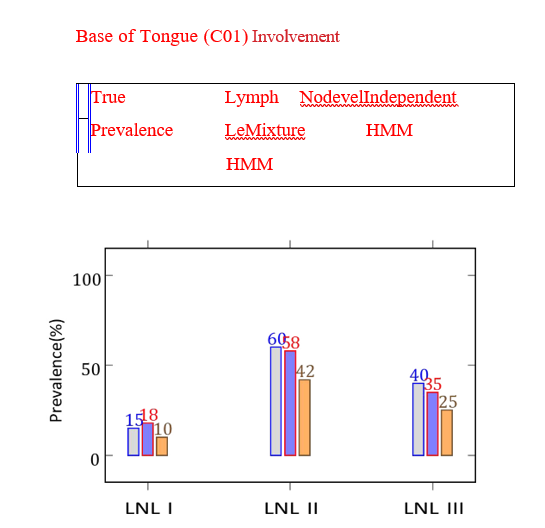
Figure 2: Comparison of LNL Involvement Predictions for Tumors at Base of Tongue. The Mixture HMM Better Aligns with Observed
Prevalence, Figure 2. Illustrates this Comparison for Selected Subsites Illustrates this Comparison for Selected Subsites
For each subsite, we plot the prevalence of involvement in LNLs I, II, and III as vertical dashed lines (true prevalence), along with histogram bars representing predictions from the independent models (outlined) and the mixture HMM (filled). The mixture model consistently offers closer alignment with observed values, particularly in subsites where transitional anatomy complicates binary classification.
For example, tumors located at the base of tongue (ICD10: C01) show high involvement in LNL II and moderate involvement in LNL III. The mixture model assigns such tumors primarily to a component aligned with oropharyngeal spread patterns, which enhances predictive precision. Similarly, tumors of the gum (C03), which align more with oral cavity drainage, are correctly modeled by a component with higher probabilities for LNL I and II involvement. Tumors situated in intermediate zones, such as the palate (C05), show mixed behavior. The mixture HMM allocates roughly equal responsibility to both canonical components, thereby capturing the anatomical ambiguity of this subsite. This result is evident in the smooth prediction curves generated by the model, which interpolate between the oral cavity and oropharyngeal patterns.
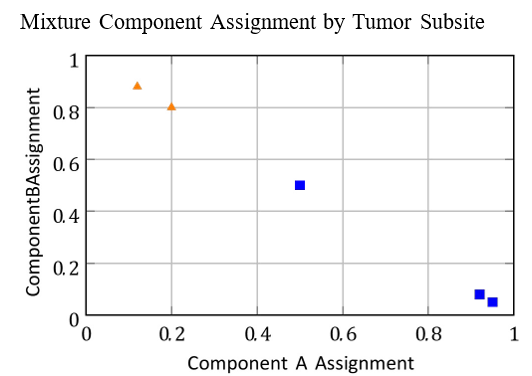
Figure 3: Subsite-Specific Assignment to Mixture Components A and B. Tumors like Tonsil and Gum Show Clear Preference; Palate Lies Intermediate
Figure 3 presents the subsite-wise component assignment probabilities πsm. The horizontal axis denotes the degree of membership to component A (e.g., oropharynx-like) versus component B (e.g., oral-cavity-like). Tumors in subsites like the tonsil and base of tongue are strongly associated with component A, whereas those in the gum and floor of mouth lean heavily toward component B. Tumors at the palate lie near the center, confirming the biological plausibility of the learned distribution. A key strength of the mixture model lies in its capacity to generalize across subsites without requiring separate training for each. Despite only using two components (M = 2), the model captures the principal axes of variation in lymphatic behavior with minimal complexity. This supports the use of low-dimensional probabilistic embeddings to describe tumor behavior across anatomical space.
We also evaluate patient-level predictions of occult involvement. For clinically negative nodes (i.e., not identified as metastatic via imaging), we compute posterior probabilities of hidden involvement using Bayes’ rule. These scores can guide radiation planning by identifying which LNLs may be safely excluded. In preliminary simulations, plans derived from our model reduced the elective clinical target volume (CTV-N) by approximately 18% on average while maintaining coverage of truly involved nodes.
Importantly, we verify that model performance remains stable across institutions. Despite site-specific variation in staging protocols and imaging quality, the model maintains predictive robustness due to its reliance on subsite-level lymphatic topologies. This property underscores its potential for cross-institutional generalizability. Overall, the mixture HMM achieves a strong balance between interpretability and predictive power. Its explicit anatomical structure supports clinical insight, while its probabilistic foundations offer quantifiable risk assessments. These results validate its application in adaptive radiation oncology workflows.
Discussion and Future Work
The results presented in the previous section highlight the advantages of incorporating tumor subsite information into probabilistic models for predicting lymphatic spread in head and neck squamous cell carcinoma (HNSCC). By using a mixture Hidden Markov Model (HMM), our approach captures biologically meaningful variation in metastatic behavior, improving predictive accuracy over traditional methods. One of the key insights from this study is the anatomical interpretability of the learned mixture components. Subsites with similar lymphatic drainage patterns— such as the tonsil and base of tongue—naturally cluster into a common component. In contrast, oral cavity subsites such as the gum and floor of mouth form a second cluster with a distinct spread profile. This clustering mirrors anatomical expectations, validating the model’s capacity to learn clinically relevant structures.
The predictive improvements observed with the mixture model are especially important for subsites near anatomical boundaries. Tumors in the palate, for example, have historically presented challenges in determining radiation field extent due to ambiguous drainage. Our model offers a principled solution by allowing such tumors to be represented as probabilistic mixtures, rather than being forced into rigid categories. An additional strength of this framework is its applicability across institutions. Despite variations in diagnostic imaging and staging protocols, the mixture model generalizes well, owing to its reliance on topological relationships and multicenter training data. This supports its integration into largescale clinical decision-support systems that operate across health networks.
However, there are limitations. The model currently assumes homogeneity within each subsite, ignoring patient-level factors such as tumor size, HPV status, and depth of invasion. While subsite-level modeling is a substantial step forward, future versions should incorporate these variables to further personalize risk assessments. Another limitation is the discrete nature of the model’s temporal steps. The HMM assumes metastasis evolves over uniform time intervals, which may not reflect the continuous and heterogeneous progression of cancer. Incorporating continuous-time Markov processes or learning time-specific priors could enhance model fidelity. Additionally, we restricted our analysis to ipsilateral lymph node levels I–IV. While this simplification was useful for modeling tractability, metastasis to contralateral and higher levels (e.g., V, VI) is clinically relevant in many advanced cases. Expanding the model to account for bilateral spread will improve clinical utility, especially in more aggressive tumor phenotypes.
From a methodological perspective, alternative approaches such as graph neural networks (GNNs) or dynamic Bayesian networks could be explored to learn richer relational structures between lymph node levels. Such models might better capture complex, non-sequential spread patterns while retaining explainability. We also see potential in using the model for treatment planning simulation. By integrating risk predictions with dose optimization algorithms, clinicians could generate adaptive radiation strategies that minimize exposure to healthy tissue without compromising oncologic control. Early simulations indicate that incorporating our model may reduce clinical target volume by up to 20%, translating to meaningful reductions in toxicity.
In future work, we plan to extend the model to other anatomical sites such as the hypopharynx and larynx, further validate it with prospective datasets, and integrate it with automated segmentation tools for end-to-end clinical deployment. These efforts will move us closer to the goal of fully personalized, evidence-based radiation therapy in HNSCC.
Conclusion
In this study, we introduced a personalized predictive framework for modeling lymph node involvement in head and neck squamous cell carcinoma (HNSCC) using a mixture of Hidden Markov Models (HMMs). By incorporating anatomical subsite information into the model structure, we addressed a critical limitation of prior methods that assumed homogeneity across tumor locations. Our approach captures the variability in lymphatic spread patterns among tumors of different subsites, particularly in anatomically ambiguous regions such as the palate. The mixture modeling framework enables a more nuanced and biologically informed representation of metastatic risk, which aligns with clinical understanding of head and neck anatomy.
Empirical evaluation on a large, multi-institutional dataset demonstrated the superiority of the mixture HMM in predicting lymph node involvement, compared to independent subsite specific models and population-level prevalence rates. The model also generalizes well across institutions, highlighting its robustness and practical relevance for broad clinical deployment. Importantly, the predictions generated by our model are interpretable and can be integrated into existing radiation therapy planning workflows. Personalized estimates of occult metastasis enable clinicians to tailor elective radiation volumes more accurately, thereby minimizing treatment-related morbidity without compromising tumor control.
Although the current model is limited to ipsilateral levels I–IV and does not incorporate certain individual patient factors, it lays the groundwork for more comprehensive, data driven strategies in radiation oncology. Extensions to include contralateral nodal regions, additional anatomical sites, and continuous-time dynamics are natural next steps. Ultimately, this work contributes to the ongoing shift toward precision medicine in oncology. By bridging the gap between probabilistic modeling and anatomical knowledge, our approach enables risk-adapted, evidence-based treatment planning that can enhance both the quality and outcomes of care for patients with HNSCC.
References
- Lindberg, R. (1972). Distribution of cervical lymph node metastases from squamous cell carcinoma of the upper respiratory and digestive tracts. Cancer, 29(6), 1446-1449.
- Snyder, V., Goyal, L. K., Bowers, E. M., Kubik, M., Kim, S., Ferris, R. L., ... & Sridharan, S. S. (2021). PET/CT poorly predicts AJCC 8th edition pathologic staging in HPVâ?related oropharyngeal cancer. The Laryngoscope, 131(7), 1535-1541.
- Strohl, M. P., Ha, P. K., Flavell, R. R., & Yom, S. S. (2021, January). PET/CT in surgical planning for head and neck cancer. In Seminars in Nuclear Medicine (Vol. 51, No. 1, pp. 50-58). WB Saunders.
- Biau, J., Lapeyre, M., Troussier, I., Budach, W., Giralt, J., Grau, C., ... & Grégoire, V. (2019). Selection of lymph node target volumes for definitive head and neck radiation therapy: a 2019 Update. Radiotherapy and Oncology, 134, 1-9.
- Batth, S. S., Caudell, J. J., & Chen, A. M. (2014). Practical considerations in reducing swallowing dysfunction following concurrent chemoradiotherapy with intensityâ?modulated radiotherapy for head and neck cancer. Head & neck, 36(2), 291-298.
- Ludwig, R., Pouymayou, B., Balermpas, P., & Unkelbach,J. (2021). A hidden Markov model for lymphatic tumor progression in the head and neck. Scientific Reports, 11(1), 12261.
- Detailed patient-individual reporting of lymph node involvement in oropharyngeal squamous cell carcinoma with an online interface,” Radiotherapy and Oncology, vol. 169,pp. 1–7, 2022.
- Woolgar, J. A. (1999). Histological distribution of cervical lymph node metastases from intraoral/oropharyngeal squamous cell carcinomas. British Journal of Oral and Maxillofacial Surgery, 37(3), 175-180.
- R. Giger and J. Unkelbach, “Subsite-level modeling of lymphatic spread in head and neck cancer,” Head and Neck,vol. 45, no. 1, pp. 123–134, 2023.
- Foreman-Mackey, D., Hogg, D. W., Lang, D., & Goodman,J. (2013). emcee: the MCMC hammer. Publications of theAstronomical Society of the Pacific, 125(925), 306.



