Research Article - (2024) Volume 3, Issue 3
Multidrug-Resistant Staphylococcus Aureus on the Hands of Healthcare Workers in Jos, Nigeria
2Department of Microbiology, University of Jos, Nigeria
3Department of Morbid Anatomy, Jos University Teaching Hospital, Nigeria
4Institute of Human Virology, Nigeria
5University of Nigeria Teaching Hospital Ituku-Ozalla, Enugu, Nigeria
6Ministry of Health and Sanitation, Sierra Leone, Nigeria
Received Date: Oct 31, 2024 / Accepted Date: Nov 25, 2024 / Published Date: Dec 04, 2024
Copyright: ©©2024 Dr. Alobu Walter Emeka, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Alobu, W. E., Emmanuel, Y., Adulugba, I. A., Eveshojiame, B. J., Giftson, M. A., et al. (2024). Multidrug-Resistant Staphylococcus Aureus on the Hands of Healthcare Workers in Jos, Nigeria. Archives Clin Med Microbiol, 3(3), 01-05.
Abstract
Introduction: Staphylococcus aureus has been implicated in a wide range of infections from mild skin infections to life threatening blood stream infections. The multidrug resistant strain is widely distributed within the hospital setting and readily transmitted by contaminated hands of healthcare workers.
Aim: The aim of this study was to investigate the susceptibility pattern and multidrug resistance index of Staphylococcus aureus isolated from hands of healthcare workers to routine antibiotics in Jos, North central Nigeria.
Methodology: Forty-eight isolates were collected from hands of 145 healthcare workers in Jos University Teaching Hospitals, Nigeria. The Staphylococcus aureus isolates were identified using standardized laboratory techniques. Antibiotic susceptibility testing was done using disc diffusion method, phenotypic method was used to detect methicillin resistance responsible for Methicillin-Resistant Staphylococcus aureus (MRSA).
Result: The in-vitro susceptibility of the Staphylococcus aureus isolates to antibiotics indicated 77.3-90% sensitivity to clindamycin, ciprofloxacin and gentamicin with 56-90% resistance to tetracycline, erythromycin and penicillin. Out of the 48 isolates tested with methicillin, 28 (58.3%) were resistant while 20 (41.7%) were sensitive. Multiple resistance was also observed to 6 frequently used antibiotics.
Conclusion: The current prevalence of multidrug resistant S. aureus present on the hands of healthcare workers is of great concern. Proper hand hygiene is key, as it is the most effective infection prevention and control (IPC) principle to curbing healthcare associated infections.
Keywords
MRSA, Hand Hygiene, Healthcare Workers, Infection, Carriage
Introduction
Staphylococcus aureus is a ubiquitous organism that is both a normal microbiota and a pathogen [1]. Hand and nasal carriage among individuals within the community ranges between 20 to 30% persistently or intermittently while within healthcare setting is between 25% to 50% [2-4]. As colonization precedes infection, colonized healthcare workers hands provide a reservoir from which S. aureus can be transmitted when host innate defences are breached (they become invasive) such as insertion of an indwelling catheter, or surgery and in secondary immunosuppression such as in HIV/AIDS [5,6]. In a study that examined S. aureus isolates from blood cultures of patients, it found out that these isolates were identical to nasal isolates in 82% of the infected patients[7]. This, therefore points to the fact that those with S. aureus infections may have become infected with their colonizing strain. Furthermore, a study within a healthcare setting demonstrated that about 17% (9–25%) of contacts between a HCWs and a MRSA-colonized patient resulted in transmission of MRSA from a patient to the gloves of a healthcare worker[8]. In another study which screened 177 healthcare workers, as much 36 (20.3%) of the HCWs were carriers of methicillin resistant Staphylococcus aureus in both their hand and anterior nares[9]. Beyond the healthcare setting, colonized patients and healthcare workers are potential source of community dissemination, thereby increasing the risk of spreading multidrug resistant pathogen within the community setting[10].
Timeline for development of resistance from the clinical introduction of penicillin in 1940, indicates that penicillinase was formally observed within a short period (less than 2 years) [11]. Resistance to penicillin was mediated by β-lactamase which hydrolyse the β-lactam rings and rendered the antibiotic non-effective[12]. Subsequently, after the introduction of methicillin between 1959 – 1960, the following year saw the emergence of methicillin-resistant Staphylococcus aureus isolates. Resistance to methicillin is mediated by the mecA gene, which encodes for a penicillin-binding protein (PBP2A) with reduced uptake for β-lactam antibiotics[13]. The aim of this study was to investigate the susceptibility pattern and multidrug resistance index of Staphylococcus aureus isolated from healthcare workers hands to routine antibiotics in Jos, North central Nigeria.
Materials and Methods
Ethical Approval
Informed consent was received for all individual participants while ethical approval for this work was also gotten from the Ethical Board/Committee of Jos University Teaching hospital. Reference number for ethical approval: JUTH/DCS/IREC/127/ XXX/2447.
Sampling
This study was a cross-sectional point prevalence study of healthcare workers (HCWs) in the medical and surgical wards, the intensive care unit and the special care baby unit of a tertiary care hospital in North-central Nigeria. Hand swabs were obtained from HCWs during clinical rounds without prior information about the procedure. These swabs were then taken to the Medical Microbiology laboratory for routine analysis. All HCWs who either recently washed their hands, used hand rub or avoided volunteering to give hand swabs were excluded from the study.
Sample size determination
The minimum sample size will be calculated using the following formula[13];
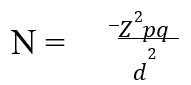
Where;
N = minimum sample size
Z = the standard normal deviation corresponding to 95% levels of significance (1.96)
P = local prevalence rate of Methicillin resistant Staphylococcus aureus in previous studies is 5.3
%14
q = 1-p
d = degree of accuracy desired, set at 5% = 0.05
Substituting the equation above;

Attrition
However, 10% of the calculated sample size was added to allow for attrition. The final sample size was approximately 85, but total sample size obtained at the end was 145.
Randomization
This was a non-randomized study.
Blinding
This study did not involve blinding.
Inclusion and Exclusion Criteria
Participants included in the study were all HCWs posted in selected wards and intensive care units (ICUs) of JUTH. Samples taken from hands of all health workers who gives consent. Participants excluded from the study includes All healthcare workers who would be found to have recently used a hand sanitizer or just washed their hands as at the time of sample collection.
HCWs who decline consent and participation
Laboratory Protocol and Phenotypic of MRSA
Forty-eight Staphylococcus aureus isolates from hands of one hundred and forty-five (145) healthcare workers taken while at their duty post using sterile water moistened swab. The swabs were then inoculated onto Mannitol Salt Agar (Oxoid Basingstoke, UK). These inoculated media plates were incubated aerobically for 24 hours at 37°C. After overnight incubation, yellow colonies which signified Staphylococcus species were examined further. The organisms on the positive plates were characterized using gram stain; catalase test and coagulase test (both slide and tube test) with standard requirements.
Antimicrobial susceptibility testing was done to observe the resistance pattern to conventional antibiotics. Antibiotics used was based on the various classes such as Inhibitors of the cell wall synthesis (penicillin and methicillin), protein synthesis (gentamycin, tetracyclines and erythromycin), and nucleic acid synthesis (ciprofloxacin). Susceptibility testing using the disc diffusion method on Muller Hinton agar according to Bauer et al. (1966) was employed[14]. The interpretation of the zone of inhibition of the single disc used was done according to manufacturers’ instruction and CLSI manuals. The percentage resistance, multiple antibiotic resistance (MAR) was calculated as follows: Percentage resistance = (No of resistant isolates/ No of isolates tested with the antibiotic) x 100. Methicillin resistance was tested using methicillin (30g) disc with cut-off zone of inhibition of 21mm on each isolate. For internal quality control a known clinical isolate of MRSA was used as a positive control and Staphylococcus aureus ATCC 25923 as a negative control.
DNA extraction was carried out following standard protocol according to Zymo DNA mini-prep extraction kit (Zymo research, USA). The detection of mecA using a primer with 162 bp was carried out according to Larsen et al., (2008). Data interpretation and statistical analysis was done using SPSS. The quantitative data are expressed as numbers and percentages.
Results
Forty-eight isolates of Staphylococcus aureus were recovered from hands of 145 healthcare workers. The demographic distribution of the healthcare workers is depicted in Table 1. Forty-three isolates (90.0%) were most resistant to beta lactam antibiotics: penicillin. As high as 29 (60.4%) and 27 (56.3%) isolates were resistant to protein inhibitors: erythromycin and tetracycline. However, other protein inhibitors showed less resistance, 12 (25%) and 13 (27%) respectively for gentamycin and clindamycin. Beta lactam stable fluoroquinolones: ciprofloxacin demonstrated less resistance with 15 (31%) isolates being resistant. Out of the 48 isolates tested with methicillin, 28 (58.3%) were resistant while 20 (41.7%) were sensitive (Table 2). Many isolates demonstrated multidrug resistances. Only one isolate was (2%) resistant to all 6 antibiotics while a total of 16 (33.3%) isolates had highest multidrug resistance to 4 antibiotics (MAR of 0.333). A multiple resistant index per isolates of 0.02 was observed for 6 tested antibiotics (Table 3). High percentage of multiple drug resistance quantified as index per isolates (Table 3) was illustrated in percentage. The resistance pattern of the isolates (figure 1) is shown.
|
Characteristics |
Frequency (n:145) |
(%) |
S. aureus (n: 48) |
|
Age group (years) |
|
|
|
|
< 47 |
113 |
84.2 |
38 |
|
> 47 |
21 |
15.7 |
10 |
|
Sex |
|
|
|
|
Female |
75 |
52.1 |
29 |
|
Male |
69 |
47.9 |
18 |
|
Department/ward |
|
|
|
|
Medical ward |
52 |
35.9 |
18 |
|
Surgical ward |
59 |
40.7 |
10 |
|
ICU |
10 |
6.9 |
7 |
|
Others |
16 |
11.0 |
6 |
|
Profession |
|
|
|
|
Doctor |
81 |
55.8 |
22 |
|
Nurse |
43 |
29.7 |
16 |
|
others |
21 |
14.5 |
10 |
Table 1: Demographic and Clinical Characteristics of Study Participants
|
Antibiotics |
Number (%) of resistance isolates |
Number (%) of susceptible isolates |
|
Methicillin |
28/48 (58.3) |
20 (41.7) |
Table 2: Methicillin Resistance
|
Number of Antibiotics to which Resistance occurred |
Number of Resistant Isolates |
Multiple Resistance Index |
|
0 |
0 |
0 |
|
1 |
6 |
0.125 |
|
2 |
10 |
0.208 |
|
3 |
12 |
0.250 |
|
4 |
16 |
0.333 |
|
5 |
3 |
0.063 |
|
6 |
0 |
0 |
|
7 |
1 |
0.021 |
|
Total |
48 |
|
Table 3: Multiple Resistance Index with Respect to Isolates
Figure 1: Resistance Pattern of Staphylococcus aureus isolates
Discussion
Staphylococcus aureus remains responsible for most clinical infections within the healthcare and community settings. Their ability to cause disease is largely based on its ubiquitous nature as a normal microbiota and an opportunistic pathogen. Multidrug-resistant infections have been associated with increased cost, prolonged length of hospital stay, and excess in-hospital mortality[15]. Most importantly, invasive infections resulting from multidrug resistance strain of S. aureus carries with it a poor patient outcome and economic implications. Acquisition of the multidrug resistance features by have further impacted on the poor outcome of healthcare-associated infections. Specifically, methicillin resistance have been directly correlated with prolonged length of stay after infection[16]. A systematic analytic study done in 254 cases of methicillin-resistant Staphylococcus aureus-Blood stream infection (MRSA-BSI), found out that death related to BSI occurred in 81 cases (31.9%) of the MRSA-BSI while only 12 cases (14.1%) of methicillin-sensitive Staphylococcus aureus-Blood stream Infection (MSSA-BSI) had a poor (death) outcome[17]. Therefore, colonization with multidrug resistant S. aureus increases the chances of infection up to 13 times[4]. Within the community setting, S. aureus, most importantly MRSA is estimated to cause up to 90% of skin and soft tissue infections (SSTI), pus and abscess formation[4,18]. In a study which surveyed 914 members of 321 households found the prevalence of S. aureus and MRSA to be 25% and 0.4% respectively with Seventy-eight (24%) households reporting serious skin infection[19]. Furthermore, acquisition of methicillin resistance was associated with resistance to macrolides, lincosamides, and streptogramin B (MLSB) rendering S[20]. aureus resistant to most classes of the readily available antibiotics. The SCCmec complex which carries the mecA gene and other gene regulatory components has been classified into twelve known SCCmec types (I–XII)[4]. This SCCmec types has been employed to group MRSA into health-care-associated MRSA (HA-MRSA) community-associated MRSA (CA-MRSA) and livestock-associated MRSA (LA-MRSA) strains[4].
High levels of resistance of Staphylococcus aureus to penicillin was noted in this study up to 90%. This in keeping with other studies, such as this which reported a high proportion (89.9%) of S. aureus isolates being resistant to penicillin[21]. This high resistance to penicillin is also seen in erythromycin and tetracycline which is keeping with acquired resistance to macrolides, lincosamides, and streptogramin B (MLSB) associated with methicillin resistance[20].
There was high susceptibility of Staphylococcus aureus to gentamicin class of the protein inhibitors and to ciprofloxacin, which is in keeping with a similar study which observed high susceptibility to amikacin and gentamicin[22]. Multidrug resistance of Staphylococcus aureus isolates gotten from healthcare-associated infection and community acquired infections show different resistant patterns.
The multiple antibiotic resistance (MAR) being an easy and cost-effective way to tract antibiotic resistance was employed to access the level of resistance using a cut off 0.2 which signifies high level of antimicrobial resistance[23]. The highest number of isolates were resistant to 4 groups of antibiotics (33.3%) which was also observed in another study with 53.8% of isolates resistant to 4 antibiotics [22]. Low MAR index was observed with resistance to one and five antibiotics while only one isolate was resistant to all 6 antibiotics. The limitations of this study include the few numbers of antibiotics discs tested against the isolates and inability to test all 48 isolates for molecular detection of A gene.
Conclusion
The high levels of multidrug resistance observed with Staphylococcus aureus isolated from hands of HCWs is a cause for concern. Regular hand hygiene audit and MRSA surveillance is critical to ensure that patient, healthcare workers and the hospital environment are kept free of this pathogen. Proper hand hygiene is key as it is the most effective infection prevention and control (IPC) principle to curbing healthcare associated infections.
Conflict of Interest
None
Funding
This work was funded through collective efforts of participating authors. No external funding was received.
Data Summary
No new data generated.
Author Contributions
Alobu Walter Emeka conceived the research idea, sample collection and processing, proposal and manuscript writing. Yemgbe Emmanuel worked on data entry and analysis, contributed to proposal and manuscript writing, correction and final input while Ifeoma Adulugba, Obinna Innocent Oke, Ben James, Musa Adah Giftson, Ayeni Christabel Oyowo, Alpha Umaru Bai-Sesayparticipated in sample collection and contributed to proposal and manuscript writing.
References
- Akter, R., Paul, S., Naher, A., Rahman, M., Sultana, S., & Sarkar, D. (2017). Nasal Carriage of Methicillin Resistant Staphylococcus aureus Among Health Care Workers At Tertiary Care Hospital In Dhaka. Bangladesh Journal of Medical Microbiology, 11(2), 17-19.
- Tigabu, A., Tiruneh, M., & Mekonnen, F. (2018). Nasal Carriage Rate, Antimicrobial Susceptibility Pattern, and Associated Factors of Staphylococcus aureus with Special Emphasis on MRSA among Urban and Rural Elementary School Children in Gondar, Northwest Ethiopia: A Comparative Cross-Sectional Study. Advances in preventive medicine, 2018(1), 9364757.
- Walana, W., Bobzah, B. P., Kuugbee, E. D., Acquah, S.,Ezekiel, V. K., Yabasin, I. B., ... & Ziem, J. B. (2020). Staphylococcus aureus nasal carriage among healthcare workers, inpatients and caretakers in the Tamale Teaching Hospital, Ghana. Scientific African, 8, e00325.
- Lee, A. S., De Lencastre, H., Garau, J., Kluytmans, J., Malhotra-Kumar, S., Peschel, A., & Harbarth, S. (2018). Methicillin-resistant Staphylococcus aureus. Nature reviews Disease primers, 4(1), 1-23.
- R S, D V. (2021). MRSA colonization in health care professionals with varying degrees of exposure to the hospital environment and its susceptibility to mupirocin in a tertiary care centre. Indian Journal of Microbiology Research. Apr 28;8:71–5.
- Hidron, A. I., Kempker, R., Moanna, A., & Rimland, D. (2010). Methicillin-resistant Staphylococcus aureus in HIV-infected patients. Infection and drug resistance, 73-86.
- Von Eiff, C., Becker, K., Machka, K., Stammer, H., & Peters,G. (2001). Nasal carriage as a source of Staphylococcus aureus bacteremia. New England Journal of Medicine, 344(1), 11-16.
- McBryde, E. S., Bradley, L. C., Whitby, M., & McElwain,D. L. S. (2004). An investigation of contact transmission of methicillin-resistant Staphylococcus aureus. Journal of Hospital Infection, 58(2), 104-108.
- Gebreyesus, A., Gebre-Selassie, S., & Mihert, A. (2013). Nasal and hand carriage rate of methicillin resistant Staphylococcus aureus (MRSA) among health care workers in Mekelle Hospital, North Ethiopia. Ethiopian medical journal, 51(1), 41-47.
- Cimolai, N. (2008). The role of healthcare personnel in the maintenance and spread of methicillin-resistant Staphylococcus aureus. Journal of infection and public health, 1(2), 78-100..
- Foster, T. J. (2017). Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. FEMS microbiology reviews, 41(3), 430-449.
- Enright MC, Robinson DA, Randle G, Feil EJ, Grundmann H, Spratt BG. (2022). The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA). Clin Microbiol. 2003 Aug;41(8):3765–3776.
- Turner, N. A., Sharma-Kuinkel, B. K., Maskarinec, S. A., Eichenberger, E. M., Shah, P. P., Carugati, M., ... & Fowler Jr, V. G. (2019). Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nature Reviews Microbiology, 17(4), 203-218.
- Hudzicki, J. (2009). Kirby-Bauer disk diffusion susceptibility test protocol. American society for microbiology, 15(1), 1-23.
- Serra-Burriel, M., Keys, M., Campillo-Artero, C., Agodi, A., Barchitta, M., Gikas, A., ... & López-Casasnovas, G. (2020). Impact of multi-drug resistant bacteria on economic and clinical outcomes of healthcare-associated infections in adults: Systematic review and meta-analysis. PloS one, 15(1), e0227139.
- Rauber, J. M., Carneiro, M., Arnhold, G. H., Zanotto, M. B.,Wappler, P. R., Baggiotto, B., ... & d'Azevedo, P. A. (2016). Multidrug-resistant Staphylococcus spp and its impact on patient outcome. American journal of infection control, 44(11), e261-e263.
- Kim, C. J., Kim, H. B., Oh, M. D., Kim, Y., Kim, A., Oh, S.H., ... & KIND Study group (Korea Infectious Diseases Study group). (2014). The burden of nosocomial staphylococcus aureus bloodstream infection in South Korea: a prospective hospital-based nationwide study. BMC infectious diseases, 14, 1-9.
- Kim, M. W., Greenfield, B. K., Snyder, R. E., Steinmaus,C. M., & Riley, L. W. (2018). The association between community-associated Staphylococcus aureus colonization and disease: a meta-analysis. BMC infectious diseases, 18, 1-11.
- Miller, M., Cook, H. A., Furuya, E. Y., Bhat, M., Lee, M. H., Vavagiakis, P., ... & Lowy, F. D. (2009). Staphylococcus aureus in the community: colonization versus infection. PLoS One, 4(8), e6708.
- Miklasinska-Majdanik, M. (2021). Mechanisms of resistance to macrolide antibiotics among Staphylococcus aureus. Antibiotics, 10(11), 1406.
- Joachim,A., Moyo, S. J., Nkinda, L., Majigo, M., Rugarabamu, S., Mkashabani, E. G., ... & Lyamuya, E. F. (2018). Nasal Carriage of Methicillin-Resistant Staphylococcus aureus among Health Care Workers in Tertiary and Regional Hospitals in Dar es Salam, Tanzania. International journal of microbiology, 2018(1), 5058390.
- Kot, B., Wierzchowska, K., Piechota, M., & Gruzewska,A. (2020). Antimicrobial resistance patterns in methicillin-resistant Staphylococcus aureus from patients hospitalized during 2015–2017 in hospitals in Poland. Medical Principles and Practice, 29(1), 61-68.
- Ayandele, A. A., Oladipo, E. K., Oyebisi, O., & Kaka, M. O. (2020). Prevalence of multi-antibiotic resistant Escherichia coli and Klebsiella species obtained from a tertiary medical institution in Oyo State, Nigeria. Qatar medical journal, 2020(1), 9.



