Research Article - (2023) Volume 7, Issue 1
Molecular Approach for Identification of Pangasianodon Hypophthalmus based on Mitochondrial Cytochrome b Oxidase subunit i (coi) Gene from Pakistan.
Received Date: Dec 21, 2023 / Accepted Date: Dec 28, 2023 / Published Date: Jan 06, 2023
Copyright: ©Â©2023: Tayyaba Malik This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Malik, T., Naeem, M. (2023). Molecular Approach for Identification of Pangasianodon Hypophthalmus based on Mitochondrial Cytochrome b Oxidase subunit i (coi) Gene from Pakistan. Stem Cell Res Int, 7(1), 01-07.
Abstract
The aquaculture industry is dependent on rich fish resources in water bodies. Human activities have led to a rapid decline of fish species. In Asia, the Pangasiidae family is highly valued for its potential for survival and its fillet meat. DNA barcoding is a taxonomic method using genetic markers in organisms mitochondrial DNA (mt DNA) for identification. The phylogeny and identification of Pangasianodon hypophthalmus in the subcontinent is of great concern. For species identification, a precise and rapid technique is DNA barcoding. This method is strongly effective for analyzing the divergence among species. DNA barcoding is more reliable as compared to external morphology. To avoid mislabeling and conservation of species, it is equally useful in juveniles as well as adult stages of fishes. As DNA bar-coding is a taxonomic method that uses small genetic markers in organisms’ mitochondrial DNA (mt DNA) for identification of particular species. In recent study MAGA X and Kimura 2 Parameter was used to evaluate genetic distance and neighbor joining tree was constructed. BOLD and GenBank reveals the nearest identity matches. As mitochondrial cyt-b gene region was successfully used for identifying species and accepted as a standard region for DNA barcoding.
Keywords
DNA Barcoding, Hypophthalmus, Mitochondrial Cyt-b, Pakistan
Introduction
Globally, fish account for more than half of all vertebrates, with more than 30,000 species. Additionally, fish provide humans with animal protein and are an important component of biodiversity. Classification and identification of fish are essential to fishery in- vestigations, nature reserve assessments, and food and drug iden- tification [1]. In order to study the huge variety of fish species, a proper taxonomical key is important and its molecular basis should be known [2]. Ontogenetic metamorphosis occurs in most fishes, resulting in a diverse range of morphological characteristics. On the path to ontogenetic development, many morphometric features changes. Species identification by Morphological characters pos- es great challenge and controversy to taxonomy as convergent or divergent evolution could cause changes in morphological char- acters [1].
DNA barcoding is a tools that helps to identify the closely related species in diverse fish fauna that is based upon the isolation of DNA segments, as early studies have already recognized that se- quence diversity in a 650-bp fragment of the mitochondrial-DNA, CO1 provides solid species level resolution for large variety of animal groups [3, 4]. DNA barcoding technique is based upon the molecular approach of DNA isolation and tag them for a single species and stores them, later this data is used for identification purposes [5]. From ovum to adult, barcoding eliminates the diffi- culties taxonomists face in identifying organisms [6].
The extraction of biomolecules, DNA, RNA, and protein, is of utmost importance in early days of experimental biology and for monitoring biological assaults [7, 8]. In contemporary biological sciences the isolation of DNA is a key step in many diagnostic kits for various diseases, DNA molecules can be extracted from various sample of dead tissues, bones, virus sample or dead organ- ic matter for the purpose of study [7]. DNA isolation is achieved by complete splintering of tissues that release cells and underly- ing organelles, denaturation of protein complexes, and activation of nucleases that release the DNA molecules [9]. Closely related intermixing of fish fauna is an important problem to solve, one species may evolve to many other species levels, and this requires a cheap and rapid method in a mass production environment [10].
DNA is relatively stable molecule at high temperature and usu- ally provides far better hereditary and molecular identification of closely related species, that provides insight into diverse fish fau- na, to keep a check on invasive and key varieties [10, 11]. It can be analyzed with a single piece of isolated gene to identify it again [12]. However, genetic information for less than 10% of the total species is yet documented [13]. PCR (polymerase chain reaction ) is an advance method of amplification of target sequence of gene and make copies for the desired DNA sequence, it is also noted that this process is very quick and specific but also very sensitive [14].
The history of Lessepsian incursions demonstrates those species that becomes problematic to categorize commonly remain unrec- ognized or misidentified [15]. In recent years the documentation of fish species has gained consideration due to the cumulative cases of deception in the fish processing industry [16]. DNA barcoding aims to provide a competent method for species-level identifica- tions using a range of species-specific molecular tags derived from the five regions of the mitochondrial cytochrome c oxidase I (COI) gene [17]. As a result, scientists worldwide have established mito- chondrial DNA databases and repositories for all animals, includ- ing fish [6]. Striped catfish (Pangasianodon hypophthalmus) is presently the principal freshwater aquaculture species, more than 90% of it is marketed as fillet out of which most is exported to over 127 countries worldwide and is often being mislabeled [18, 19].
Materials and Methods
In recent work, for identification through DNA barcoding spec- imens of P. hypophthalmus were obtained from Tawakkal Fish Hatchery near Muzaffargarh, Punjab, Pakistan. As the standard taxonomic key was utilized to recognize the fish at morphologi- cal basis. Phenol Chloroform extraction method was executed for isolation of DNA from fish tissues. The weight of tissues was ap- proximately 50mg, they were grinded by adding extraction buffer. 12µl Proteinase-K was added and incubated at 37°C and 55°C for 60mins respectively. Then centrifuged at 5000rpm for 10mins. The supernatant was collected; Phenol, Chloroform and Isoamyl alco- hol were added in 25:24:1 ratio and centrifuge at 12000rpm for 10min. Top aqueous layer was collected; Chloroform and Isoamyl alcohol in the ratio 24:1 were added, after gently mixing again centrifuge at 12000rpm for 10min. 0.1 volume of 3M sodium ac- etate and equal volume of ice cold ethanol (100%) was added to the supernatant and mixed thoroughly until DNA pallet was ob- tained. After incubation at -20°C for 1hr it was again centrifuged at 1000rpm for 10min and decant the supernatant. After washing with 70% ethanol air dry DNA pellets were suspended in 100µl of distilled water and stored at -20°C for further study. Finally sample was electrophoresed on 0.7% agarose gel at 80volt for 30mins. DNA concentration and purity was evaluated by Thermo-Scien- tific Nanodrops. DNA sequence was amplified from region of mi- tochondrial cytochrome b oxidase I (COI) gene using the primers pair as follows:
Cyt-b Forward primer:5’AGCCTACGAAAAACCCACCC 3’
Cyt-b Reverse primer: 5’AAACTGCAGCCCCTCAGAATGA- TATTTGTCCTC 3’
SensQuest Labcycler was used for PCR amplification. For this processes 25µl of reaction mixture was used in PCR, containing distilled water 11.3µl, master mix 12.5µl, forward primer 0.1µl, reverse primer 0.1µl and DNA template sample 1µl. The PCR thermal cycling conditions includes an initial denaturation of 95°C (5 min), followed by denaturation at 95°C (30 sec; 40 cycles), an- nealing at 55°C (30 sec) and extension at 72°C (30 sec), with final extension at 72°C (7 min). After amplification, PCR products were run on 1.5% agarose gel for 50 min and then visualized using UV transilluminator to assess the quality of the amplified product. The most clarified samples were selected for the sequencing purpose.
Results
The purified PCR products was for sequencing. Blast results by NCBI helps to determine the best match homology. The barcode was deposited to GenBank under Accession number MT441543.1(Pangasianodon hypophthalmus) with 364bp and BLAST results in table.1.1 and pairwise genetic distance in ta- ble.1.2.
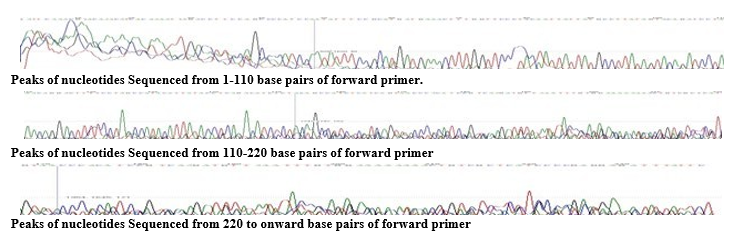
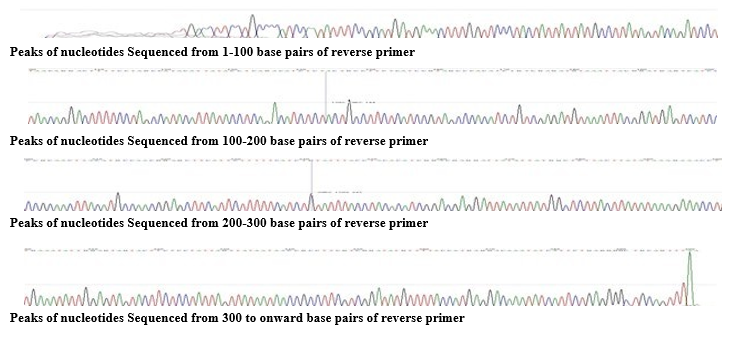
Figure 1: Results of Electropherogram showing Peaks of nucleotides
A distinctive electropherogram was formed after DNA sequencing to show nucleotides in colorful peaks(fig.1.1). By means of MEGA X software; genetic distances and phylogenetic relationship was analyzed as NJ trees and K2P genetics were produced Evolution- ary analyses of the aligned sequences were conducted by program MEGA X (fig.1.3). The phylogenetic tree (fig.1.2) was rooted among P hypophthalmus Accession No: MT441543.1 and close- ly related KY586022.1, KY586028.1, KY5860251, KY586008.1. The relationship was analyzed as NJ tree and K2P genetics were rooted among above (fig.1.2 and 1.3)
Figure 2: Phylogenetic distances
Figure 3: BLAST Neighbor joining tree by blast inquiry with Distance Evaluating of P. hypophthalmus
Table 1: Accession No. and BLAST results analysis of Pangasianodon hypopthalmus
|
Query ID |
NCBI: tax- onomy ID |
Sequence Length |
Accession No. |
Percentage similarity |
E Value |
Max Score |
Total Score |
Accession No. of best match & % match |
|
43205 |
310915 |
364 |
MT441543.1 |
100% |
0.0 |
673 |
673 |
KY586022.1 |
|
|
|
|
|
|
|
|
|
(100%) |
|
|
|
|
|
|
|
|
|
KY586028.1 |
|
|
|
|
|
|
|
|
|
(99.73%) |
|
|
|
|
|
|
|
|
|
KY586025.1 |
|
|
|
|
|
|
|
|
|
(99.73%) |
|
|
|
|
|
|
|
|
|
KY586008.1 |
|
|
|
|
|
|
|
|
|
(100%) |
Table 2: Pairwise Genetic Divergence between Species of Pangasianodon hypophthalmus
|
|
MT441543.1 P.hypophthalmus |
KY586022.1 P.hypophthalmus |
KY586028.1 P.hy- pophthalmus |
KY586025.1 P.hy- pophthalmus |
KY586008.1 P.hypophthalmus |
|
MT441543.1 P.hypophthalmus |
|
|
|
|
|
|
KY586022.1 P.hypophthalmus |
0.0000 |
|
|
|
|
|
KY586028.1 P.hy- pophthalmus |
0.0028 |
0.0049 |
|
|
|
|
KY586025.1 P.hy- pophthalmus |
0.0028 |
0.0026 |
0.0000 |
|
|
|
KY586008.1 P.hy- pophthalmus |
0.0055 |
0.0099 |
0.0099 |
0.0077 |
|
Discussion
For inferring fish phylogenetic relationships and understanding speciation, only morphometric characteristics were used before because Pangasiidae species are highly similar, it is difficult to distinguish them [20]. Molecular analysis based on sequence of mitochondrial-DNA for is helpful in accurate recognition of un- known species [21, 22]. Cyt-b(cytochrome) gene sequence of 650 base pair are usually used in study for species identification and sequence of 649bp was used for identification of Labeo bata from Pakistan [23, 24]. Present study on Pangasianodon hypophthal- mus was conducted to examine the species by DNA barcoding. Amplified PCR product was sequenced, and complete barcode’s fragments of 364 base pairs of P. hypophthalmus (Accession No: MT441543.1) were obtained. Earlier reports fragments of base pairs extending 157 to 541 bp in humans, which lies close to re- cently identified base pairs(364bp) [25]. The most appropriate way of distinguishing the DNA depend diversity of intra and in- terspecific is barcode analysis used to distinguish diverse species, which correspond with neighbor joining tree. Besides that many standardized methods are still used for classification and distance measurement [26]. Comparison of quality and quantity of DNA were accessed by Nanodrop and absorbance were noted for 1.6 to 2 for purified DNA product after extraction as the value of absor- bance were found 1.6-1.76 study revealed that present extraction quantification were in general agreement [24, 27].
Study of molecular biology by Cytochrome b also useful for mo- lecular taxonomy of fish species [28]. Identification sequence as Cytochrome b gene is significantly used for identification of animals and plants diversity [29]. So the present study used Cy- tochrome b for the identification of fish species. DNA barcoding marked as very useful for evolutionist, taxonomist and phyloge- netic study. Previous studies were in debate about the importance of these studies for taxonomists in DNA barcoding processes to identifying species and its bio-diversity [30]. Cyt b oxidase was successfully used for DNA barcoding of fishes, as it effectively mark the differences among specific species [31]. The barcoding could be useful in a taxonomic study but not as a whole; as it pro- vides additional information to make the research significant [32].
Species identification was accessed by using by BOLD and NCBI database for this study [33]. Sometimes the results from Barcode of life database (NCBI and BOLD) had significant differences in identification percentage and phylogenetics. This database may also have some unknown species absent in record like P. tenuisp- inis. Recent results reveal that DNA barcoding is more specifically accessing fishes on basis of diversity [4]. Phylogenetic differences and species identification were noticeably explained on basis of cytochrome b genes, considered as the preeminent identification tool. The genetic divergence was found in their sequences due to variations. Those variation percentages are measured on the basis of knowing species similarity index [34]. Alignment of nitrogenous bases in present study was close to [35]. Several countries require that species be identified when fish are supplied to souks without proper labeling [36]. DNA-barcoding can be used for confirmation results of fish species that were under consideration, as previously studied by [37]. As these DNA bands clearly indicates the size of sequence that were under consideration for study. In many species of the catfish the specimen were mislabeled reasoned lacking in proper DNA based identification. Species were observed to be in- terconnected, such analysis of closeness were assessed by measur- ing of distance by neighbor joining tree, as identical relationship was marked in present study where closely related species were grouped together to formulate a tree [38]. As the general reference sequence, BOLD and NCBI are operative, and data-base helps in explaining sequences [39]. As this study also mentioned it [40-50].
Conclusion
Identification of fishes on molecular bases, in Pakistan is not being considered an accustomed exercise. Numerous techniques prac- ticed, for classifying documentation of fish species traditionally; are less authenticated in contrast to approach of techniques aligned at molecular roots. Recent study was an attempt to provoke the fu- ture of these modern approaches. Prior techniques utilized for eras were confined; as they do not show effective outcomes for misla- beling of fish-fillet and fish species, besides spoiled specimens. Substantiations among nucleic divergences amongst species; gen- era and family could be examined with molecular approaches.
Disclosure statement
No potential conflict of interest was reported by the authors. Prof. Dr. Muhammad Naeem designed and invigilate the work. Tayyaba Malik performs and analyzed work and write the manu- script.
References
- Bingpeng, X., Heshan, L., Zhilan, Z., Chunguang, W., Yan- guo, W., & Jianjun, W. (2018). DNA barcoding for identifi- cation of fish species in the Taiwan Strait. PloS one, 13(6), e0198109.
- Vartak, V. R., Narasimmalu, R., Annam, P. K., Singh, D. P., & Lakra, W. S. (2015). DNA barcoding detected improper la- belling and supersession of crab food served by restaurants in India. Journal of the Science of Food and Agriculture, 95(2), 359-366.
- Ramadan, H. A., & Baeshen, N. A. (2012). Biological Iden- tifications through DNA barcodes. Biodiversity conservation and utilization in a diverse World, InTech Publisher, Repub- lika Hrvatska, 109-129.
- Takahara, T., Minamoto, T., & Doi, H. (2013). Using environ- mental DNA to estimate the distribution of an invasive fish species in ponds. PloS one, 8(2), e56584.
- Hebert, P. D., & Gregory, T. R. (2005). The promise of DNA barcoding for taxonomy. Systematic biology, 54(5), 852-859.
- Ayesha, U. R., Shafi, N., Akhtar, T., Zareen, A., & Ayub, H. (2019). DNA barcoding of cyprinids (Labeo rohita, Catla catla and Cirrhinus mrigala), mitochondrial CO1-based study. Mi- tochondrial DNA Part B, 4(1), 405-407.
- Wink, M., Yousef, A. E., & Carstrom, C. An Introduction to Molecular Biotechnology.
- Armstrong, K. F., & Ball, S. L. (2005). DNA barcodes for biosecurity: invasive species identification. Philosophical Transactions of the Royal Society B: Biological Sciences, 360(1462), 1813-1823.
- Doyle, K. (1996). The source of discovery: protocols and ap- plications guide. Madison: PROMEGA.
- Pardo, M. A., & Pérez-Villareal, B. (2004). Identification of commercial canned tuna species by restriction site analysis of mitochondrial DNA products obtained by nested primer PCR. Food Chemistry, 86(1), 143-150.
- Darling, J. A., & Blum, M. J. (2007). DNA-based methods for monitoring invasive species: a review and prospectus. Biolog-ical Invasions, 9(7), 751-765.
- Ram, J. L., Ram, M. L., & Baidoun, F. F. (1996). Authenti- cation of canned tuna and bonito by sequence and restriction site analysis of polymerase chain reaction products of mito- chondrial DNA. Journal of Agricultural and Food Chemistry, 44(8), 2460-2467.
- Golani, D., & Appelbaum-Golani, B. (2010). Fish invasions of the Mediterranean Sea: change and renewal.
- Lockley, A. K., & Bardsley, R. G. (2000). DNA-based meth- ods for food authentication. Trends in Food Science & Tech- nology, 11(2), 67-77.
- Azzurro, E., Goren, M., Diamant, A., Galil, B., & Bernardi, G. (2015). Establishing the identity and assessing the dynamics of invasion in the Mediterranean Sea by the dusky sweeper, Pempheris rhomboidea Kossmann & Räuber, 1877 (Pempher- idae, Perciformes). Biological Invasions, 17(3), 815-826.
- Espiñeira, M., Gonzalez-Lavín, N., Vieites, J. M., & San- taclara, F. J. (2009). Development of a method for the identi- fication of scombroid and common substitute species in sea- food products by FINS. Food Chemistry, 117(4), 698-704.
- Hubert, N., Hanner, R., Holm, E., Mandrak, N. E., Taylor, E., Burridge, M., ... & Bernatchez, L. (2008). Identifying Canadi- an freshwater fishes through DNA barcodes. PLoS one, 3(6), e2490.
- Tung, N. T., Tuan, N., Tuan, T. T., & Hoa, N. D. (2001). Devel- opment situation of two fish species of Pangasiidae cultured in the Mekong data of Viet Nam (Pangasianodon hypophthalmus and Pangasius bocourti). Assessment of Mekong Fisheries Project. Unpublished document. Mekong River Commission, Phnom Penh.
- Dung, N. H. (2008, December). Vietnam Pangasius and World Markets. In Presentation Given at the International Sympo- sium on Catfish Aquaculture in Asia: Present Status and Chal- lenges for Sustainable Development. Can Tho University.
- AHMED, M. S., DINA, S. R., NAHAR, L., ISLAM, N. N.,& Al REZA, H. (2018). Molecular characterization of Chan- na species from Bangladesh based on Cytochrome c Oxidase Subunit I (COI) gene. FishTaxa, 3(4), 87-93.
- Dawnay, N., Ogden, R., McEwing, R., Carvalho, G. R., & Thorpe, R. S. (2007). Validation of the barcoding gene COI for use in forensic genetic species identification. Forensic sci- ence international, 173(1), 1-6.
- Kerr, K. C., Lijtmaer, D. A., Barreira, A. S., Hebert, P. D., & Tubaro, P. L. (2009). Probing evolutionary patterns in Neo- tropical birds through DNA barcodes. PLoS One, 4(2), e4379.
- Lohman, D. J., Prawiradilaga, D. M., & Meier, R. (2009). Im- proved COI barcoding primers for Southeast Asian perching birds (Aves: Passeriformes). Molecular Ecology Resources, 9(1), 37-40.
- Naeem, M., & Hassan, S. (2019). Molecular approach for identification of Labeo bata based on COI gene sequence from Pakistan. Mitochondrial DNA Part B, 4(1), 244-246.
- Ferri, G., Alu, M., Corradini, B., Licata, M., & Beduschi, G. (2009). Species identification through DNA “barcodes”. Ge- netic testing and molecular biomarkers, 13(3), 421-426.
- Bhattacharjee, M. J., Laskar, B. A., Dhar, B., & Ghosh, S. K. (2012). Identification and re-evaluation of freshwater catfish- es through DNA barcoding. PloS one, 7(11), e49950.
- Cawthorn, D. M., Steinman, H. A., & Witthuhn, R. C. (2011). Comparative study of different methods for the extraction of DNA from fish species commercially available in South Afri- ca. Food Control, 22(2), 231-244.
- Luo, A., Zhang, A., Ho, S. Y., Xu, W., Zhang, Y., Shi, W., ... &Zhu, C. (2011). Potential efficacy of mitochondrial genes for animal DNA barcoding: a case study using eutherian mam- mals. BMC genomics, 12(1), 1-13.
- MALIK, T., & NAEEM, M. (2022). Molecular approach for identification of Pangasianodon hypophthalmus based on Mi- tochondrial Cytochrome b Oxidase Subunit I (COI) gene from Pakistan.
- Hebert, P. D., Cywinska, A., Ball, S. L., & DeWaard, J. R. (2003). Biological identifications through DNA barcodes. Proceedings of the Royal Society of London. Series B: Bio- logical Sciences, 270(1512), 313-321.
- Al-Zafiri, B., Magdy, M., Ali, R. A. M., & Rashed, M. A. S. (2018). DNA Barcoding of Common Commercial Sea Catfish (Genus: Plicofollis) from Kuwait. American Journal of Mo- lecular Biology, 8(2), 102-108.
- Ebach, M. C., & Carvalho, M. R. D. (2010). Anti-intellectu- alism in the DNA barcoding enterprise. Zoologia (Curitiba), 27, 165-178.
- Ha, T. T. T., Huong, N. T., Hung, N. P., & Guiguen, Y. (2018).Species identification using DNA barcoding on processed panga catfish products in Viet Nam revealed important mis- labeling. Turkish Journal of Fisheries and Aquatic Sciences, 18(3), 457-462.
- Falade, M. O., Opene, A. J., & Benson, O. (2016). DNA barcoding of Clarias gariepinus, Coptodon zillii and Saroth- erodon melanotheron from Southwestern Nigeria. F1000Re- search, 5.
- Pouyaud, L., & Paradis, E. (2009). The phylogenetic structure of habitat shift and morphological convergence in Asian Clar- ias (Teleostei, Siluriformes: Clariidae). Journal of Zoological Systematics and Evolutionary Research, 47(4), 344-356.
- Marko, P. B., Lee, S. C., Rice, A. M., Gramling, J. M., Fitzhen-ry, T. M., McAlister, J. S., ... & Moran, A. L. (2004). Mislabel- ling of a depleted reef fish. Nature, 430(6997), 309-310.
- Panprommin, D., Soontornprasit, K., Tuncharoen, S., Pithak- pol, S., & Keereelang, J. (2019). DNA barcodes for the identi- fication of species diversity in fish from Kwan Phayao, Thai- land. Journal of Asia-Pacific Biodiversity, 12(3), 382-389.
- Iyiola, O. A., Nneji, L. M., Mustapha, M. K., Nzeh, C. G.,Oladipo, S. O., Nneji, I. C., ... & Adeola, A. C. (2018). DNA barcoding of economically important freshwater fish species from north-central Nigeria uncovers cryptic diversity. Ecolo- gy and evolution, 8(14), 6932-6951.
- Wong, L. L., Peatman, E., Lu, J., Kucuktas, H., He, S., Zhou, C., ... & Liu, Z. (2011). DNA barcoding of catfish: species authentication and phylogenetic assessment. PLoS One, 6(3), e17812.
- Ardura, A., Planes, S., & Garcia-Vazquez, E. (2013). Appli- cations of DNA barcoding to fish landings: authentication and diversity assessment. ZooKeys, (365), 49.
- Avise, J. C., Arnold, J., Ball, R. M., Bermingham, E., Lamb, T., Neigel, J. E., ... & Saunders, N. C. (1987). Intraspecific phylogeography: the mitochondrial DNA bridge between population genetics and systematics. Annual review of ecolo- gy and systematics, 489-522.
- FAO, Fisheries and Aquaculture. 2009. Review of the state of the World Fishery Resources, Circular No. 942, Rev.2 FIRF/ C942, Rev. 2 (En), pp.110.
- Hogan, Z., Baird, I. G., Radtke, R., & Vander Zanden, M. J. (2007). Long distance migration and marine habitation in the tropical Asian catfish, Pangasius krempfi. Journal of Fish Bi- ology, 71(3), 818-832.
- Ko, H. L., Wang, Y. T., Chiu, T. S., Lee, M. A., Leu, M. Y.,Chang, K. Z., ... & Shao, K. T. (2013). Evaluating the accura- cy of morphological identification of larval fishes by applying DNA barcoding. PLoS One, 8(1), e53451.
- Lenormand, S. (1996). Les Pangasiidae du delta du Mekong (Viet Nam): description preliminaire des pecheries, elements de biologie, et perspectives pour une diversification des el- evages. Cantho: Memoire de fin d.
- Nwani, C. D., Becker, S., Braid, H. E., Ude, E. F., Okogwu, O. I., & Hanner, R. (2011). DNA barcoding discriminates fresh- water fishes from southeastern Nigeria and provides river sys- tem-level phylogeographic resolution within some species. Mitochondrial DNA, 22(sup1), 43-51.
- Rasmussen, R. S., Morrissey, M. T., & Hebert, P. D. (2009). DNA barcoding of commercially important salmon and trout species (Oncorhynchus and Salmo) from North America. Journal of agricultural and food chemistry, 57(18), 8379-8385.
- Victor, B. C., Hanner, R., Shivji, M., Hyde, J., & Caldow, C. (2009). Identification of the larval and juvenile stages of the Cubera Snapper, Lutjanus cyanopterus, using DNA barcod- ing. Zootaxa, 2215(2), 24.
- Ward, R. D., Zemlak, T. S., Innes, B. H., Last, P. R., & Hebert,P. D. (2005). DNA barcoding Australia's fish species. Philo- sophical Transactions of the Royal Society B: Biological Sci- ences, 360(1462), 1847-1857.
- Zhang, J., & Hanner, R. (2012). Molecular approach to the identification of fish in the South China Sea. PLoS One, 7(2), e30621.


