Review Article - (2022) Volume 3, Issue 1
Microbial Clock: A review on forensic microbiology for crime scene investigations
2Professor, Head of the Department, Department of Biochemistry and Forensic Science,, Gujarat University, Ahmedabad 380009, India
Copyright: ©Copyright: �2022 Dr. Rakesh Rawal. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Khushboo Gautam, Rakesh Rawal. (2022). Microbial Clock: A review on forensic microbiology for crime scene investigations. In J Fore Res, 3(1), 112-120.
Abstract
As with other branches of forensic science, forensic microbiology is the study of the microbes associated with a cadaver to determine Time Since Death, Post Mortem Interval, and also identify the cause of death and personal identification. The advent of forensic microbiology and metagenomics has been aided by advances in genetic sequencing technologies, which have improved microbiological sample techniques and dramatically evolved bioinformatics pipelines. Humans carry a variety of microbial communities in and on their bodies, which constantly interact with and change their surroundings.
Main Body of the Abstract: We reviewed more than 80 papers that detailed various aspects of the Post-Mortem Microbiome in terms of the Microbial Clock to identify such as geolocation, personal identification, trace evidence, manner and cause of death, postmortem interval and Time Since Death. The fact that much of the research is carried out in mouse and swine models, which are often hard to emulate in human and real-time circumstances, is a key challenge for forensic medical and microbiological research.
Short Conclusion: To fully grasp its promise of individual Post-Mortem microbiome characterization in forensic sciences, we must include technology innovations that are either currently accessible or are being developed. Sophisticated microbial fingerprint and DNA recoverability would allow the detection of both victims and perpetrators, which are all greatly facilitated by a thorough understanding of the decomposition process. The review to address these obstacles is continuing, and microbiome-based evidence would be expected to play a role in crime scene investigations in the soon.
Keywords
Microbial clock, metagenomic, postmortem interval (PMI), Time since death (TSD), Microbiome, Forensic Microbiology, Trace Evidence, geolocation, Microbial fingerprint, decomposition
Background
What happens, after death? The formation of a huge, sophisticated ecosystem occurs as a result of death. After death, the natural process of decomposition includes physical and physiological changes that reveal crucial information about the Time Since Death (TSD), Post-Mortem Interval (PMI), as well as the location and manner of death. The corpse begins to decompose within 2–3 minutes of death. The start is controlled by a mechanism known as autolysis-Alternatively, self-digestion. This process begins and proceeds more quickly and that have a lot of enzymes. Autolysis normally takes a few days to manifest it visually. The onset of fluid-filled blisters on the skin and skin slippage, in which huge sheets of skin slough off the body, are the initial signs. (Roy et al., 2021) However, the body has adjusted to its surroundings (algor mortis), blood has accumulated in the body, generating skin darkening (livor mortis), and Because of the increasing alkalinity, the cytoplasm of cells has dissolved (rigor mortis). When a large enough number of cells burst, nutrition fluids emerge available, and the putrefaction might occur (Vass, 2001). Microorganisms (bacteria, fungi, and protozoa) break down the body's soft tissues, causing tissue catabolism in gases, liquids, and simple molecules. The activity of microbes (bacteria, fungi,and protozoa) upon its body's fatty tissue causes putrefaction, which is the catabolism of tissue within gases, liquids, and simple molecules. The production of sulfhaemoglobin in settled blood usually causes a greenish tint of the skin as the first obvious sign of putrefaction. As a result of the production of gas, the process can cause tissue distension like (hydrogen sulphide, carbon dioxide, methane, ammonia, sulphur dioxide, and hydrogen) in the bowels, but also seen it in the skin , several body parts, including the face, lips, and mouth and abdomen. Anaerobic fermentation is correlated to this. Predominantly in the intestines, yielding high- fat by-products Butyric and propionic acids are the most volatile fatty acids.(Pechal et al., 2018) The intestines are shown to be an overthrow of gas and liquids. Putrefaction is mainly from the rectum; however it can be severe enough to pull the skin apart, inflicting more damage. Further crack-up of protein and lipids phenolic compounds and glycerols are produced. Compounds, indole, 3-methylindole (skatole), putrescine and others. It has been observed that there is a protein like cadaverine as well as other volatile fatty acids present at the time of decomposition. At such a phase in the decomposition cycle, electrolytes are rapidly depleted and the body excretes both aerobic and anaerobic byproducts. There are a lot of bacteria, and there's a lot of insect activity in this phase (Tsokos, 2005). Even without activities of scavengers, bacteria, archaea, fungi, and Protista, the chemical breakdown of biodegradable waste would be tremendously slow, resulting in biochemical waste repositories.(Belk et al., 2018a) In the last millions of years, the reliability of mammalian carcasses has co- evolved with the evolution of microbial decomposing organic matter is relied upon to bring about preservation of key biochemical metabolic pathways and cross-realm biological collaborations for productive reusing of supplement saves(Belk et al., 2018a). After the host has died, the thought of the microbiome composition functioning as a biomarker has been utilized, and it is especially significant in the field of forensic sciences.(Can et al., 2014) .We have sought a systematic overview of the potential applicability of the Post-mortem Microbiome in forensic medicine and Forensic Microbiology, emphasizing on the various microbial marker profiles, and their robustness as well as limits, with an insistence on the TSD, PMI , manner of death and personal identification. The assessment of TSD is critical in creating relevant information about the time of death, which is required information to be submitted in a court of law. It's also one of the biggest greatest difficulties for forensic experts all over the world, because there are just too many variables that influence postmortem alterations, making it impossible to depend on a single approach. In this review paper, we look at various elements of current Post-Mortem Microbiome research. To begin, we examine the methodologies utilized to profile the Post-Mortem Microbiome and compare them to state- of-the-art approaches employed in other microbiome research. Following that, we go through significant discoveries in Post- Mortem Microbiome community features, focusing on the variety and resilience of microbial communities, suggested microbial markers, and the utility or lack thereof proposed prediction models and we are also focusing on knowledge gaps and limitation of the study. We also discussed the different technologies and methodologies that have been used for microbial profiling, as well as the novel approaches and techniques that are emerging in Post- mortem microbiome research.
Main Text
The Microbiome of the Human Postmortem
The utilization of cadaver associated microbial populations to determine PMI is a relatively novel technique for calculating TSD that has been investigated by numerous researchers across the globe. The factors that induce mammalian disintegration are identical and predictable(DeBruyn and Hauther, 2017a). At least for few ambits across distinct hosts and settings ambits. Because mammalian cadavers are such a valuable resource. Communities of microbes are likely to be a source of nutrients in an environment. Bacteria have evolved over millions of years ago to become efficient. These micronutrients should be recycled. As a result, mammalian bre(Metcalf, 2019) (Hyde et al., 2015). As a result, mammalian disintegration might contain a predictable sequence of microorganisms that represents the various stages of metabolic activity and disintegration routes.(Adserias-Garriga et al., 2017) (Pechal et al., 2018) (Johnson et al., 2016). Actually, microbiome data (epigenetic composition of microorganisms) has been used in a lot of researches. During the decomposition of modern artiodactyls, the species composition changes (Metcalf et al., 2016) (Javan et al., 2019) (Damann, Williams and Layton, 2015). A majority of studies with appropriate power have used this method. Approaches to Computational learning to construct(Johnson et al., 2016). Several studies have employed computational methodologies with sufficient power to generate prediction models that would be used to estimate the TSD of samples with unknown post mortem sample. According to (Metcalf, 2019) Several Methods rely on the microbiome have established accurate PMI estimations during some PMI ranges, as elevated as +/- 2–3 days, and measurable inaccuracy rates.
Microbial clock
There is still forensic entomology (Tomberlin et al., 2011)and an entomological clock which have been utilized such that the adult flies come in and lay their eggs on the first day(Catts and Goff, 1992)(Catts and Goff, 1992). After three to seven days, the eggs hatch into initial larval instars, which are somewhat larger than the pupae that emerge after Eight to Nine days. This clock is temperature dependent, " which means it’s quicker in warm temperatures and shorter in cold weather, but irrespective, if the maggots from within a body, the species, and the size, you can often calculate and if you know the temperature on how long those flies larvae have been on the body, and that has been recycled in court 100 times(Megyesi, Nawrocki and Haskell, 2005). Furthermore, because of temperature, seasonal, and developmental variations, they are challenging to implement in all situations. Most of these forensically significant insects have little information on their geolocation variation(Lindgren et al., 2011). Even though they are inevitable or have pre-determined ecologies, the postmortem microbiome has already shown a ton of potential as an Even if they are unavoidable or have predetermined ecologies, the postmortem microbiome has already demonstrated a great deal of promise as a novel, conventional, and accurate and consistent approach to playing an important role in the prediction of TSD and PMI, based on the idea that microbial succession changes in a determined and clock-like context throughout vertebrate species, even within the same eco system, As a result of the decomposition cycle (He et al., 2019). The cycling of various microbial species, such as entomology(Wells, 2019), might signify PMI and TSD.

Figure 1: Illustration of Microbial Clock
Distinct sampling locations have different microbial clocks
Research on microbial community change is also addressed in samples correlated to mammalian corpse skin. Based on the stage of decomposition as well as the complexity of the death scene, various sampling location intends to assess may be more or less accessible(Gulnaz T. Javan et al., 2016). The gut microbes(Jovel et al., 2016) (Yatsunenko et al., 2012) in your resident microbes are the ones that are active in decomposition from day one, converting all those cellular products and so they are quite occupied, and subsequently they are flushed into the soil as well as all the decomposition fluid when they go down(Yatsunenko et al., 2012)(DeBruyn and Hauther, 2017a)(Schloissnig et al., 2013). The following community we have to consider is the resident soil microbes, which are microbes that are just normally present in the soil and also have received to flush nutrient rich into the system, so microbes that can take advantage of this that grow quickly and can bloom under these scenarios and may be are really not sensitive to the elevated ammonium content will absolutely do so well, then , get a bloom of such opportunist soil microbes that participate in the third party to consider is all the visitors to the cadaver , as well as the entire ecosystem, the network of different species that will come to feed off of the cadaver, and they all have their own commensal microorganisms, so some of those may be getting into the system as well(Blum, Zechmeister-Boltenstern and Keiblinger, 2019)(Zhou et al., 2018)(Daniel, 2005).Additionally, throughout a death enquiry, certain sampling locations would be less intrusive. Skin samples, for example, might be a little more approachable and a little less intrusive then gastrointestinal samples during a death enquiry. While utilizing microbial clocks as nothing more than a forensic tool, all of these elements must be taken into account. Throughout the sections below, we go over these factors in further depth for each sampling type. (Ford, 1901)
Epinecrotic Community
The epinecrotic population(Benbow et al., 2019) was described by (Pechal et al., 2014) as microbes that live or move on the surface of decomposing corpses, such as the mucous membrane (Bizzarro et al., 2013)(e.g., mouth, buccal cavities). Therefore, as there are so many exterior sites to sample from, it's important to consider when bacteria from diverse locations decompositions in similar ways. In skin (Johnson et al., 2016) (Kodama et al., 2019) areas have been studied to see whether they can enhance the precision of PMI estimations and thus the microbial clock and he found that skin (Grice et al., 2008)sampling from the inner ear canals had lower error rates than skin (Grice and Segre, 2011)samples from the inner nasal surfaces when using bioinformatics pipelines (Kaszubinski et al., 2020).Although such encouraging findings imply whether the skin(Tozzo et al., 2020) could develop a precise microbial clock.
Internally location
An extrinsic variable like pH, temperature, insects, and scavengers may not have the same impact on the internal sites as they do on external sites. The intrinsic Post-Mortem Microbiome is also important would not be harmed by gut-related microorganisms that continue to thrive after death.(Can et al., 2014). The gut microbial community is also being looked into as a viable site of microbial activities changes.(DeBruyn and Hauther, 2017b). Even as a consequence, accumulating a time series of microorganisms into the bones of a cadaver could be used to follow the progression of bone samples, allowing for the creation of a microbiological clock for something like a prolonged period of death.(Damann, Williams and Layton, 2015) and his colleagues studied more than 1 corpses intended to decay over a long period of time. This allowed for the identification of microorganisms, and decomposition took place in a series over a long period of time. Compared to what seems to be achievable if other sorts of samples, like brain, and skin, are also absorbed by microbial decomposers, earlier Accumulated degree days (ADDs).
What and how to expect barriers and gaps in knowledge and how to address them
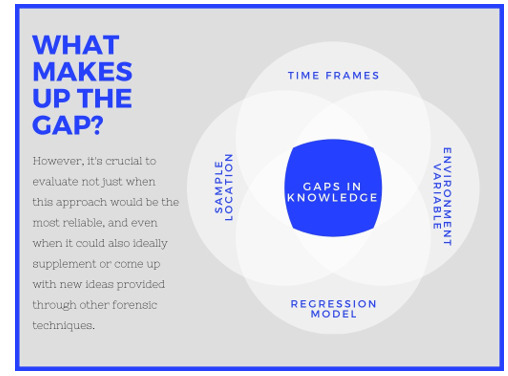
Figure 2: Knowledge gaps in Post-Mortem microbiome
Throughout the realm of forensic microbiology, there are a few issues to overcome. (Metcalf 2018) Recently identified certain gaps that must be addressed before the post-mortem microbiome can be used to estimate PMI (Metcalf 2018). Further possible medicolegal uses can be applied in the same way.
Time frames
In fresh stage: It's unclear when a cycle of decomposing microbes begins to eradicate the human skin microbiome's distinctive traits, but considerable evidence shows it is not prolonged (Fierer et al., 2010)(Grice et al., 2008) (Grice and Segre, 2011).(Lauber et al., 2014)
From the beginning of decomposition : Even though community productivity is prolonged and would seem to be a comparatively universally applicable approach-related groups of microbes to become predominant in a continuous path which is associated with time, temperature, and possibly those certain factors-such a time frame is effective for effectively utilizing the habitat of microbial cascades(Metcalf et al., 2016) (Metcalf et al., 2013)(Amendt et al., 2007). As a consequence, ancillary strategies such as using microbes as a protein supplement are becoming more popular. Although microorganisms can participate actively for long periods of time, chronological study is significant (Metcalf et al., 2016)
Skeleton Stage : Many of the most complex cases to investigate are those where the corpse has been decomposed for a long time (Damann, Williams and Layton, 2015). As a result, this historical period is favourable with opportunities to develop new forensic science approaches.
Environmental factor: Temperature is probably the sole breakdown robust model (Carter, Yellowlees and Tibbett, 2008) Mark Eric Benbow et al. Perhaps as an outcome, other abiotic factors such as gas, temperature, atmospheric conditions, and the existence or non-existence of bugs would all be taken into account. (Belk et al.)(Carter, Yellowlees and Tibbett, 2008)(Metcalf et al., 2016)(Pechal et al., 2018) In future research, these attributes will be evaluated to see if they are relevant.
Sample location
Microbial community changes in mammalian corpse surface (Hyde et al., 2015)(Metcalf et al., 2013)(DeBruyn and Hauther), buccal cavity(Pechal et al., 2018) (Hyde et al., 2015), nose(Johnson et al., 2016), ocular, (Pechal et al., 2018), bone (Johnson et al., 2016) (Gulnaz T Javan et al., 2016) (Damann, Williams and Layton, 2015), and corpse-associated soils (Metcalf et al., 2016)(Cobaugh, Schaeffer and DeBruyn, 2015)(Gulnaz T Javan et al., 2016)(Pechal et al., 2013) have all been studied. Each of these sites has shown potential, and a research that compared skin and soil samples found that neither sample was obviously superior to the other.(Grice and Segre, 2011). Certain locations may be more appropriate within specific times since death time periods.
Regression Model
The need for computational methods to create model-based predictive models is intriguing, data differences have been observed due to the limited amount of training data with an adequate sample size to generate a viable model(Schouls, Schot and Jacobs, 2003) (Belk et al.)
An overview of technology (past, present, and future)
Forensic Medicine and genetics have an ancient legacy of joint efforts. Because of breakthroughs in metagenomics (Mason-Buck et al., 2020), there's also been a lot of literature on the human microbiome. Sir “Alec Jeffrey’s discovered DNA(Deoxyribonucleic Acid) fingerprinting (Roewer, 2013)for human identification (Børsting and Morling, 2016). Years prior to the actual discovery of the short tandem repeat (STR) assays(Sullivan, Hopgood and Gill, 1992)(Mitchelson, 2003). The precise polymerase chain reaction (PCR)–based mtDNA (mitochondrial DNA)sequencing approach has become the ideal means of interpreting limited amounts of deteriorated sample materials, such as hair(Sullivan, Hopgood and Gill, 1992) (Susanne and Kristufek, 1993) and ancient bones, micrograms of desired adhering DNA are needed for restriction fragment length polymorphism analysis(Malla et al., 2019). However, contemporary approaches to sequencing of post-mortem microbiomes have exhibited a significant change toward better and more sophisticated molecular methods. DNA extraction kits, PCR techniques(Matsuki, 2007) , Next-generation Sequencing, Pyrosequencing (Lauber et al., 2009), metagenomics, and whole-genome sequencing are among them. The QIAamp DNA minikit (Qiagen) , QIAamp DNA microbiome Kit , Gentra Puregenes Kits is one of the widely viable(Bag et al., 2016) Dna extraction procedures (Claassen et al., 2013) (Soliman et al.,2017). Group-specific primers(Matsuki et al., 2002) have indeed been created and effectively utilized to determine the relative numbers of major microbial species for the applicability of TSD (Matsuki et al., 2004).
Figure 3: A Glimpse of handful of the exiting techiniques for post- microbiome Analysis
A sequence of modifications employing usable common refrain applications is necessary for the data analysis making judgments. These judgments have an impact on the data's outcomes and interpreting, therefore any use of the Post-mortem microbiome in the courts of law would necessitate shortened policies and protocols.(Zolfo et al., no date) (Gulnaz T Javan et al., 2016). Previous research has mostly used 16SrRNA(DeBruyn and Hauther) sequencing since it provides reliable data, is cost- effective and worthwhile, and can be used on a variety of data, from screening microbial signatures to predicting the place and time of death(Chakravorty et al., 2007). (Kuiper, 2016) whole genome sequencing (mWGS) , multilocus sequence (Belén, Pavón and Maiden, no date)method (Fiehn, 2002)(Bikel et al., 2015) data analysis validity and reliability(Kaszubinski et al., 2020). Tracing evidences, bio-crimes and mode of death are the major impacts by the microbial community. In contrast, a study of the literature identified a wide range of research investigating systematic microbial ecological succession variations in the tissue expression of genes in phases of decomposition in humans(Finley, Benbow and Javan, 2015). Throughout the last 20 years, technological advancements, especially massively parallel sequencing, metagenomics, and computational biology advances, have made it possible to interpret microorganisms for a diversity of human forensic applications, including personal identification, body fluid characterization, PMI estimation, time of death, geolocations(Hedrick et al., 2000), and bio-crimes encompassing potentially pathogenic real-time monitoring (Fu et al., 2018)(Kim et al., 2017). Therefore, as a result, applied scientific approaches to the study of microbiology evidence in civil and criminal cases for the purpose of crime scene investigation must be classified as forensic microbiology. (Robinson et al., 2020).
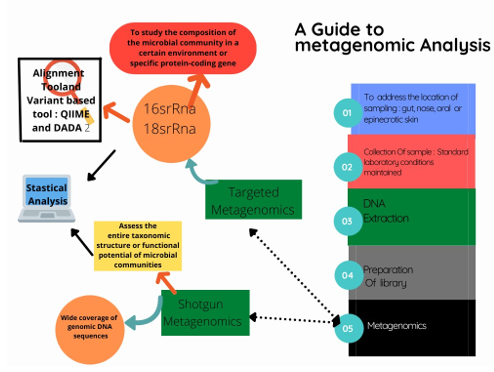
Figure 4: A schematic Guide to Metagenomic Analysis
Limitation of Technology
Next-generation sequencing (Trivedi et al., 2014) have made it feasible to conduct more cost-effective analyses of microbial communities. Nucleic acid extraction procedures used in sample preparation continue to be a significant source of bias toward specific microbial taxa. Identifying microorganisms is one of the most difficult challenges in huge, sophisticated metagenomic samples(Quince et al., 2017), distinguishing those of involvement from nearby microbial species and the immense cluttered background that makes up a microbial sample (Jo, Oh and Park, 2020) taken from different body sites, and determining the level of certainty that could be given to a promising species identified in just such a wide variety of complex samples.
Figure 5: Estimated Classification of Machine Learning Model
An Innovative Step up The Ladder to Forensic Microbiology:
The validity of microbiological evidence and the collecting of samples are the key steps in introducing the Postmortem- microbiome into the "Palais de Justice" proceeding(Ventura Spagnolo et al., 2019). There are currently no regulations or acts governing further use of post-mortem microbiome evidence in court cases(Schmedes, Sajantila and Budowle, 2016) (Hao, Pei and Brown, 2017). The postmortem microbiome can also be used as expert testimony since it has been thoroughly validated, has widely accepted regulatory pathways, is widely accepted in the scientific establishment, and has widely recognized significant discernible accuracies or errors. Another challenge that is inclined to express itself as a stumbling block once microbiological evidence is brought out of the trial is the statistical analysis' intricacy. Prior to getting implemented, so each approach should be approved, not all of which are confined to its acuity, precision, reliability, quantitation limit, and interpretability.

Figure 6: An illustration of the way to integrate new technologies into the workplace
Innovative, advanced, customized equipment is indeed being developed that would be used by cops at a scene of crime to provide instant solutions about just how long the person has already been dead and to aid in the discovery of hidden burials.(D’Argenio, 2018). Regarding dataset clarification of complicated metagenomic datasets, more sophisticated sequencing technologies could be designed and standardized(Ye et al., 2012)Other omic studies like metagenomic(Whon et al., 2021), metabolomics(Pesko et al., 2020) and transcriptomics(Bikel et al., 2015) are perhaps more approachable as previously, and Post-mortem Microbiome(Gulnaz T. Javan et al., 2016) study within those domains must be explored more as it will provide insights(Mason-Buck et al., 2020).
Conclusion
The main advantage of this the Existing TSD and PMI estimation methods, as well as gross postmortem alterations and entomological clocks, have the least probability as decomposition progresses but The above mentioned newest advanced technologies, such as Next Generation Sequencing, profile microbial communities and computational method that would use this information for creation of models which are generalizable across various environments similarly as internal external etc. However determination of TSD has never been simple but not impossible too. The main advantage is to include universally applicable microbiological models that is peculiar for every setting into the legal framework and contributes in crime scene investigation in the nearish future.
Abbreviations
TSD: Time Since death
PMI: Post-Mortem Interval
DNA: Deoxyribo Nucleic Acid
PCR: Polymerase Chain Reaction
mtDNA: mitochondrial DNA
Ethics Declarations
Ethics Approval and Consent to Participate
Not Applicable
Consent for Publication
Not Applicable
Availability of Data And Material
Not Applicable
Competing Interests
The Author declare that they have no competing interests
Funding
There is no such funding for this project. It is completely self- funded.
Authors Contribution
Khushboo Gautam reviewed literature; collected data and drafted manuscript. Dr. Rakesh Rawal edited the data and made necessary corrections. The Authors have read and accepted final manuscript Acknowledgement Authors acknowledge the Department of Biochemistry and Forensic Science, Gujarat University, INDIA. Thanks to Dr. Rakesh Rawal for his guidance and prompt feedback throughout the Process.
References
- Adserias-Garriga, J. et al. (2017) ‘Daily thanatomicrobiome changes in soil as an approach of postmortem interval estimation: An ecological perspective’, Forensic Science International, 278, pp. 388–395. doi: 10.1016/j. forsciint.2017.07.017.
- Amendt, J. et al. (2007) ‘Best practice in forensic entomology- Standards and guidelines’, International Journal of Legal Medicine, 121(2), pp. 90–104. doi: 10.1007/S00414-006- 0086-X/FIGURES/2.
- Bag, S. et al. (2016) ‘An improved method for high quality metagenomics DNA extraction from human and environmental samples’, Scientific Reports, 6. doi: 10.1038/SREP26775.
- Belén, A., Pavón, I. and Maiden, M. C. J. (no date) ‘Chapter11 Multilocus Sequence Typing’, Business, 551. doi: 10.1007/978-1-60327-999-4.
- Belk, A. et al. (2018a) ‘Microbiome data accurately predicts the postmortem interval using random forest regression models’, Genes, 9(2). doi: 10.3390/genes9020104.
- Belk, A. et al. (2018b) ‘Microbiome Data Accurately Predicts the Postmortem Interval Using Random Forest Regression Models’, Genes 2018, Vol. 9, Page 104, 9(2), p. 104. doi: 10.3390/GENES9020104.
- Benbow, M. E. et al. (2015) ‘The Potential of High-throughput Metagenomic Sequencing of Aquatic Bacterial Communities to Estimate the Postmortem Submersion Interval’, Journal of Forensic Sciences, 60(6), pp. 1500–1510. doi: 10.1111/1556- 4029.12859.
- Benbow, M. E. et al. (2019) ‘Necrobiome framework for bridging decomposition ecology of autotrophically and heterotrophically derived organic matter’, Ecological Monographs, 89(1). doi: 10.1002/ecm.1331.
- Bikel, S. et al. (2015) ‘Combining metagenomics, metatranscriptomics and viromics to explore novel microbial interactions: Towards a systems-level understanding of human microbiome’, Computational and Structural Biotechnology Journal, 13, pp. 390–401. doi: 10.1016/j.csbj.2015.06.001.
- Bizzarro, S. et al. (2013) ‘Subgingival microbiome in smokers and non-smokers in periodontitis: An exploratory study using traditional targeted techniques and a next-generation sequencing’, Journal of Clinical Periodontology, 40(5), pp. 483–492. doi: 10.1111/jcpe.12087.
- Blum, W. E. H., Zechmeister-Boltenstern, S. and Keiblinger,K. M. (2019) ‘Does Soil Contribute to the Human Gut Microbiome?’, Microorganisms, 7(9). doi: 10.3390/ MICROORGANISMS7090287.
- Børsting, C. and Morling, N. (2016) ‘Genomic Applications in Forensic Medicine’, Medical and Health Genomics, pp. 295–309. doi: 10.1016/B978-0-12-420196-5.00022-8.
- Can, I. et al. (2014) ‘Distinctive thanatomicrobiome signatures found in the blood and internal organs of humans’, Journal of microbiological methods, 106, pp. 1–7. doi: 10.1016/J. MIMET.2014.07.026.
- Carter, D. O., Yellowlees, D. and Tibbett, M. (2008) ‘Temperature affects microbial decomposition of cadavers (Rattus rattus) in contrasting soils’, Applied Soil Ecology, 40(1), pp. 129–137. doi: 10.1016/J.APSOIL.2008.03.010.
- Catts, E. P. and Goff, M. L. (1992) ‘Forensic entomology in criminal investigations’, Annual Review of Entomology, 37(1), pp. 253–272. doi: 10.1146/ANNUREV. EN.37.010192.001345.
- Chakravorty, S. et al. (2007) ‘A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria’, Journal of microbiological methods, 69(2), pp. 330–339. doi: 10.1016/J.MIMET.2007.02.005.
- Claassen, S. et al. (2013) ‘A comparison of the efficiency of five different commercial DNA extraction kits for extraction of DNA from faecal samples’, Journal of Microbiological Methods, 94(2), pp. 103–110. doi: 10.1016/J. MIMET.2013.05.008.
- Cobaugh, K. L., Schaeffer, S. M. and DeBruyn, J. M. (2015) ‘Functional and structural succession of soil microbial communities below decomposing human cadavers’, PLoS ONE, 10(6), pp. 1–20. doi: 10.1371/journal.pone.0130201.
- D’Argenio, V. (2018) ‘Human Microbiome Acquisition and Bioinformatic Challenges in Metagenomic Studies’, International journal of molecular sciences, 19(2). doi: 10.3390/IJMS19020383.
- Damann, F. E., Williams, D. E. and Layton, A. C. (2015) ‘Potential Use of Bacterial Community Succession in Decaying Human Bone for Estimating Postmortem Interval’, Journal of forensic sciences, 60(4), pp. 844–850. doi: 10.1111/1556-4029.12744.
- Daniel, R. (2005) ‘The metagenomics of soil’, Nature Reviews Microbiology, 3(6), pp. 470–478. doi: 10.1038/ NRMICRO1160.
- DeBruyn, J. M. and Hauther, K. A. (2017a) ‘Postmortem succession of gut microbial communities in deceased human subjects’, PeerJ, 2017(6). doi: 10.7717/peerj.3437.
- DeBruyn, J. M. and Hauther, K. A. (2017b) ‘Postmortem succession of gut microbial communities in deceased human subjects’, PeerJ, 2017(6), p. e3437. doi: 10.7717/PEERJ.3437/ SUPP-2.
- DeBruyn, J. M. and Hauther, K. A. (2017c) ‘Postmortem succession of gut microbial communities in deceased human subjects’, PeerJ, 2017(6), pp. 1–14. doi: 10.7717/peerj.3437.
- Fiehn, O. (2002) ‘Metabolomics - The link between genotypes and phenotypes’, Plant Molecular Biology, 48(1–2), pp. 155–171. doi: 10.1023/A:1013713905833.
- Fierer, N. et al. (2010) ‘Forensic identification using skin bacterial communities’, Proceedings of the National Academy of Sciences of the United States of America, 107(14), pp. 6477–6481. doi: 10.1073/pnas.1000162107.
- Finley, S. J., Benbow, M. E. and Javan, G. T. (2015) ‘Microbial communities associated with human decomposition and their potential use as postmortem clocks’, International Journal of Legal Medicine, 129(3), pp. 623–632. doi: 10.1007/s00414- 014-1059-0.
- Ford, W. W. (1901) ‘On the bacteriology of normal organs’, Journal of Hygiene, 1(2), pp. 277–284. doi: 10.1017/ S0022172400000231.
- Fu, X. L. et al. (2018) ‘Application of High-throughput Sequencing in Researches of Cadaveric Microorganisms and Postmortem Interval Estimation’, Journal of Forensic Medicine, 34(5), pp. 475–481. doi: 10.12116/j.issn.1004-5619.2018.05.004.
- Grice, E. A. et al. (2008) ‘A diversity profile of the human skin microbiota’, Genome Research, 18(7), pp. 1043–1050. doi: 10.1101/GR.075549.107.
- Grice, E. A. and Segre, J. A. (2011) ‘The skin microbiome’, Nature Reviews Microbiology, 9(4), pp. 244–253. doi: 10.1038/nrmicro2537.
- Hao, Y., Pei, Z. and Brown, S. M. (2017) ‘Bioinformatics in Microbiome Analysis’, Methods in Microbiology, 44, pp. 1–18. doi: 10.1016/BS.MIM.2017.08.002.
- He, J. et al. (2019) ‘Potential use of high-throughput sequencing of bacterial communities for postmortem submersion interval estimation’, Brazilian journal of microbiology : [publication of the Brazilian Society for Microbiology], 50(4), pp. 999– 1010. doi: 10.1007/S42770-019-00119-W.
- Hedrick, D. B. et al. (2000) ‘Measuring soil microbial community diversity using polar lipid fatty acid and denaturing gradient gel electrophoresis data’, Journal of Microbiological Methods, 41(3), pp. 235–248. doi: 10.1016/ S0167-7012(00)00157-3.
- Hyde, E. R. et al. (2015) ‘Initial insights into bacterial succession during human decomposition’, International Journal of Legal Medicine, 129(3), pp. 661–671. doi: 10.1007/ s00414-014-1128-4.
- Javan, Gulnaz T et al. (2016) ‘Human Thanatomicrobiome Succession and Time Since Death’, Nature Publishing Group. doi: 10.1038/srep29598.
- Javan, Gulnaz T. et al. (2016) ‘The thanatomicrobiome: A Missing Piece of the Microbial Puzzle of Death’, Frontiers in Microbiology, 7(FEB). doi: 10.3389/fmicb.2016.00225.
- Javan, G. T. et al. (2019) ‘An interdisciplinary review of the thanatomicrobiome in human decomposition’, Forensic Science, Medicine, and Pathology, 15(1), pp. 75–83. doi: 10.1007/s12024-018-0061-0.
- Jo, J., Oh, J. and Park, C. (2020) ‘Microbial community analysis using high-throughput sequencing technology: a beginner’s guide for microbiologists’, Journal of Microbiology, 58(3),pp. 176–192. doi: 10.1007/s12275-020-9525-5.
- Johnson, H. R. et al. (2016) ‘A machine learning approach for using the postmortem skin microbiome to estimate the postmortem interval’, PLoS ONE, 11(12), pp. 1–23. doi: 10.1371/journal.pone.0167370.
- Jovel, J. et al. (2016) ‘Characterization of the gut microbiome using 16S or shotgun metagenomics’, Frontiers in Microbiology, 7(APR), pp. 1–17. doi: 10.3389/ fmicb.2016.00459.
- Kaszubinski, S. F. et al. (2020) ‘Evaluating Bioinformatic Pipeline Performance for Forensic Microbiome Analysis*,†,‡’, Journal of Forensic Sciences, 65(2), pp. 513–525. doi: 10.1111/1556-4029.14213.
- Kim, B. R. et al. (2017) ‘Deciphering diversity indices for a better understanding of microbial communities’, Journal of Microbiology and Biotechnology, 27(12), pp. 2089–2093. doi: 10.4014/jmb.1709.09027.
- Kodama, W. A. et al. (2019) ‘Trace Evidence Potential in Postmortem Skin Microbiomes: From Death Scene to Morgue’, Journal of Forensic Sciences, 64(3), pp. 791–798. doi: 10.1111/1556-4029.13949.
- Kuiper, I. (2016) ‘Microbial forensics: nextâ?generation sequencing as catalyst’, EMBO reports, 17(8), pp. 1085– 1087. doi: 10.15252/embr.201642794.
- Lauber, C. L. et al. (2009) ‘Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale’, Applied and environmental microbiology, 75(15), pp. 5111–5120. doi: 10.1128/AEM.00335-09.
- Lauber, C. L. et al. (2014) ‘Vertebrate decomposition is accelerated by soil microbes’, Applied and Environmental Microbiology, 80(16), pp. 4920–4929. doi: 10.1128/ AEM.00957-14/SUPPL_FILE/ZAM999105551SO1.PDF.
- Lindgren, N. K. et al. (2011) ‘Exclusion of forensically important flies due to burying behavior by the red imported fire ant (Solenopsis invicta) in southeast Texas’, Forensic science international, 204(1–3). doi: 10.1016/J. FORSCIINT.2010.05.016.
- Malla, M. A. et al. (2019) ‘Exploring the human microbiome: The potential future role of next-generation sequencing in disease diagnosis and treatment’, Frontiers in Immunology, 10(JAN), p. 2868. doi: 10.3389/FIMMU.2018.02868/ BIBTEX.
- Mason-Buck, G. et al. (2020) ‘DNA Based Methods in Intelligence - Moving Towards Metagenomics’, Preprints, (February). Available at: https://search.proquest.com/ docview/2413932290?accountid=42404.
- Matsuki, T. et al. (2002) ‘Development of 16S rRNA- gene-targeted group-specific primers for the detection and identification of predominant bacteria in human feces’, Applied and environmental microbiology, 68(11), pp. 5445– 5451. doi: 10.1128/AEM.68.11.5445-5451.2002.
- Matsuki, T. et al. (2004) ‘Use of 16S rRNA gene-targeted group-specific primers for real-time PCR analysis of predominant bacteria in human feces’, Applied and Environmental Microbiology, 70(12), pp. 7220–7228. doi: 10.1128/AEM.70.12.7220-7228.2004.
- Matsuki, T. (2007) ‘Development of quantitative PCR detection method with 16S rRNA gene-targeted genus- and species-specific primers for the analysis of human intestinal microflora and its application’, Nippon saikingaku zasshi. Japanese journal of bacteriology, 62(2), pp. 255–261. doi: 10.3412/jsb.62.255.
- Megyesi, M. S., Nawrocki, S. P. and Haskell, N. H. (2005) ‘Using accumulated degree-days to estimate the postmortem interval from decomposed human remains.’, undefined, 50(3),pp. 1–9. doi: 10.1520/JFS2004017.
- Metcalf, J. L. et al. (2013) ‘A microbial clock provides an accurate estimate of the postmortem interval in a mouse model system’, eLife, 2, pp. 1–19. doi: 10.7554/elife.01104.
- Metcalf, J. L. et al. (2016) ‘Mammalian Corpse Decomposition’, Science, 351(6269), pp. 158–162.
- Metcalf, J. L. (2019) ‘Estimating the postmortem interval using microbes: Knowledge gaps and a path to technology adoption’, Forensic Science International: Genetics, 38(November 2018), pp. 211–218. doi: 10.1016/j.fsigen.2018.11.004.
- Mitchelson, K. R. (2003) ‘The use of capillary electrophoresis for DNA polymorphism analysis’, Molecular Biotechnology 2003 24:1, 24(1), pp. 41–68. doi: 10.1385/MB:24:1:41.
- Pechal, J. L. et al. (2013) ‘Microbial community functional change during vertebrate carrion decomposition’, PLoS ONE, 8(11), pp. 1–11. doi: 10.1371/journal.pone.0079035.
- Pechal, J. L. et al. (2014) ‘The potential use of bacterial community succession in forensics as described by high throughput metagenomic sequencing’, International journal of legal medicine, 128(1), pp. 193–205. doi: 10.1007/S00414-013-0872-1.
- Pechal, J. L. et al. (2018) ‘A large-scale survey of the postmortem human microbiome, and its potential to provide insight into the living health condition’, Scientific Reports, 8(1), pp. 1–15. doi: 10.1038/s41598-018-23989-w.
- Pesko, B. K. et al. (2020) ‘Postmortomics: The Potential of Untargeted Metabolomics to Highlight Markers for Time since Death’, OMICS A Journal of Integrative Biology, 24(11), pp. 649–659. doi: 10.1089/OMI.2020.0084/ASSET/IMAGES/ LARGE/OMI.2020.0084_FIGURE7.JPEG.
- Quince, C. et al. (2017) ‘Shotgun metagenomics, from sampling to analysis’, Nature biotechnology, 35(9), pp. 833– 844. doi: 10.1038/NBT.3935.
- Robinson, J. M. et al. (2020) ‘Forensic Applications of Microbiomics: A Review.’, Frontiers in microbiology, 11, p. 608101. doi: 10.3389/fmicb.2020.608101.
- Roewer, L. (2013) ‘DNA fingerprinting in forensics: Past, present, future’, Investigative Genetics, 4(1), pp. 1–10. doi: 10.1186/2041-2223-4-22.
- Roy, D. et al. (2021) ‘Microbiome in Death and Beyond: Current Vistas and Future Trends’, Frontiers in Ecology and Evolution, 9(March), pp. 1–24. doi: 10.3389/fevo.2021.630397.
- Schloissnig, S. et al. (2013) ‘Genomic variation landscape of the human gut microbiome’, Nature, 493(7430), pp. 45–50. doi: 10.1038/nature11711.
- Schmedes, S. E., Sajantila, A. and Budowle, B. (2016) ‘Expansion of microbial forensics’, Journal of Clinical Microbiology, 54(8), pp. 1964–1974. doi: 10.1128/ JCM.00046-16.
- Schouls, L. M., Schot, C. S. and Jacobs, J. A. (2003) ‘Horizontal Transfer of Segments of the 16S rRNA Genes between Species of the Streptococcus anginosus Group’, Journal of Bacteriology, 185(24), pp. 7241–7246. doi: 10.1128/JB.185.24.7241-7246.2003.
- Soliman, T. et al. (2017) ‘Profiling soil microbial communities with next-generation sequencing: the influence of DNA kit selection and technician technical expertise’, PeerJ, 5(12). doi: 10.7717/PEERJ.4178.
- Sullivan, K. M., Hopgood, R. and Gill, P. (1992) ‘Identification of human remains by amplification and automated sequencing of mitochondrial DNA’, International journal of legal medicine, 105(2), pp. 83–86. doi: 10.1007/BF02340829.
- Susanne, R. U. T. C.- and Kristufek, D. (1993) ‘Applied andenvironmental’.
- Tomberlin, J. K. et al. (2011) ‘A roadmap for bridging basicand applied research in forensic entomology’, Annual Review of Entomology, 56, pp. 401–421. doi: 10.1146/annurev- ento-051710-103143.
- Tozzo, P. et al. (2020) ‘Skin microbiome analysis for forensic human identification: What do we know so far?’, Microorganisms, 8(6), pp. 1–19. doi: 10.3390/ microorganisms8060873.
- Trivedi, U. H. et al. (2014) ‘Quality control of next-generation sequencing data without a reference’, Frontiers in Genetics, 5(MAY), p. 111. doi: 10.3389/FGENE.2014.00111/BIBTEX.
- Tsokos, M. (2005) ‘Postmortem Changes and Artifacts Occurring During the Early Postmortem Interval’, Forensic Pathology Reviews, pp. 183–238. doi: 10.1007/978-1-59259- 910-3_5.
- Vass, A. (2001) ‘Beyond the grave – understanding human decomposition’, Microbiology Today, 28(28), pp. 190–192.
- Ventura Spagnolo, E. et al. (2019) ‘Forensic microbiology applications: A systematic review’, Legal medicine (Tokyo, Japan), 36, pp. 73–80. doi: 10.1016/J. LEGALMED.2018.11.002.
- Wells, J. D. (2019) ‘A Forensic Entomological Analysis Can Yield an Estimate of Postmortem Interval, and Not Just a Minimum Postmortem Interval: An Explanation and Illustration Using a Case’, Journal of Forensic Sciences, 64(2), pp. 634–637. doi: 10.1111/1556-4029.13912.
- Whon, T. W. et al. (2021) ‘Omics in gut microbiome analysis’, Journal of Microbiology, 59(3), pp. 292–297. doi: 10.1007/ s12275-021-1004-0.
- Yatsunenko, T. et al. (2012) ‘Human gut microbiome viewed across age and geography’, Nature, 486(7402), pp. 222–227. doi: 10.1038/nature11053.
- Ye, D. D. et al. (2012) ‘[A review on the bioinformatics pipelines for metagenomic research].’, Dong wu xue yan jiu = Zoological research / ‘Dong wu xue yan jiu’ bian ji wei yuan hui bian ji, 33(6), pp. 574–585. doi: 10.3724/ sp.j.1141.2012.06574.
- Zhou, D. et al. (2018) ‘Soil is a key factor influencing gut microbiota and its effect is comparable to that exerted by diet for mice [version 1; peer review: 1 approved, 1 approved with reservations]’, F1000Research, 7. doi: 10.12688/ F1000RESEARCH.15297.1.
- Zolfo, M. et al. (no date) ‘Profiling microbial strains in urban environments using metagenomic sequencing data’. doi: 10.1186/s13062-018-0211-z.


