Research Article - (2021) Volume 2, Issue 3
LAMBEDA PHAGE
Received Date: Sep 07, 2021 / Accepted Date: Sep 15, 2021 / Published Date: Sep 24, 2021
Copyright: ©Copyright: �2021 Mahdi Tajalifar. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Citation: Mahdi Tajalifar (2021) LAMBEDA PHAGE. In j Fore Res, 2(3), 76-79.
Abstract
Background: Recognition of bacteriophages in many aspects plays an important role such as controlling the number and variation of bacteria and participation in horizontal gene transfer, which is an important process in bacterial evolution. Bacteriophages use small proteins to take over the host molecular machinery and thus interfere with central metabolic processes in infected bacteria. In general, phages inhibit or reverse host transcription to transcribe their genome. Mechanical and structural studies of phage host transcription may lead to the development of new antibacterial agents.
Result: The result shows that phage vB_Eco4M-7 must be a lytic virus. This was confirmed by monitoring faglitic development by a one-step growth test. In addition, phage of relatively small uniform plaques (1 mm in diameter) occur without lysogenesis. Electron microscopic analysis showed that vB_Eco4M-7 belongs to the family Myoviridae. Based on mass spectrometric analysis, including the fragmentation pattern of unique peptides, 33 vB_Eco4M-7 phage proteins are assigned to the annotated reading frames. The results indicate that the phage studied is a potential candidate for phage treatment and / or food protection against E. coli, which produce Shiga toxin, as most of these strains belong to the O157 serotype.
Conclusion: In general, phages inhibit or reverse host transcription to transcribe their genome. Mechanical and structural studies of phage host transcription may lead to the development of new antibacterial agents. The result shows that phage vB_Eco4M-7 must be a lytic virus. This was confirmed by monitoring faglitic development by a one-step growth test.
Background
Currently, only 9,083 phage genomic sequences are available from the National Biotechnology Information Center (NCBI) database, of which 325 are Pseudomonas phage sequences (as of January 14, 2020). Obviously, the number of isolated phages is very limited. In addition, the vast majority of them are lytic and the number of lysogenic phages is less. In addition, in the case of obtained phages, we know only a very small part of the function of the phage genes, which are mainly related to morphology and the lysis process. The genes responsible for regulating the interaction between phages and their hosts are highly unknown. Therefore, isolation of new temperate phages is required, and analysis of genomes and their life cycle leads to a better understanding of the relationship and simultaneous evolution between phages and bacterial hosts [1].
Here we introduce a new mild phage, YA3, and perform evolutionary analysis. Which indicates that YA3 is a new mild phage of P. aeruginosa and a new member of the lambdoid family. Isolation and in-depth study of the new lambdoid phage, YA3, helps to better understand the evolutionary relationship between temperate phages and their host [2]. An alternative strategy is to modify the function of the bacteria in the native environment. For example, one approach has been to develop drugs that target the virulence of antibiotic-resistant pathogens in order to specifically counteract their harmful effects while minimizing the choice of resistance. Although a number of antiviral drugs are being studied, inhibition targets are generally limited to those available by small, biological molecules (e.g., surface and secreted proteins) that may require multiple drugs. To target several pathogens. Target effects on other microbes and host [1].
Moderate phage using genetic engineering has been reported to suppress Stx from E. coli-based populations cloned in mammalian intestines. First, it shows that genetic hybrids between lambidoid phages can overcome phage resistance mechanisms by maintaining function. A Stx transcription repressor is then genetically encoded in its engineering phage, indicating that the Stx produced by E. coli MG1655 is significantly reduced in vitro. Finally, it shows that engineered phage, when injected into mice before colonization by this strain of E. coli, can be dispersed from a single dose throughout the rat intestine to significantly increase the concentration of fecal Stx. Reduce. This work describes a new therapeutic framework for in situ modification of intestinal bacteria for genetically based antiviruses [1].
Phage therapy is a promising alternative to antibiotics, there are several factors that limit the use of this treatment. First, a wide range of different phages is needed to provide appropriate treatment options for many patients with various infectious diseases. Second, not every phage is suitable for phage therapy. In particular, mild phages should not be used in this method because they can lyse the host cells instead of lysis. Third, some phages carry toxic genes into their genomes and therefore cannot be used as therapeutic agents. Fourth, it is possible that bacteria are resistant to phages. Despite the limitations described above, the antibacterial activity of bacteriophages is very attractive in combating unwanted bacteria. Therefore, apart from their use in the treatment of human diseases, the use of these viruses in the protection of food, agriculture and industry is also considered. Although the vast majority of Escherichia coli strains are common to humans, pathogenic strains also belong to this species. Among them, E. coli producing Shiga toxin (STEC), including the enterohemorrhagic E. coli group (EHEC), appear to be some of the most dangerous pathogens. While the first signs of infection by these bacteria may be unpleasant but not dangerous (dysentery), complications such as hemolytic uremic syndrome, thrombocytopenia, and bleeding colitis can cause severe symptoms or even death in patients. In fact, local STEC epidemics could be devastating, such as the one in Germany in 2011, which killed more than 50 people [3].
Bacterial transcription machines have been a constant target for antibiotics. In addition to RNA polymerase (RNAP) itself, bacteria also have the transcriptional regulation of a number that is needed. While there are sequences and construction similarities between bacterial and eukaryotic RNAPs, the molecular machine consists of transcriptional regulatory layers between very different areas of life. In fact, only the NusG / Spt5 factor appears to have been saved from the overall transcription lengthening level. Thus, can transcriptional regulatory factors or transcriptional complexes modified by specific regulatory agents be very attractive to develop new antimicrobial agents. However, because only this agent has controllable enzymatic activity, it is difficult to target this regulatory layer with traditional inhibitor screening marketing. Due to the increasing number of three-dimensional structures with high resolution of the transcription factor and their complexes, silicon screening methods can be set up. Bacteriophages create large reservoirs of unique proteins that connect different molecular bacterial machines to allow the release of phages. Many of these phage-inducing protein modulators do not show any conformity with the proteins produced in all three kingdoms of life. So you target such molecules of effect of the host molecular machinery with the metabolic pathways you need, ultimately leading to the complete shutdown of the host metabolism and the destruction of the host bacteria. The need for host molecular machinery is one of the main criteria that should be considered when selecting a potential drug target, and the identification of new phage proteins can lead to the need for an interesting new drug in pathogenic disease [4].
Result
1- Several phage lysogenic genes have been found in the YA3 genome, including orf22 (site-specific integrase), orf23 (oxygenase), and orf35 (exonuclease). Using HHpred analysis, these three lysogen-related genes are similar to lambdoid phage proteins. Site-specific integrase (ORF22) is similar to enterobacteriaceae phage lambda with (100% probability) and 1-enterobacterial phage P2 with (100% probability). Excisionase (ORF23) is similar to Enterobacteria phage lambda excisionase (98.56% probability) and Enterobacteria phage P2 (98.68% probability). Exonuclease (ORF35) is similar to Enterobacteria phage lambda exonuclease (100% probability). In addition, three ORF products are proteins that, after analysis using HHpred, have a sequence with the CI, Cro, and Q proteins of lambdoid phages. Homologous CI (ORF50, CI suppressor protein) is a 236 amino acid protein that is similar to CI Enterobacteria phage lambda (99.58% probability) and Enterobacteria phage 186 (98.67% probability), respectively. Cro homolog (ORF51, which regulates Cro transcription) is a 59 amino acid protein, which is similar to the Cro regulatory protein of Enterobacteria phage P22 (probability 98.76%), phage inhibitor protein N15 (probability 98.95%), and lambda suppressor Cro of Enterobacteria phage lambda (98.46 probability probability), respectively. The homologous Q (ORF61, Q protein against Q protein) is a 139 amino acid protein that is similar to the anti-terminator Q protein of Enterobacteriaceae (99.8% reliability).
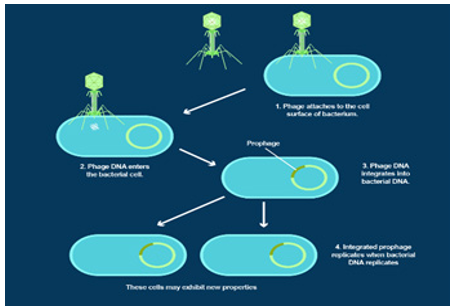
Figure 1: Lysogenic Phage
2- Modifying the function of bacteria with species and genetic characteristics requires great care, especially in their natural ecosystem. While the complexity and heterogeneity of the mammalian gut make it a challenging environment for targeted changes, phages have adapted to the high bacterial host characteristics, making them an attractive therapeutic tool. The use of phages is not just to kill bacteria, but phages are used in the phage therapy approach. The specific function of the desired bacteria is modified in the mammalian intestine. Because antibacterial approaches can increase resistance, the goal is to engineer mild copper phages.
Figure 2: Phage Lambeda
As an alternative to bacteriolytic strategies aimed at preventing pathogenesis by killing bacteria, an alternative approach is proposed that aims to reduce the expression of the pathogen. By introducing phage to lysogenize the bacteria in the mammalian intestine and expressing a repressor, transcription of the pathogen is performed. (B) (i) E. coli933W genetic designs showing the 933W profile to show cI to maintain a lysogenic state in which stx2 is not expressed; and (ii) induction that degrades cI protein and results in The expression of lytic genes includes cro. And stx2. (iii) This leads to cell lysis and the release of phage and Stx2 proteins. Expression of an unanalyzed CI for the 933W, 933W. cIind- prophage from a genomically engineered phage moderated (antiviral prophylaxis) can induce the 933W progelgal to remain lysogenic despite induction and degradation of endogenous cI protein.
3- Bioinformatics analysis of the vB_Eco4M-7 genome shows that it contains 96 open frames (ORF), of which 62 ORF are in the direct strand of the virus genome and 34 ORF are in the complementary strand. Among all the identified ORFs, 35 were assigned based on their amino acid sequence compliance to known proteins or evolutionary conserved protein domains and roles, hypothetical functions (36.5). Approximately 63.5% of ORFs (61 ORFs) were classified as hypothetical proteins with unknown function. The vB_Eco4M-7 phage genome does not contain sequences of genes encoding integrase, recombinant, inhibitors, and oxygenase, which are major markers of moderate viruses. As a result, phage vB_Eco4M-7 should be considered a lytic virus. In addition, the results obtained after genetic screening of the vB_Eco4M-7 genome based on the pathogenicity database of pathogenic bacteria indicate that this E. coli phage lacks genes encoding toxins and other pathogens that may affect cells. Eukaryotic effect. This is an important sign because the detection of any of the genes associated with pathogenesis by genetic screening directly rejects the vB_ Eco4M-7 phage for therapeutic use.
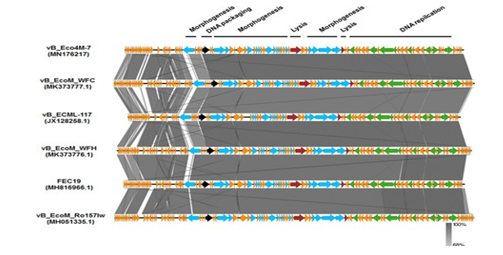
Figure 3: ECO4M
4- Lambda phages have long been known to use strategies to modulate host RNAP stop and stop when transcribing the phage genome. For example, most of these phages use N and Q proteins to facilitate the shift of early gene expression to early delay and to support the expression of late genes throughout their lytic life cycle. Therefore, N and Q proteins interact with host RNAP, transcription factors, RNA, and / or DNA, and modify ECs steadily to stop and terminate Ruse resistance, thus allowing it to pass through Read endogenous and genetic terminals, even if the terminators are located downstream of the site where N or Q are initially loaded on the EC (process termination). Map a portion of the λ phage genome (thick black line) including early and late control regions. The size of areas and elements and their position are not plotted on a scale. Open boxes with names-protein encoding regions; Narrow black boxes, bead DNA; Black angle arrows, promoters; Red stop signs, intrinsic terminals (“i”) and ρ-dependent (“ρ”); Green markers, regulatory regions active as DNA (QBE and SDPE); Dark yellow boxes, vertebral regulatory regions active as RNA; Turquoise butter, anti-termination proteins; Turquoise angled arrows, the sites of adsorption of anti-termination proteins in the EC. Dark yellow lines, transcription. Map adapted to changes.
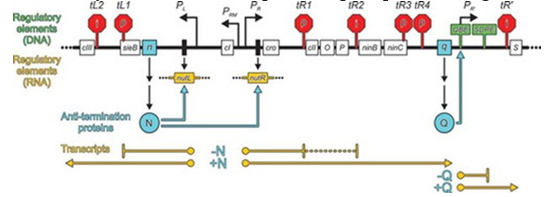
Figure 4: Biological activities of lambdoid phage N and Q proteins.
Conclusion
Currently, only 9,083 phage genomic sequences are available from the National Biotechnology Information Center (NCBI) database, of which 325 are Pseudomonas phage sequences (as of January 14, 2020). Here we introduce a new mild phage, YA3, and perform evolutionary analysis. Which indicates that YA3 is a new mild phage of P. aeruginosa and a new member of the lambdoid family. Phage therapy is a promising alternative to antibiotics, there are several factors that limit the use of this treatment. First, a wide range of different phages is needed to provide appropriate treatment options for many patients with various infectious diseases. Second, not every phage is suitable for phage therapy.
Back ground: Bacterial transcription machines have been a constant target for antibiotics. In addition to RNA polymerase (RNAP) itself, bacteria also have the transcriptional regulation of a number that is needed. While there are sequences and construction similarities between bacterial and eukaryotic RNAPs, the molecular machine consists of transcriptional regulatory layers between very different areas of life. In fact, only the NusG / Spt5 factor appears to have been saved from the overall transcription lengthening level.
Result: Several phage lysogenic genes have been found in the YA3 genome, including orf22 (site-specific integrase), orf23 (oxygenase), and orf35 (exonuclease). Using HHpred analysis, these three lysogen-related genes are similar to lambdoid phage proteins. Modifying the function of bacteria with species and genetic characteristics requires great care, especially in their natural ecosystem. While the complexity and heterogeneity of the mammalian gut make it a challenging environment for targeted changes, phages have adapted to the high bacterial host characteristics, making them an attractive therapeutic tool. This is an important sign because the detection of any of the genes associated with pathogenesis by genetic screening directly rejects the vB_Eco4M-7 phage for therapeutic use. Lambda phages have long been known to use strategies to modulate host RNAP stop and stop when transcribing the phage genome. For example, most of these phages use N and Q proteins to facilitate the shift of early gene expression to early delay and to support the expression of late genes throughout their lytic life cycle.


