Research Article - (2024) Volume 2, Issue 3
Karyological Study of Fresh Water Fish Channa Punctatus with G and NOR Banding Pattern
Received Date: Jan 10, 2024 / Accepted Date: Feb 15, 2024 / Published Date: Mar 13, 2024
Copyright: ©Â©2024 Verma V K. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Verma, V. K. (2024). Karyological Study of Fresh Water Fish Channa Punctatus with G and NOR Banding Pattern. Int Internal Med J, 2(3), 01-07.
Abstract
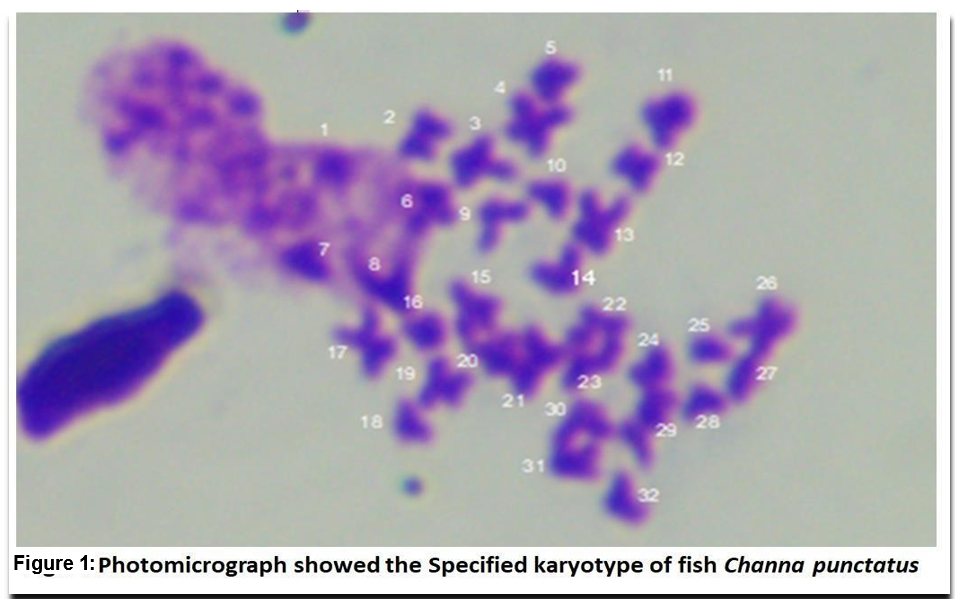
Karyotype analysis is the most classical method in the field of cytogenetics. It includes the determination of 2n chromosome numbers, relative length of chromosomes, centromeric formula, centromeric index, total chromatin length, etc. It describes a technique for taking pictures of a stained preparation in which the chromosomes are arranged in regular centromere patterns. In the present study, the significant karyotype of Channa punctatus were observed by analysing the metacentric, submetacentric, subtelocentric, and telocentric chromosomes. Among these the diploid chromosome number was 32 which were identified to be 14-metacentric, 12- submetacentric and 7- sub telocentric and 7- telocentric chromosomes. Thus, such studies shall be promoted to extend the analysis on the physiological health of fishes in changing environment.
Keywords
Karyotype, Cytogenetics, Centromere, Physiological Health.
Introduction
The species that make up the fish family are incredibly diverse. The fishes have chromosomal complements that vary greatly in both character and number, demonstrating their diversity. The topic of fish chromosomes might easily fill volumes, so this article's focus has been restricted to a broad overview of fish cytogenetics and how it can be used to further aquaculture and fish biotechnology. Current genetic research uses fish cytogenetics for a wide range of purposes, including studies into the evolutionary history of fishes, environmental toxicology, experimental ploidy manipulation, and transgenic fish production [1]. Cytogenetic techniques like taxonomic studies and karyological analyses are essential to providing fundamental knowledge on fish breeding programmes like inter-specific hybridization, chromosome manipulation techniques, and genetic improvement of commercial fish [2- 6]. To date, many karyotyping techniques have been developed to visualise fish chromosomes at different developmental phases, including tissue cultures, squashing techniques, and cell suspensions of the tissues going through mitosis [7,8].
Fish chromosome research dates back to study of the fish Salmo truttofario's chromosomes by Post's 1965. The progress of several fish cytogenetic research has been summarised by Blaxhall (1975) and Bloom (1977) [9,21]. 32 numbers of chromosomes were found in Channa punctatus in both sexes of normal diploid cells. We found no sex chromosomes in this fish species. These findings are consistent with the earlier data that is currently available [10-12]. Channel catfish (Ictalaris punctatus) standardised karyotypes were created by Zhang and Tiersch in (1998) [13]. Like with mammals, research on fish chromosomes began in the 1960s. Karyological investigations enabled us to gain a basic understanding of chromosome with their size, number, and morphology.
Numerous researchers have studied various species, such as crayfish. Phimphamet al., (2017) studied the five species of lutjanid fish. Yadav and Neeru (2018) studied constitutive heterochromatin and NOR banding in Three species of Indian major carps [14- 18]. In modern methods like karyo-taxonomy, chemotaxonomy, DNA barcoding, and DNA polymorphism aid in fish identification by adding new characters to the morphological character-based system [19].
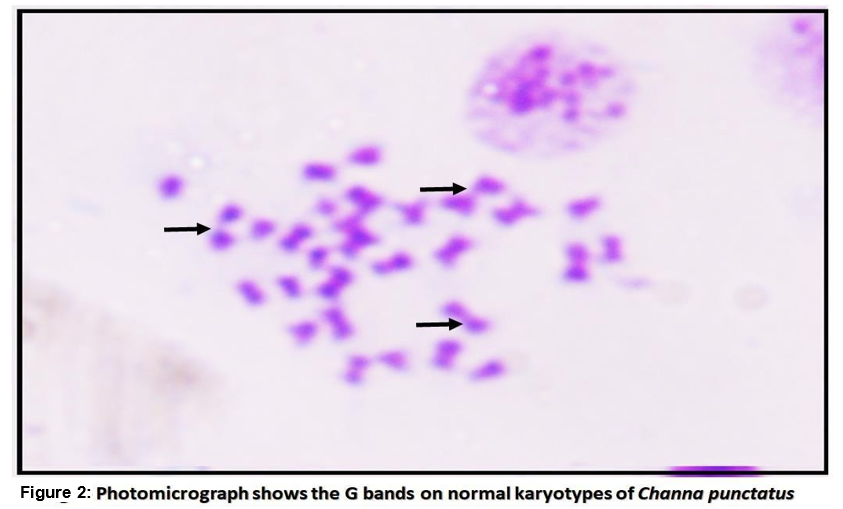
The purpose of the current study is to investigate the normal metaphase chromosomes spreads with G-banding which represents the heterochromatic region which is rich in AT (Adenine and Thymine)stains more darkly, whereas less condense region of euchromatin which is rich in GC (Guanine and Cytosine) appears as light bands. Although the precise mechanism of silver staining is unknown, it is thought to bind to the non-histone proteins of the transcriptionally active NOR region.
Material and methods
The experimental fishes (Channa punctatus (05-15 gms)) were used for chromosomal studies. which were collected Bithoor side local market Kanpur district. Fish that weren't sick were acclimated to the lab environment. We chose a total of 16 fishes with various body weights and sizes. After 4 hours of colchicine (0.1%) therapy, the abdomen area was dissected, the kidney tissues were collected, immediately transferred to different Petri dishes, chopped into small pieces, and then stored in a hypotonic solution for 30 minutes at room temperature. A pipette was then used to aspirate out the hypotonic solution. Each tube's contents were re-suspended after the addition of the fresh, refrigerated 5.0 ml of fixative (3:1 ration of methanol and glacial acetic acid). After standing for 10 minutes, the fixative was pipetted out. Three times of the aforementioned procedure were required until a mass of tissues and a supernatant fell at the bottom of the tube. After fixing, the button was moved to filter paper to dry before being suspended in 45% glacial acetic acid solution and dropped from 12" height above pre-warmed slides. Slides were air dried, given a 24-hour maturation period, and then stained with 4% Giemsa according to Klingerman and Bloom (1977). G banding pattern was carried out as described by Sumner (1990) and NOR banding methods according to Howell and Black (1980) [20-22].
Results
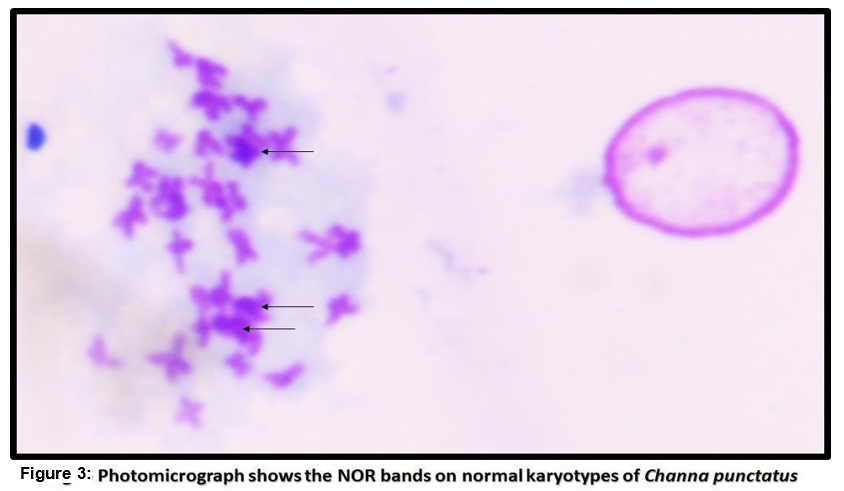
In the present study, fish were kept without feeding 1 day before the starting of the experiment [23]. The significant karyotype of fish Channa punctatus were observed by analyzing through karyotypic position like metacentric, submetacentric, subtelocentric, and telocentric chromosomes. Among these the diploid chromosome number was 32 (Fig. 1). In which 13metacentric, 7-submetacentric 5-subtelocentric and 7-telocentric were identified in experimental animal. The G bands' structural analysis revealed that after banding, transverse bands remained on them. These were brought on by heterochromatin's presence as well as the presence and absence of bases (Fig. 2). In the metaphase spreads, the stain showed up as a dark patch by using silver staining, the NOR bands on the chromosomes were found. In the metaphase spreads, these bands were seen in one to four locations. The transverse bands of the chromosomes were structurally evident after banding. Heterochromatin and the presence and lack of bases both contributed to the banding. The homologous chromosomes were paired using the bands. At two or three locations in the metaphase spread, the NOR regions were clearly visible. The transverse bands of the chromosomes are structurally evident after banding. Heterochromatin and the presence and lack of bases both contribute to the banding. The homologous chromosomes and the bands aid in chromosomal pairing. As can be seen in 2-3 locations in the metaphase spreads (Fig. 3), NOR binds to the nonhistone proteins of the transcriptionally active NOR region following the silver staining.
|
N o |
Remark |
Meta-centric |
Submetacentric |
Atelo-centric |
Telo-centric |
Total (2n) |
|
1 |
Karyotypic |
1,2,3,4,11,1 |
5,6,8,14,15, |
7,9,18,24,2 |
10,22,23,25, |
|
|
|
Specification |
2,13,16,17, |
21,30 |
9 |
27,28,32 |
|
|
|
|
19,20,26, |
|
|
|
|
|
|
|
31 |
|
|
|
|
|
|
|
|
|
|
|
32 |
|
2 |
Karyotypic |
|
|
|
|
|
|
|
Counts |
13 |
7 |
5 |
7 |
|
Table 1: Represents the Karyotypic Specification of fish Channa Punctatus
Conclusion and Discussion
Fish are particularly vulnerable to environmental dangers because they live in aquatic systems, which can harm them either directly or indirectly. Some pollutants build up in the fish's system, where they interfere at the molecular and cellular levels, causing aberrations in the genetic code that could be passed down to the next generation [24,25]. In the current investigation Channa punctatus was utilized as test material for karyotyping because of its G and NOR banding pattern.
The quantity of cells undergoing mitosis division closely correlates with the achievement of a desired number of mitotic chromosomes spread. Species and environmental factors also have an impact on the mitotic rate [26]. Shao et al. (2010) reported in their investigations by contrasting the number of metaphase chromosomes and mitotic rates in various life stages of Cynoglossussemilaevis, that the larvae had a greater mitotic rate than the juveniles and adults [27].
Similar to how older C. gariepinus and D. rerio larvae behaved differently to the colchicine concentration and duration and to the hypotonic solutions in our investigation, changes in mitotic division rates may be the cause. In a different study, the importance of environmental conditions in determining a trustworthy number of chromosome spreads was demonstrated by Wakahara (1972), who found that the ventral tail-fin epidermis of the larval African clawed frog (Xenopuslaevis) had a higher mitotic rate at night than it did during the day [28]. The most common microtubular toxin is colchicine. Colchicine has been shown to block spindle microtubules and distribute metaphase chromosomes in the cytoplasm in metaphase cells prior to nuclear envelope breakdown (NEB) [29].
In general, a high number of small-sized fish are difficult to work with when using the chromosomal spread approach. The majority of fish chromosome preparation methods involve the collection of tissues from various fish body parts, including the intestine, blood, skin fibroblasts, bone marrow, liver and spleen, fin epithelium, kidney, gills and gonads [30-39]. The kidney tissue of the fish Channa punctatus was used to prepare the chromosomal metaphase spreads in the present study. Alternate dark and bright spots running the length of the euchromatic region of chromosomes is known as the G banding technique. In 1990, Sumner [20,21]. In vertebrates, particularly lower vertebrates like fish, this method has been applied frequently. Balxhall (1983), Gold and Li [40,41]. In addition, Leitao et at. (1999 and 2000) described the G banding pattern in three species of oysters, Crassostreagigas, Crassostrea angulate, and Crassostreavirginica, and they noted the dark and light band patterns on the two arms of seven pairs of chromosomes. G banding is hypothesised to be the outcome of DNA and protein interaction and involves protease-treated chromosomes with giemsa dye [42,43]. Wendy (2001). Karhan and Ergen (2010 discovered a sizable heterochromatin area on the metacentric chromosome [44,45].
The initial silver staining techniques used to see AgNO3 proteins used two phases of AgNO3 impregnation followed by a stage of development using reducing agents such ammonia or formic acid [46]. A group of acidic non-histone proteins that bind silver ions and are specifically visible by silver techniques in routinely processed cytohistological material are found in NORs, a chromosomal tract that contains ribosomal genes 22]. For the study of chromosomal evolution in insects, Rufas et at. (1987), Bedo (1991), and Bella et at. (1993) reported that the NOR is a useful marker [47-49]. The quantity and position of NORs are typically indicative of a population's species within a certain insect species. In the current investigation, two pairs of medium-sized telocentric chromosomes were stained with silver to identify the NOR bands on experimental fish chromosomes. It is thought that silver staining interacts with the non-histonic proteins of the transcriptionally active NOR, which was discovered by Wiberg (1983) and Thodeet at. (1985) to be positioned terminally on the short arms of tiny submetacentric chromosomes in European eel Anguilla. In the Congo fish Conger, Sushana et at. (1994) discovered silver markings terminally on the short arm of an acrocentric chromosome [50-52]. These NORs were uniform in size across all specimens. The nuclear organising areas on the chromosomes of four species of the genus Diplodus were described as a phenotypic by Vitturiet at. in 1996 [53]. With the exception of one chromosome, which had clear bands situated right below the centromere, Morescalchiet at. (1998) applied the g banding patterns to the chromosomes of Macacasylvanus and noticed that all chromosomes appeared to have an identical banding pattern to most macaque species [54]. Sharma and Sharma (1998) also noted the use of G bands on numerous Indian fish species [55]. According to Wang et at. (2004), the quantities and locations of the NORs are frequently changeable and prevent correct localization on the crosstree chromosomes [56]. Review of 20-year history of karyotypes with NOR banding patterns in Bivales by Leitao and Chaves (2008) [57]. While Arzu and Ergene (2010) identified the NOR area were observed by many chromosome pairs by silver staining [45]. Valic et at. (2010) found the NORs in the telomeres of the two pairs of medium-sized submetacentric eleostesukliva chromosomes When Kaewsri et at. (2014) examined 74 sparrow fish karyotypes, they clearly saw the NOR in 6 pairs of acrocentric chromosomes. Nandini and Arokia (2014) discovered the NORs on one pair of submetacentric chromosomes in Labeorohita,. Neeruet at. (2018) investigated the Ag-NOR bands found on the homologous chromosomal pairs 15 and 11 in L. rohita and C. catla, respectively [18,58-60]. Ag-NOR staining revealed the presence of darkly stained NORs on the terminal area of C. mrigala. Nuclear organizing region were characterized in fresh water teleost heteropneustefossilis by Vermaet al. 2019 while Verma et al. (2020) identified the NOR bands with G bands in chromosomes of fish Clariasbatrachus [61,62]. Additionally, chromosomal analyses of economically significant fish are crucial for enhancing fish output genetically through chromosomal mapping and genetic selection. In fact, it is crucial to identify each chromosome individually and establish exact karyotypes. In order to increase fish production genetically through chromosomally set alterations and genetic selection, chromosomal research on commercially significant fish are crucial. This calls for the accurate karyotyping of each individual chromosome.
Acknowledgments
This paper was supported by letter No- 9/2022/447/ Seventy-4-2022(17)2022 Date 15/03/2022
UPCOE Grant, Uttar Pradesh Government Lucknow (U.P.) and letter no. CSJMU/R&D/CVR/14/2022under C.V. Raman Minor Research Project Scheme of CSJMU, Kanpur. Special thanks go to administration of Chhatrapati Sahu ji Maharaj University, Kanpur (U.P.). to provide me support during the experiments.
References
- Larry, C., Kent, H. B, & Penny K. R..(1990) Chromosomes of Fish. Adav. Veter. Sci. Comparative medicine Vol 34:209.
- Crego-Prieto, V., Danancher, D., Campo, D., Perez, J., Garcia- Vazquez, E., & Roca, A. (2013). Interspecific introgression and changes in population structure in a flatfish species complex after the Prestige accident. Marine pollution bulletin, 74(1), 42-49.
- Christopher, J. S. G., Murugesan, A. G., & Sukumaran, N. (2010). Induction of meiotic gynogenesis in the stinging catfish Heteropneustes fossilis (Bloch) and evidence for female homogamety. Aquaculture Research, 42(1), 129-138.
- Gilna, B., Kuzma, J., & Otts, S. S. (2014). Governance of genetic biocontrol technologies for invasive fish. Biological invasions, 16, 1299-1312.
- Thresher, R. E., Hayes, K., Bax, N. J., Teem, J., Benfey,T. J., & Gould, F. (2014). Genetic control of invasive fish: technological options and its role in integrated pest management. Biological invasions, 16, 1201-1216.
- Gui, J., & Zhu, Z. (2012). Molecular basis and genetic improvement of economically important traits in aquaculture animals. Chinese Science Bulletin, 57, 1751-1760.
- Armstrong, S. J., & Jones, G. H. (2003). Meiotic cytology and chromosome behaviour in wildâ?type Arabidopsis thaliana. Journal of experimental botany, 54(380), 1-10.
- Fan, Z., & Fox, D. P. (1990). A new method for fish chromosome preparation. Journal of fish biology, 37(4), 553- 561.
- Blaxhall, P. C. (1975). Fish chromosome techniques–a reviewof selected literature. Journal of Fish Biology, 7(3), 315-320.
- Rishi, K. K., & Haobam, M. S. (1990). A chromosomal study on four species of snake-heads (Ophiocephalidae: Pisces) with comments on their karyotypic evolution. Caryologia, 43(2), 163-167.
- Rishi, K. K., & Grewal, S. (1995). Chromosome aberration test for the insecticide, dichlorvos, on fish chromosomes. Mutation Research/Genetic Toxicology, 344(1-2), 1-4.
- Saxena, K.K. and Rana, R. (2008). Genotoxicity of permethrinin Channa punctatus by using chromosomal aberration test. Proceedings of 78th Annual Session and Symp. Nat. Acad. Sci., India. 41.
- Zhang, Q., & Tiersch, T. R. (1998). Standardization of the channel catfish karyotype with localization of constitutive heterochromatin and restriction enzyme banding. Transactions of the American Fisheries Society, 127(4), 551-559.
- Tan, X., Qin, J. G., Chen, B., Chen, L., & Li, X. (2004).Karyological analyses on redclaw crayfish Cherax quadricarinatus (Decapoda: Parastacidae). Aquaculture, 234(1-4), 65-76.
- Kavalco, K. F., Pazza, R., Bertollo, L. A., & Moreira-Filho,O. (2005). Molecular cytogenetics of Oligosarcus hepsetus (Teleostei, Characiformes) from two Brazilian locations. Genetica, 124, 85-91.
- Unal, S., GaffaroÄ?lu, M., Ayata, M. K., & Yüksel, E. (2014). Karyotype, C-banding and AgNORs of two endemic leuciscine fish, Pseudophoxinus crassus (Ladiges, 1960) andP. hittitorum Freyhof & Özulug, 2010 (Teleostei, Cyprinidae). Comparative Cytogenetics, 8(4), 249.
- Phimphan, S., Supiwong, W., Tanomtong, A., Pinthong, K., Sangpakdee, W., & Kaewsri, S. (2017). Karyotypic study of five Lutjanid species using conventional and Ag-NORs banding techniques. Cytology and Genetics, 51, 315-324.
- Neeru, N., Bhatnagar, A., & Yadav, A. S. (2018). A study of constitutive heterochromatin and NOR banding in three species of Indian major carps from the State of Haryana, India. Journal of Applied and Natural Science, 10(2), 535-539.
- Kesavan, S. (2021). Modern Taxonomic Tools in Fish Identification. AgriCos e-Newsletter Open Access Multidisciplinary Monthly Online Magazine: (2) 11:39-43.
- Sumner, A.T. (1990). Chromosome banding. Unwin HymanLtd.
- Kligerman, A. D., & Bloom, S. E. (1977). Rapid chromosome preparations from solid tissues of fishes. Journal of the Fisheries Board of Canada, 34(2), 266-269.
- Howell, W. T., & Black, D. A. (1980). Controlled silver- staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. experientia, 36(8), 1014-1015.
- OECD (2019) Test No. 203: fish, acute toxicity test, OECD guidelines for the testing of chemicals, section 2. OECD Publishing, Paris, pp 1–24.
- Verma, V. K., Gupta, N., Gupta, D. K., Krishna, G., & Shalaby,S. I. (2008). Clastogenic effects of lead on the chromosomal complements of Clarias batrachus. Bulletin of the National Research Centre, 33(5), 537-548.
- Verma, V. K., Gupta, N., Gupta, D. K., & Krishna, G. (2010). Chromosomal aberration in Heteropneustes fossilis due to lead intoxication. Bioresources for rural livelihood, 2, 247- 254.
- Shelton, W. L., Mims, S. D., Clark, J. A., Hiott, A. E., & Wang, C. (1997). A temperatureâ?dependent index of mitotic interval (τ0) for chromosome manipulation in paddlefish and shovelnose sturgeon. The Progressive Fishâ?Culturist, 59(3), 229-234.
- Shao, C. W., Wu, P. F., Wang, X. L., Tian, Y. S., & Chen, S. L.(2010). Comparison of chromosome preparation methods for the different developmental stages of the half-smooth tongue sole, Cynoglossus semilaevis. Micron, 41(1), 47-50.
- Wakahara, M. (1972). Daily variation in mitotic rate in tail- fin epidermis of larval Xenopus laevis and its modification by pineal organ-subcommissural organ system and photoperiods. Neuroendocrinology, 9(5), 267-277.
- Caperta, A. D., Delgado, M., Ressurreição, F., Meister, A., Jones, R. N., Viegas, W., & Houben, A. (2006). Colchicine- induced polyploidization depends on tubulin polymerization in c-metaphase cells. Protoplasma, 227, 147-153.
- Gold, J. R., Li, Y. C., Shipley, N. S., & Powers, P. K. (1990).Improved methods for working with fish chromosomes with a review of metaphase chromosome banding. Journal of Fish Biology, 37(4), 563-575.
- Jacobina, U. P., Cioffi, M. D. B., Souza, L. G. R., Calado,L. L., Tavares, M., Manzella, J., ... & Molina, W. F. (2011). Chromosome mapping of repetitive sequences in Rachycentron canadum (Perciformes: Rachycentridae): implications for karyotypic evolution and perspectives for biotechnological uses. BioMed Research International, 2011.
- Tantithakura, O., Anders, F., Anders, A., & Förster, W. (1993). High resolution technique: A new approach to study fish chromosomes. Biotrop Spec Publ, 52, 45-53.
- Manosroi, J., Petchjul, K., Mevatee, U., & Manosroi, A. (2003). Karyotype analysis of the hybrid, thai red Tilapia (Oreochromis niloticus Linn. X Oreochromis mossambicus Linn.).
- Affonso, P. R. A. D. M., & Galetti, P. M. (2005). Chromosomal diversification of reef fishes from genus Centropyge (Perciformes, Pomacanthidae). Genetica, 123, 227-233.
- Fontana, F., Lanfredi, M., Kirschbaum, F., Garrido-Ramos, M. A., Robles, F., Forlani, A., & Congiu, L. (2008). Comparison of karyotypes of Acipenser oxyrinchus and A. sturio by chromosome banding and fluorescent in situ hybridization. Genetica, 132, 281-286.
- Blanco, D. R., Lui, R. L., Bertollo, L. A. C., Diniz, D., & Filho, O. M. (2010). Characterization of invasive fish species in a river transposition region: evolutionary chromosome studies in the genus Hoplias (Characiformes, Erythrinidae). Reviews in Fish Biology and Fisheries, 20, 1-8.
- Poletto, A. B., Ferreira, I. A., Cabral-de-Mello, D. C., Nakajima, R. T., Mazzuchelli, J., Ribeiro, H. B., ... & Martins,C. (2010). Chromosome differentiation patterns during cichlid fish evolution. BMC genetics, 11(1), 1-12.
- Martinez, P. A., Jacobina, U. P., & Molina, W. F. (2011). Comparative cytogenetics and heterochromatic patterns in two species of the genus Acanthostracion (Ostraciidae: Tetraodontiformes). Marine genomics, 4(3), 215-220.
- Liu, S., Hui, T. H., Tan, S. L., & Hong, Y. (2012). Chromosome evolution and genome miniaturization in minifish. PLoS One, 7(5), e37305.
- JR, G., & YC, L. (1991). Trypsin G-banding of North American cyprinid chromosomes: phylogenetic considerations, implications for fish chromosome structure, and chromosomal polymorphism. Cytologia, 56(2), 199-208.
- Luo, C. (1998). Multiple chromosomal banding in grass carp,Ctenopharyngodon idellus. Heredity, 81(5), 481-485.
- Leitão, A., Chaves, R., Santos, S., Guedes-Pinto, H., & Boudry, P. (2007). Interspecific hybridization in oysters: restriction enzyme digestion chromosome banding confirms Crassostrea angulata× Crassostrea gigas F1 hybrids. Journal of Experimental Marine Biology and Ecology, 343(2), 253- 260.
- Leitao, A., Chaves, R., Santos, S., Boudry, P., Guedes Pinto, H., & Thiriot Quievreux, C. (2002). Cytogenetic study ofOstrea conchaphila (Mollusca: Bivalvia) and comparative karyological analysis within Ostreinae. Journal of Shellfish Research, 21(2), 685-690.
- Wendy, A. B. (2001). Karyotypes analysis and chromosome banding. Encyclopedia of Life Sciences, Nature Publishing Group:1-7
- Karahan, A., & Ergene2t, S. (2010). Cytogenetic analysis of Garra variabilis (Heckel, 1843)(Pisces, Cyprinidae) from Savur Stream (Mardin), Turkey. Turkish Journal of Fisheries and Aquatic Sciences, 10(4).
- Goodpasture, C., & Bloom, S. E. (1975). Visualization of nucleolar organizer regions in mammalian chromosomes using silver staining. Chromosoma, 53(1), 37-50.
- Rufas, J. S., Giménez-Abián, J., Suja, J. A., & Vega, C. G.D. L. (1987). Chromosome organization in meiosis revealed by light microscope analysis of silver-stained cores. Genome, 29(5), 706-712.
- Bedo, D. G. (1991). Cytological characterisation of heterochromatin in mitotic and meiotic chromosomes of the Old World screwworm fly, Chrysomya bezziana (Diptera: Calliphoridae). Genome, 34(4), 631-637.
- Bella, J. L., Serrano, L., Hewitt, G. M., & Gosálvez, J. (1993). Heterochromatin heterogeneity and rapid divergence of the sex chromosomes in Chorthippus parallelus and C. p. erythropus (Orthoptera). Genome, 36(3), 542-547.
- Wiberg, U. H. (1983). Sex determination in the European eel (Anguilla, L.) A hypothesis based on cytogenetic results, correlated with the findings of skewed sex ratios in eel culture ponds. Cytogenetic and Genome Research, 36(4), 589-598.
- Thode, G., Alvarez, M. C., Giles, V., & Garcia, E. (1985). Chromosome complement, C-banding and Ag-NOR location in Ophysurus serpens (Ophichthidae, Anguilliformes). Cytobios, 43(171), 73-77.
- Susana, S., Angelo, C., Angela, M. & Anna, M.P. (1994). Karyotypes, C- and G-banding and nuclear organizing regions Conger- conger (Osteichthyes, Anguilliformes). Bull. Zool., 16, 59-63.
- Vitturi, R., Libertini, A., Mazzola, A., Colomba, M. S., & Sara,G. (1996). Characterization of mitotic chromosomes of four species of the genus Diplodus: karyotypes and chromosomal nucleolar organizer region phenotypes. Journal of fish biology,49(6), 1128-1137.
- Morescalchi, A. M., Ciani, A. C., & Stanyon, R. (1998). Chromosome banding and molecular cytogenetics of the Barbary macaque, Macaca sylvanus. Italian Journal of Zoology, 65(1), 101-107.
- Sharma, A., & Sharma, M. S. (1994). Toxic effect of zinc smelter effluent to some developmental stages of freshwater fish, Cyprinus carpio(Linnaeus). Journal of environmental biology, 15(3), 221-229.
- Wang, Y., Xu, Z., & Guo, X. (2004). Differences in the rDNA- bearing chromosome divide the Asian-Pacific and Atlantic species of Crassostrea (Bivalvia, Mollusca). The Biological Bulletin, 206(1), 46-54.
- Leitão, A., & Chaves, R. (2008). Banding for chromosomal identification in bivalves: A 20-year history. Aquaculture, 1, 44-49.
- ValiÄ?, D., KapetanoviÄ?, D., Zanella, D., MrakovÄiÄ?, M., TeskeredžiÄ?, E., Besendorfer, V., ... & Ráb, P. (2010). The karyotype and NOR phenotype of Telestes ukliva (Cyprinidae). Folia Zoologica, 59(2), 169-173.
- Kaewsri, S., Tanomtong, A., Getlekha, N., Saenjundaeng, P., Suksuwan, R., & Supiwong, W. (2014). Standardized karyotype and idiogram of Quoy's parrotfish, Scarus quoyi (Perciformes: Scaridae) by conventional staining and Ag- NOR banding techniques. Cytologia, 79(4), 429-435.
- Nandini, S., & Arockia Rita, J. J. (2014). Karyomorphology of two major carps, Catla and Labeo rohita. Biores. Bull, 1, 1-7.
- Verma V.K., Gupta, N. & Gupta, D. K (2019). The characterization of somatic chromosome of fresh water catfish Heteropneustes fossils by nuclear organizing region, Proc. Zool. Soc. India, ISSN: 0973-6683, 18/ (2): 81-88.
- Verma V.K., Gupta, N., Gupta, D. K &Shalaby, S.I., (2020) Chromosomal metaphase spreads with G and NOR bandings of fresh water teleostsClariasbatrachus., Egypt.J. Vet. Sci., ISSN:1110-0222, 51(1): 61-67. DOI10.21608/ ejvs.2020.17868.1169.
- Post, A. (1965). Vergleichende, Untersuchungen der chromosomenzohenbeisuswasser. Z Zool. Syst. Evolution Torsch., 3, 47-93.