Research Article - (2025) Volume 4, Issue 1
In Silico Investigation of Mapk1 Mediated Key Protein Interactors in Lung Squamous Cell Carcinoma
2Department of Zoology, Rawalpindi Women University, Rawalpindi-46300, Pakistan
Received Date: Feb 13, 2025 / Accepted Date: Mar 17, 2025 / Published Date: Mar 24, 2025
Copyright: ©Ã?©2025 Fareeha Ambreen, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Shah, Z. A., Ambreen, F., Ejaz, R. (2025). In Silico Investigation of Mapk1 Mediated Key Protein Interactors in Lung Squamous Cell Carcinoma. J Surg Care, 4(1), 01-08.
Abstract
MAPK1 signaling pathway promotes development and survival of epithelial tissue. MAPKs family controls the activity of micro- environment by triggering the intracellular signaling. The focus of this study to understand the MAPK1 PPI by using STRING database and retrieve top twenty interaction network to analyze the significance of MAPK1 related proteins in lung Squamous cell carcinoma. We used cBioPortal a cancer platform to perform gene alteration analysis to uncover their participation in lung SqCC progression. We identified TP53 gene showed high ratio alteration among eleven lung SqCC linked disease causing MAPK1 mediated pathway genes in comparative study of lung squamous cell carcinoma TCGA, nature 2012 dataset. In pres- ent study by meta-analysis we characterized vital MAPK1 signaling pathway altered interactions that are MYLK, BCL2, TP53, BCL6, RAF1, CTTN, FGF19, FGF3, FGF4, ATR, TP63, HES1, CDKN2A, RASA, MRAS, TNFSF10, IL12A, TEK, PTEN, AGTR1, DVL3, EPHB1, TRIO, RPS6KA1, RPS6KA2, FOS, FGFR1, PAK2, PRKCI, PIK3CA, PIK3CG and KDR that involved in cell proliferation, growth, differentiation, tumor suppressed activities and apoptosis. In aberrant conditions they act as oncogenic and anti-apoptotic agents. The empirical validation of MAPK1 signaling cascade key interactions in lung SqCC in future that gives fruitful selection of therapeutic targets for squamous cell carcinoma.
Introduction
The most frequent cancer causing deaths related to lung cancer and elevated mortality ratio annually worldwide [1]. Lung cancer has two major groups i.e small-cell lung carcinoma (SCLC) and non- small-cell lung carcinoma (NSCLC). The NSCLC further divided into squamous cell carcinoma (SCC), adenocarcinoma (ADC) and large cell carcinoma (LCC) [2]. Lung cancer progression mainly initiated by bronchiolar and alvoliar epithelia by aberrent proliferation and differentiative capabilites [3]. The PIK3 amplification trigger the overexpression of EGFR in lung SCC. The TP53 gene deregulation mostly found in lung SCC. Cyclin D1 overexpression also reported in lung SCC progression [4]. In several studies KRAS, FBXW7, RUNX1T1, ERBB4, BA13, KEAP1, GRM8 and TP53 gene alterations are reported in lung SCC [5].
The high-throughput sequencing techniques generate huge amount of sequence secretes from DNA single molecule. The DNA sequences contained specific alterations, mutations, aberrations, methylation abnormalities, unregulated RNA splicing and editing observed that produce expression level change which leads to promotion of tumorigenesis [6]. The cells which contained altered genomic molecular frame undergo rapid proliferation and change into multiple clones [7]. Majority of cancer data containing databases presents genomic heterogeneity of diverse genes with unique oncogenic signaling pathways [8]. The progression of cancer initiated through abnormal gene expression which mediated by epigenetic activities in various studies [9]. The system level understanding of cancer gain the insight of aberrant processes that triggered by altered proteins of complex networks which play critical role in DNA methylation and gene expression [10].
The organism physiology at cellular level regulated by complicated interconnected networks of kinases containing multi layered messenger system that trigger the initiation and transduction of signals from receptors to targets [11]. The cell division, differentiation and maintenance like key activities regulated by serine-threonine containing mitogen-activated protein kinases [12]. The MAPK network maintains the microenvironment by regulating and controlling the input/output of system. The network cascade designs the fate of MAPK regulation, translocalization and target specificity [13]. The cascade components shared between various pathways such cross-talk through phosphorylation form a huge MAPK complex [14]. In this work we investigated top 20 protein link components of MAPK network in lung squamous cell carcinoma based on high ratio alterations in cross-cancer samples to identify drug targets of relevant connected pathways.
Bioinformatics Apparatus Environment
We construct Insilico protocol to discover co-targets of interacted proteins with MAPK1 in lung SqCC. We retrieve the MAPK1 PPIs from STRING database (www.string-db.org) that stored huge amount of interactions networks based on scoring pattern for greater reliability of interactions. We selected only top twenty interactions with more than 0.9 confidence score. We perform Gene Ontology enrichment analysis on retrieved network through WebGestalt toolkit (www.webgestalt.org) that explores the linking of network components with biological, molecular, localization, drugs and disease processes. We set the Hypergeometric test, Benjamin and Hochberg method and minimum two genes per category to achieve valuable enrichment results. For further meta-analysis to refine the GO results and find their contribution in lung SqCC promotion by cBioPortal (www.cbioportal.org) that perform mutation analysis of disease genes with cancer dataset. The cancer portal also gives the frequency and ratio of alterations. We infer the involvement of multiple functional processes genes with respective alterations that are the key targets in lung SqCC.
Results
Retrieval of MAPK1 Mediated Key Protein Interactions from STRING Database
We retrieve the MAPK1 mediated top 20 proteins interaction network from STRING database. The network contain 21 proteins with 48 interactions, PPI enrichment P-value is 1.52e- 05, average node degree is 4.57, clustering coefficient is 0.817 and high confidence interactions of 0.900. Network interactions showed by green (neighborhood evidence), blue (cooccurrence), red (fusion evidence), purple (experiment evidence), yellow (text mining), light blue (database) and black indicate the (coexpression evidence). The line thickness and sign indicates confidense score and action respectively (Figure. 1.1).
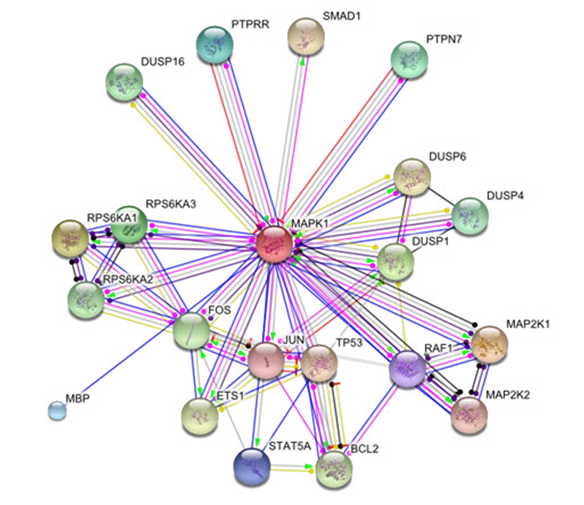
Figure 1.1: MAPK1 Mediated Key Protein Interactors Network Of 21 Nodes And 48 Edges
STRING database allows us to construct clusters based functional association by the method of K-MEANS algorithm. Each cluster has two types of interactions i.e. intra/interclusters which is the significanct view of functional drug targets. The MAPK1 is the largest cluster that interact with multiple key proteins of cellular processes including proliferation, growth, differentiation, migration and apoptosis (Figure 1.2)
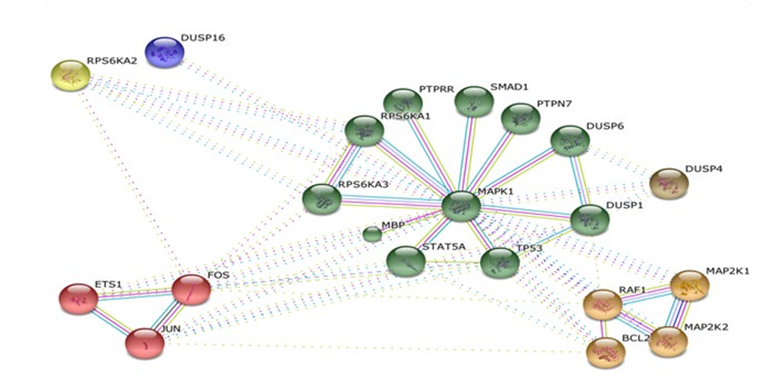
Figure 1.2: MAPK1 Network Clusters
MAPK1 Mediated Key Protein Interaction Go Analysis By Webgestalt
This result showed the vitality of network components in various cell processes that are essential in cell survival. The molecular functions of network proteins in cell microenvironment gain the imprortance at metabolic level. The localization of proteins gives their significance in signal transduction (Table 1).
|
S. NO |
GO Processes, Functions and Localization |
No. Genes |
|
1 |
TRIF-dependent toll-like receptor signaling pathway |
10 |
|
2 |
MYD88-independent toll-like receptor signaling pathway |
10 |
|
3 |
MYD88-dependent toll-like receptor signaling pathway |
10 |
|
4 |
Toll signaling pathway |
10 |
|
5 |
Positive regulation of innate immune response |
11 |
|
6 |
Phosphoric ester hydrolase activity |
7 |
|
7 |
MAP kinase phosphates activity |
4 |
|
8 |
Protein tyrosine phosphates activity |
7 |
|
9 |
Kinase bonding |
7 |
|
10 |
Phosphates activity |
7 |
|
11 |
Nucleoplasam |
13 |
|
12 |
Cytosol |
14 |
|
13 |
Nuclear part |
15 |
|
14 |
Nucleus |
19 |
|
15 |
Organelle part |
18 |
|
16 |
TF complex |
5 |
|
17 |
Intracellular organelle lumen |
14 |
Table 1: Biological, Molecular And Cell Localization Enrichment Analysis Of Mapk1 Network
This result explores the involvement of MAPK1 network components in other pathways in KEGG database. The linked pathways are involved in cell division, growth, migration, differentiation, transformation and neoplastic events (Table 2).
|
S. No |
KEGG Pathways |
No. Genes |
|
1 |
MAPK signaling pathway |
16 |
|
2 |
Neurotrop signaling pathway |
10 |
|
3 |
Pathways in cancer |
7 |
|
4 |
Long term potentiation |
7 |
|
5 |
Renal cell carcinoma |
6 |
|
6 |
Chronic myeloid leukemia |
6 |
|
7 |
B cell receptor signaling pathway |
6 |
|
8 |
ErbB signaling pathway |
6 |
|
9 |
Progesterone mediated oocyte maturation |
6 |
|
10 |
Colorectal cancer |
7 |
Table 2: The MAPK1 Network KEGG Pathway Enrichment Analysis
The result showed the number of drugs that target the components of the network (Table 3).
|
S. No |
Drugs |
No. Genes |
|
1 |
Collagenase |
4 |
|
2 |
Glutathione |
5 |
|
3 |
Masoprocol |
2 |
|
4 |
Cisplatin |
3 |
|
5 |
Fludarabine |
2 |
|
6 |
Sorafenib |
2 |
|
7 |
Boriezomib |
2 |
|
8 |
Genistein |
2 |
|
9 |
Alzeroxylon |
2 |
|
10 |
Cyclophosphamide |
2 |
Table 3: The Drugs Enricment Analysis of MAPK1 Protein Network
The most significant result of this study to identify the involvement of network components in diseases especially in cancer related events. Through these findings we perform further analysis to gain deep insight of such disease relevent proteins in lung cancer progression (Table 4).
|
S. No |
Diseases |
No. Genes |
|
1 |
Drug interaction with drug |
8 |
|
2 |
Noonan syndrome |
4 |
|
3 |
Cancer or viral infection |
9 |
|
4 |
Stress |
7 |
|
5 |
Death |
6 |
|
6 |
Drug toxicity |
5 |
|
7 |
Pulmonary valve stenoses |
3 |
|
8 |
Coffin-lowry syndrome |
5 |
|
9 |
Cell transformation, neoplastic |
5 |
|
10 |
Melanosis |
3 |
Table 4: The MAPK1 Pathway Disease Enrichment Analysis
MAPK1 Disease Causing Genes Comparative Charatriza- tion With Lung Squamous Cell Carcinoma Dataset
The Cancer Genome Atlas (TCGA) provides genetic alterations in 178 lung squamous cell cancers. The dataset contain 360 mutations in exonic regions, 165 rearrangements and 323 CNAs. MAPK1 mediated pathway 92.1% altered in dataset. we obtain 11 genes encoding proteins from cancer or viral infection, death and cell transformation neoplasam by disease enrichment analysis of Webgestalt (Figure 3.1a-b).
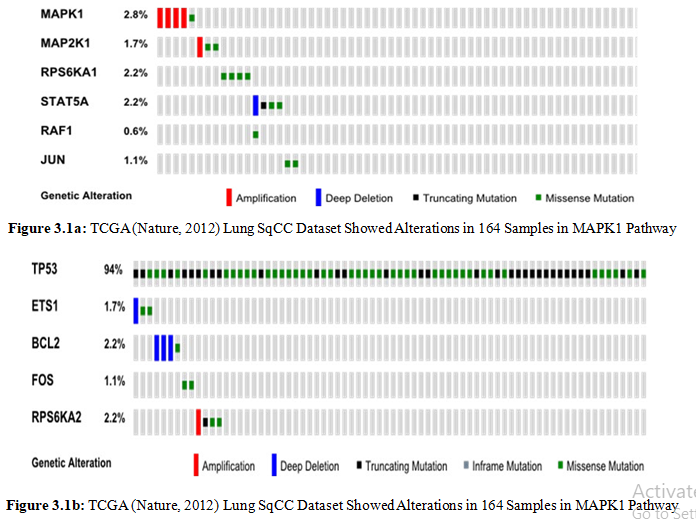
We further assess the altered PPI linkers to identify wide range of targets to control the aberrent cellular signaling. We use cbioportal network tab to generate state change reaction network of related components.
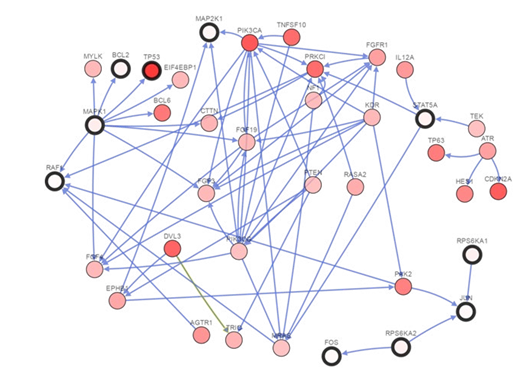
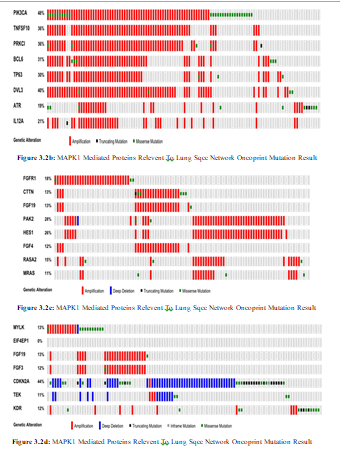
Figure 3.2a: MAPK1 Mediated Proteins Relevent To Lung Sqcc Network By Cbioportal That Involve In State Change Reactions

Discussion
MAPKs regulate cell division, maintainace and migration. The ERK1 to ERK8, p38-α to p38-δ and JUNK1 to JUNK3 are included in MAPKs family [15]. The signaling cascade of MAPKs initiates by three components that are MAPK, MAP2K and MAP3K which phosphorylate and regulate the MAPKs by MAP2Ks phosphorylation. Transcription factors like p53, c-Jun and ATF2 phosphorylated by MAPKs [16]. The MAPKs mediated network regulated by various linked kinases such as KSR, MP1 and JIPs [17]. The JNK, MKK7 and MLK members interact with JIP proteins and their expression increase the JNK regulation that promote type-2 diabetes and obesity [18]. The ERK/MAPK pathway gain high profile importance in cancer therapy. Human malignancies commonly has mutated Ras-Raf-ERK/MAPK pathway [19].
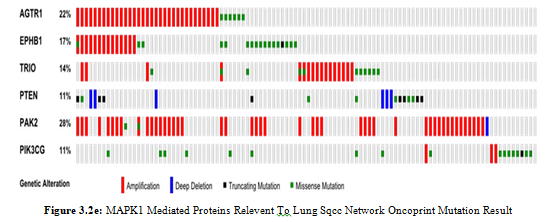
In present study MAPK1 showed 2.8% amplification which in- volved as a signaling integrating unit that regulate cellular prolif- eration and moleular target phosphorylation. We identified several MAPK1 interactors such as MYLK, BCL2, TP53, BCL6, RAF1, CTTN, FGF19, FGF3 and FGF4 that showed 8% amplification, 2.2% loss of function, 94% loss of function, 29% amplification, 0.6% loss of function, 11% amplification, 13% amplification, 11.8% amplification and 12% amplification respectively that are involved in regulation of cellular migration, regulation of apoptot- ic events, apoptotosis activaton, cellular morphogenesis, cell cycle controller, regulator of cytoskeleton and cell survival events. Inter- estingly all those proteins which are involved in cell proliferation are overexpressed that leads to tumriogenesis and such proteins that are controller of cell cycle and apoptotic causing agents un- dergo loss of function. We identified KDR which showed 8% am- plification that involved in endothlial morphogensis, survival and proliferation. It has several interactions such as FGFR1, PAK2, PRKCI, PIK3CA and PIK3CG which showed amplification of 17%, 27%, 35%, 43% and loss of function 11% respectively that involved in regulation of downstream signaling, reorganization of cytoskeleton, activation of microtubule, catalyst the regulatory ac- tivites and immune response.
In our study various unique differential expressions are observed such as ATR, TP63, HES1, CDKN2A, RASA, MRAS, TNFSF10, IL12A, TEK, PTEN, AGTR1, DVL3, EPHB1, TRIO, RPS6KA1, RPS6KA2and FOS that expressed 17% amplification, 29% amplification, 26% amplification, 44% loss of function, 13%
amplification, 10% amplification, 36% amplification, 20.5 amplification, 10% loss of function, 11% loss of function, 20% amplification, 38% amplification, 11% amplification, 12% amplification, 2.2% loss of function, 2.1% loss of function and 1.1% loss of function. These proteins are involved in kinases checkpoint phosphorylation, maintainance of stem cell, cell migration, TP53 stabilizer and checker of cell proliferation. The amplified proteins has overexpression in cell microenvirnoment that mostly trigger the cell cycle pathways which goes to rapid uncontrollable cell division and misssense mutated genes has underexpression of proteins that are the regulators of cell cycle arrest.
In this study we analyzed lung SqCC on the base of MAPK1 PPI network mutation and copy number alteration that explore the drug targets in therapeutic environment. The MAPK1 cluster is the main drug target to deactivate and inhibit its dynamics with other protein interactions which are key indicators of cell proliferation. We identified various cascade components that are reasonable therapeutic targets to overcome and control the carcinogenesis progression.
Conclusion
MAPK1 signaling pathway control various cellular processes such as growth, division, differentiation, survival and migration. The signaling network biological approach gain deep insight into disease invovled pathway components. For ideal understanding of disease the related protein interactions uncover the core mechnism that stimulate the cell proliferation and anti-apoptotic acitivty. In the present work MAPK1 mediated signaling pathway showed high level alteration frequency in apoptotic and tumor suppressor proteins that are unable to check and control the cell proliferation in aberrent conditions that leads to lung SqCC. These MYLK, BCL2, TP53, BCL6, RAF1, CTTN, FGF19, FGF3, FGF4, ATR,
TP63, HES1, CDKN2A, RASA, MRAS, TNFSF10, IL12A, TEK, PTEN, AGTR1, DVL3, EPHB1, TRIO, RPS6KA1, RPS6KA2, FOS, FGFR1, PAK2, PRKCI, PIK3CA, PIK3CG and KDR identified significant interactions gain deep therapeutic attention. We can desigen novel therapeutic techniques against these interactions which contribute in lung SqCC. This study in future provide therapeutic targets that are mediated by MAPK1 induced carcinogenesis
References
1. Parkin, M. (2004). Lung cancer epidemiology and etiology. Pathology and Genetics, Tumours of Lung, Pleura, Thymus and Heart, pp.12-15.
2. Travis, W. D., Brambilla, E., Muller-Hermelink, H. K., & Harris, C. C. (2004). World Health Organization classification of tumours. Pathology and genetics of tumours of the lung, pleura, thymus and heart, 10, 179-84.
3. Otto, W. R. (2002). Lung epithelial stem cells. The Journal of Pathology: A Journal of the Pathological Society of Great Britain and Ireland, 197(4), 527-535.
4. Hammar, S. P., Brambilla, C., Pugatch, B., Geisinger, K., Fernandez, E. A., Vogt, P., ... & Harris, C. C. (2004). Squamous cell carcinoma. World Health Organization Classification of Tumours, Pathology & Genetics, Tumours of the Lung, Pleura, Thymus and Heart, 26-34.
5. Kan, Z., Jaiswal, B. S., Stinson, J., Janakiraman, V., Bhatt, D., Stern, H. M., ... & Seshagiri, S. (2010). Diverse somatic mutation patterns and pathway alterations in human cancers. Nature, 466(7308), 869-873.
6. Yates, L. R., & Campbell, P. J. (2012). Evolution of the cancer genome. Nature Reviews Genetics, 13(11), 795-806.
7. Landau, D. A., Carter, S. L., Stojanov, P., McKenna, A.,
Stevenson, K., Lawrence, M. S., ... & Wu, C. J. (2013). Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell, 152(4), 714-726.
8. Ellis, M. J., & Perou, C. M. (2013). The genomic landscape of breast cancer as a therapeutic roadmap. Cancer discovery, 3(1), 27-34.
9. Easwaran, H., & Baylin, S. B. (2013). Epigenetic abnormalities in cancer find a “home on the range”. Cancer Cell, 23(1), 1-3.
10. Peifer, M., Fernández-Cuesta, L., Sos, M. L., George, J., Seidel, D., Kasper, L. H., ... & Thomas, R. K. (2012). Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nature genetics, 44(10), 1104-1110.
11. Lewis, T. S., Shapiro, P. S., & Ahn, N. G. (1998). Signal transduction through MAP kinase cascades. Advances in cancer research, 74, 49-139.
12. Dhillon, A. S., Hagan, S., Rath, O., & Kolch, W. (2007). MAP kinase signalling pathways in cancer. Oncogene, 26(22), 3279-3290.
13. Yasuda, J., Whitmarsh, A. J., Cavanagh, J., Sharma, M., & Davis, R. J. (1999). The JIP group of mitogen-activated protein kinase scaffold proteins. Molecular and cellular biology.
14. Frost, J. A., Steen, H., Shapiro, P., Lewis, T., Ahn, N., Shaw, P. E., & Cobb, M. H. (1997). Crossâ?cascade activation of ERKs and ternary complex factors by Rho family proteins. The EMBO journal.
15. Schaeffer, H. J., & Weber, M. J. (1999). Mitogen-activated protein kinases: specific messages from ubiquitous messengers. Molecular and cellular biology, 19(4), 2435- 2444.
16. Morrison, D. K., & Davis, R. J. (2003). Regulation of MAP kinase signaling modules by scaffold proteins in mammals. Annual review of cell and developmental biology, 19(1), 91- 118.
17. Whitmarsh, A.J., 2006. The JIP family of MAPK scaffold proteins.Biochemical Society Transactions, 34(5), pp.828- 832.
18. Davis, R. J. (2000). Signal transduction by the JNK group of MAP kinases. Cell, 103(2), 239-252.
19. Roberts, P. J., & Der, C. J. (2007). Targeting the Raf-MEK- ERK mitogen-activated protein kinase cascade for the treatment of cancer. Oncogene, 26(22), 3291-3310.