Research Article - (2026) Volume 9, Issue 1
Identification of Cassava Mosaic Virus Strains in Zimbabwe Using Nanopore Sequencing
2University of Zimbabwe, Zimbabwe
Received Date: Feb 02, 2026 / Accepted Date: Feb 27, 2026 / Published Date: Mar 09, 2026
Copyright: ©2026 Tapiwa Nyakauru, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Nyakauru, T., Robertson, F. (2026). Identification of Cassava Mosaic Virus Strains in Zimbabwe Using Nanopore Sequencing. J Agri Horti Res, 9(1), 01-07.
Abstract
Cassava (Manihot esculenta Crantz: Euphorbiaceae) is the only species in its genus that is grown as a food crop. Africa is the largest centre for cassava production, producing over 100 million tonnes per year. However, cassava is affected by several diseases including Cassava Mosaic Disease (CMD) which is caused by Cassava Mosaic Viruses (CMVs). Symptoms of CMD include distortion of leaf lamina, mottling, unordered growth and malformation of the leaves, formation of chlorotic mosaics, and narrowing of the leaves. Several strains of CMVs that have been recorded include Africa cassava mosaic virus (ACMV), East African cassava mosaic virus (EACMV), East African cassava mosaic Malawi virus (EACMMV), East African cassava mosaic Ugandan virus (EACMV-UG), East African cassava mosaic Cameroon virus (EACMCV), East African cassava mosaic Zanzibar virus (EACMZV), East African cassava mosaic Kenya virus (EACMKV) and South African cassava mosaic virus (SACMV). However, limited information is available with regards to CMV strains in Zimbabwe. This work focused on identifying specific strains of CMV that are affecting cassava plants in Zimbabwe. Using nanopore sequencing, several strains of CMV including ACMV, EACMV, EACMV-UG, EACMV-K, and SACMV were detected in cassava plants from Zimbabwe. These findings provide the first comprehensive evidence of CMV strain diversity in Zimbabwe and highlight the potential of nanopore sequencing as a rapid and cost-effective tool for virus surveillance, early detection, and management of CMD in cassava production systems.
Keywords
Cassava Mosaic Virus, Cassava Mosaic Disease, Nanopore Sequencing, Cassava, Bioinformatics
Introduction
Cassava (Manihot esculenta Crantz) is the sole species within the genus Manihot cultivated extensively for human consumption. Globally, it serves as a major source of dietary carbohydrates, particularly in tropical and subtropical regions. Africa is the leading centre of cassava production, with annual outputs exceeding 100 million metric tons, and more than 70 million people on the continent rely on cassava as a staple food source [1,2]. While the production of cassava is set to increase as its tubers can be used for ethanol production and livestock feed, cassava is affected by several diseases, including Cassava Mosaic Disease (CMD) caused by Cassava Mosaic Viruses (CMVs) [3,4]. CMVs belong to the family Geminiviridae of the genus Begomovirus [5]. While symptoms of CMD vary by season and cassava variety, general symptoms of CMDs include distortion of the leaf lamina, mottling, unordered growth and malformation of the leaves, formation of chlorotic mosaics, and narrowing of the leaves, which all results in reduction in photosynthesis [6,7].
CMVs strains recorded in Africa include African cassava mosaic virus (ACMV), East African cassava mosaic virus (EACMV), East African cassava mosaic Malawi virus (EACMMV), East African cassava mosaic Cameroon virus (EACMCV), East African cassava mosaic Zanzibar virus (EACMZV), East African cassava mosaic Kenya virus (EACMKV), and South African cassava mosaic virus (SACMV) [8-10,5]. Rapid, accurate, and sensitive detection of CMV is essential for effective control and management of CMD. Early identification of CMV infections, prior to the appearance of visible symptoms, allows for timely implementation of protective measures, thereby reducing the risk of virus transmission to healthy cassava plants [11]. The detection of CMVs is performed using different methods, including enzyme-linked immunosorbent assay (ELISA) and polymerase chain reaction (PCR) [12,13]. In Zimbabwe, the presence of CMVs in cassava plants grown in the northern region was identified using novel degenerate primers capable of amplifying ACMV, EACMV, and SACMV [14]. With the emergence of new CMV strains, ELISA has become less effective for detecting these diverse variants. Although PCR is still widely used in viral detection, in some circumstances, it is incompatible in detecting new strains of the virus due to viral mutation or absence of sequences for primer design [15].
Oxford Nanopore MinION sequencing has gained much attention in identifying plant viruses because of its capability to generate whole genome sequences which enables the identification of known and novel viral strains [16,11]. With the increase in cassava production in Zimbabwe, there is a need to investigate the prevalence and presence of CMV strains, hence, the objective of the project was to detect strain/s of CMV affecting cassava plants in Chiredzi, southeast of Zimbabwe, using the latest MinION nanopore sequencing technology.In this study, Oxford Nanopore sequencing was employed to screen cassava plants for the presence of CMVs. Multiple strains were identified in samples from Zimbabwe, including ACMV, EACMV, EACMV- UG, EACMV-K, and SACMV, demonstrating the capability of oxford nanopore to be used as a rapid diagnostic tool for early virus detection in plants. These findings provide important insights into the diversity and distribution of CMVs in Zimbabwe and contribute to the development of effective disease management strategies.
Methodology
Sample Preparation and DNA Extraction
Leaf samples exhibiting cassava mosaic disease symptoms were collected from a field in Chiredzi, Zimbabwe. Samples were flash- frozen in liquid nitrogen and stored at −80 °C until DNA extraction. Total genomic DNA (gDNA) was extracted following the cetyl trimethylammonium bromide (CTAB) protocol described by Doyle and Doyle (1990). Briefly, 300 mg of leaf tissue was rinsed with distilled water, blotted dry with a sterile paper towel, and surface-sterilized with 70% ethanol. The tissue was then ground to a fine powder in liquid nitrogen using a sterile mortar and pestle and transferred to 1.5 mL microcentrifuge tubes containing 400 μL of preheated (65 °C) CTAB extraction buffer [2% (w/v) CTAB, 1% (w/v) polyvinylpyrrolidone (PVP), 100 mM Tris-HCl (pH 8.0), 1.4 M NaCl, 20 mM EDTA, and 5% (v/v) β-mercaptoethanol]. Samples were mixed by gentle inversion and incubated at 65 °C for 20 minutes, with mixing every 5 minutes.
Following incubation, 400 μL of chloroform:isoamyl alcohol (24:1, v/v) was added, and tubes were shaken horizontally on a rotary shaker (200 rpm) at room temperature for 15 minutes. The mixtures were centrifuged at 10,000 rpm for 5 minutes, and the aqueous phase was transferred to fresh tubes. DNA was precipitated by adding 400 μL of isopropanol, mixing thoroughly, and incubating at −20 °C for 1 h, followed by centrifugation at 10,000 rpm for 5 minutes. The supernatant was discarded, and the DNA pellet was washed with 400 μL of cold 100% ethanol, centrifuged for 5 minutes at 10,000 rpm, and air-dried. DNA was resuspended in 100 μL of nuclease-free deionized water and further purified using a Genomic DNA Clean & Concentrator kit (D4010, Zymo Research). DNA concentration and purity were determined using a NanoDrop spectrophotometer, and DNA integrity was confirmed by electrophoresis on a 1% agarose gel stained with ethidium bromide and visualized using an E-Gel Imager (Life Technologies, Israel).
Whole Genome Sequencing and Bioinformatics Analysis
DNA sequencing libraries were prepared using the Rapid Barcoding Kit (SQK-RBK004) with R9.4.1 flow cells (Oxford Nanopore Technologies), following the manufacturer’s instructions. Libraries were loaded onto a MinION device connected to a MinIT for real-time basecalling. Raw fastq sequences, generated via the ONT Albacore pipeline, were demultiplexed using Porechop (https://github.com/rrwick/Porechop) and was deposited in SRA NCBI database under accession PRJNA1308336 under biosample accessions SAMN50694092-3. Sequences were filtered using NanoFilt 2.8.0 and BLASTn searches against a custom CMV database were performed to identify CMV reads, which were assembled de novo in Geneious Prime v11.0.5 to produce viral contigs which were deposited in SRA NCBI database under accession SAMN50736739. The novelty of CMV contigs was assessed using BLASTn searches against the NCBI database. Phylogenetic relationships of CMV sequences were inferred using the Maximum Likelihood method in MEGA-X, incorporating published CMV reference genomes.
Results
Genomic DNA was successfully extracted, and DNA yield and purity were evaluated with a NanoDrop spectrophotometer. Whole- genome sequencing was performed on the Oxford Nanopore MinION platform. Basecalled reads were demultiplexed using Porechop, yielding a total of 109574 raw reads across all samples (Figure 1).
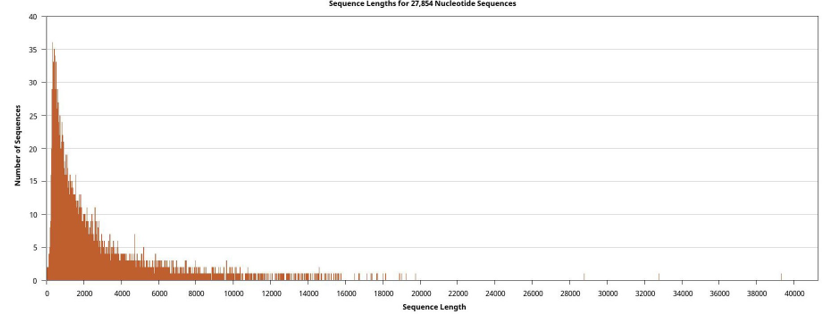
Figure 1: Cumulative Density Curves Showing Sequenced Reads and their Lengths
For sample 96/0037, sequencing generated a mean read length of 5604 bp, with the longest read spanning 113,791 bp. For sample XM6, a comparable mean read length of 3718 bp was observed, with the maximum read length also reaching 39,327 bp. Sequences were deposited in SRA NCBI database under accession PRJNA1308336.
Nanopore Sequencing and Bioinformatics Analysis
BLAST analysis was performed against a custom CMV database under default parameters and the number of CMV-assigned reads is presented in Table 1.
|
Sample name |
CMD severity score |
Total reads |
Max. seq length |
CMV-A BLAST hits |
Number of contigs formed |
|
96/0037 |
2 |
81720 |
113791 |
911 |
47 |
|
XM6 |
5 |
27845 |
39327 |
4571 |
215 |
Table 1: Nanopore Sequencing Results and Custom BLAST Analysis Obtained after Whole Genome Sequencing of Cassava Plants Affected with CMV
De novo assembly, defined as the reconstruction of genomes in the absence of a reference sequence, was subsequently performed on reads with high similarity to CMV, as identified by the custom BLAST search. BLAST analysis of the resulting contigs showed strong sequence homology with previously published CMVs documented in other studies (Table 2). Cassava variety 96_0037 was found to be infected with three strains: SACMV, EACMV-Ke, and EACMV-Malawi. Variety XM6 was infected with SACMV, EACMV, and EACMV variants from Kenya, Malawi, and Uganda. Furthermore, phylogenetic analysis confirmed the close relationship between known CMV sequences and novel contigs (Figure 2).
|
Source of novel contigs |
Description |
Query Coverage |
Pairwise Identity |
Accession |
|
XM6 |
South African cassava mosaic virus |
75.00-100.00% |
78.00-85.60% |
OP971522.1, KJ887649, AJ575560 |
|
East African cassava mosaic virus-Kenya |
76- 93.00% |
79.20-85.36% |
JF909222.1, JF909092.1, JF909174, JF909096, JF909188, JF909069, JF909069, JF909157, JF909163 |
|
|
East African cassava mosaic virus-Malawi |
100.00% |
76.60-84.00% |
KP890350.1, KT869119, KY885005, KY885004, KT869121, NC_022645, MT821892.1 |
|
|
East African cassava mosaic virus |
68.89-72.75% |
77.70-81.5% |
AJ717558, AJ717554, MZ494486 |
|
|
East African cassava mosaic virus-Uganda |
93.32-100.00% |
80.30-82.5% |
FN668380, AM502332 |
|
|
96_0037 |
East African cassava mosaic Malawi virus |
99.81-100.00% |
81.70-86.10% |
MT821892 |
|
South African cassava mosaic virus |
96.04-100% |
83.00-85.90% |
OP971522 |
|
|
East African cassava mosaic virus-Kenya |
100.00% |
78.10- 82.60% |
JF909069, JF909177 |
Table 2: BLAST Analysis of Novel Contigs Obtained from De Novo Assembly of Cmv Reads Obtained through Nanopore Sequencing
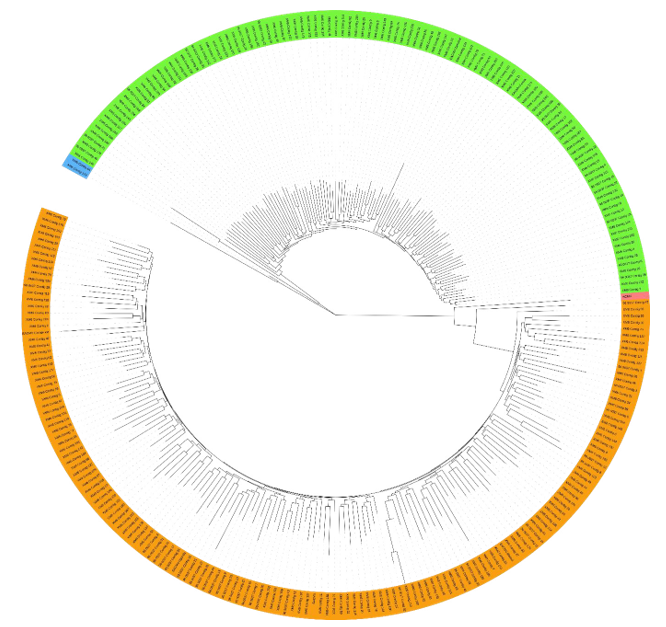
Figure 2: A Maximum Likelihood Phylogenetic Tree Showing the Evolutionary Relationships Between Novel Contigs and Known CMV Sequences
Discussion
Cassava is a critical food security crop in sub-Saharan Africa, providing a staple source of carbohydrates for millions of people. However, cassava production is severely threatened by cassava mosaic viruses which are responsible for cassava mosaic disease (CMD) [6]. CMD is one of the most devastating viral diseases of cassava, causing significant yield losses and threatening both food and income security [17]. The accurate and timely identification of CMV strains is therefore essential for implementing effective disease management strategies [18]. Historically, the detection of CMV has relied on methods such as polymerase chain reaction (PCR) and enzyme-linked immunosorbent assay (ELISA), which have been widely applied in cassava research [19,20]. However, these techniques are often limited by their dependency on prior knowledge of viral sequences, relatively long turnaround times, and the inability to detect novel or recombinant strains [21]. In many farming communities, diagnosis is still largely based on visual assessment of symptoms on cassava leaves, which can be misleading due to symptom similarity among different viral strains and environmental stressors [6].
In Zimbabwe, the information available on CMV strains is particularly outdated, with the most recent study dating back to 2004, which reported the occurrence of South African cassava mosaic virus (SACMV) in the country [6,22]. Since then, farmers have continued to report CMD outbreaks, with some CMV strains exhibiting higher virulence and adaptability, showing the urgent need for updated molecular surveillance of CMVs circulating in the country. The identification of CMV strains currently affecting cassava in Zimbabwe is crucial for designing targeted management strategies to reduce yield losses and curb viral spread.
This study applied nanopore sequencing, one of the latest next- generation sequencing platforms, to characterize cassava viruses in Zimbabwe. Nanopore sequencing offers distinct advantages over traditional detection and sequencing methods, including rapid, real-time data generation, portability, relatively low cost, and the ability to generate long reads that improve genome assembly [11]. Importantly, nanopore sequencing can be deployed directly in the field, enabling virus detection and preliminary analysis within hours, compared to days or weeks required for conventional methods [16]. For a country like Zimbabwe, where cassava cultivation is expanding and resources for plant health monitoring are often limited, the ability to identify viral pathogens within hours represents a transformative step toward proactive disease management. Our bioinformatic analyses, including BLAST searches and phylogenetic tree construction, revealed that cassava variety 96_0037 was infected with three strains: SACMV, EACMV-Ke, and EACMV-Malawi, and variety XM6 was infected with SACMV, EACMV, and EACMV variants from Kenya, Malawi, and Uganda. Such mixed infections have been associated with more severe disease symptoms and increased potential for recombination, thereby accelerating the emergence of novel, more virulent variants [24].
Interestingly, several viral contigs from different plants were found to be highly similar to known CMV strains, suggesting that novel variants may be emerging due to point mutations or recombination events, a phenomenon previously documented in plant viruses [25-27]. Even though nanopore sequencing technology is known to have high error rates, the detection of such potential new strains often highlights the evolutionary dynamics of CMV populations in Zimbabwe and reinforces the need for continuous genomic surveillance. This study is particularly important for Zimbabwe as it provides the first updated molecular data on CMV strains in nearly two decades. The findings will inform cassava breeding programs by identifying the most prevalent and virulent strains to target in the development of resistant varieties. Furthermore, real-time diagnostic capacity using nanopore sequencing can enable agricultural extension services to make rapid, evidence- based decisions, reducing the spread of infection and safeguarding farmers’ livelihoods. Importantly, the viral contigs published in this study on NCBI database can serve as a valuable reference resource for future research aimed at monitoring, comparing, and identifying emerging CMV strains in Zimbabwe and other regions of Southern Africa. This data can be used to track viral evolution, improve regional surveillance systems, and guide integrated disease management strategies [28]. In the long term, the application of such sequencing-based surveillance systems can strengthen Zimbabwe’s preparedness against viral threats not only in cassava but also in other staple crops.
Conclusion
Nanopore sequencing offers a rapid, reliable, and field-deployable tool for cassava virus detection in Zimbabwe. Several strains of cassava mosaic virus were detected, including ACMV, EACMV, EACMV-UG, EACMV-K, and SACMV. By generating updated genomic data, this study provides essential insights that will benefit cassava farmers, breeders, and policymakers in Zimbabwe and the broader Southern African region. The potential of new strains of CMV in Zimbabwe highlights the need to further investigate the presence of new strains in Zimbabwe.
Data Availability Statement
The genomic data generated and analysed during the current study are available in the NCBI SRA repository under accession PRJNA1308336 (https://www.ncbi.nlm.nih.gov/bioproject/PRJ- NA1308336/) The raw sequencing data generated and analysed in this study are publicly available in the NCBI Sequence Read Ar- chive (SRA) under BioProject PRJNA1308336. The datasets can be accessed through the following SRA Experiment accessions:
SRX30193202
SRX30161564
SRX30161563
Authors’ Contribution: Tapiwa Nyakauru: Conceptualization, formal analysis, investigation, methodology, visualization, writing original draft, review and editing. Fiona Robertson: Conceptual- ization, investigation, methodology, supervision, visualization, re- viewing draft and editing.
Funding: This work was funded by the Alliance for Global Health of the Centre for Emerging and Neglected Diseases, USA project managed by the University of Zimbabwe, College of Health Sciences, Research Support Centre.
Conflict of Interest: The authors declare that they have no known conflict of interest
Ethics Declaration: Not applicable.
Consent to Publish Declaration: Not applicable
References
- Adebayo, W. G. (2023). Cassava production in africa: A panelanalysis of the drivers and trends. Heliyon, 9(9), e19939.
- Mohidin, S. R. N. S. P., Moshawih, S., Hermansyah, A., Asmuni, M. I., Shafqat, N., & Ming, L. C. (2023). Cassava ( Manihot esculenta Crantz): A Systematic Review for the Pharmacological Activities, Traditional Uses, Nutritional Values, and Phytochemistry. Journal of Evidence-Based Integrative Medicine, 28.
- Borku, A. W. (2025). Cassava (Manihot esculenta Crantz): its nutritional composition insights for future research and development in Ethiopia. Discover Sustainability, 6(1), 404.
- Fathima, A. A., Sanitha, M., Tripathi, L., & Muiruri, S. (2023). Cassava ( Manihot esculenta ) dual use for food and bioenergy: A review. Food and Energy Security, 12(1).
- Dye, A. E., Muga, B., Mwangi, J., Hoyer, J. S., Ly, V., Rosado, Y., Sharpee, W., Mware, B., Wambugu, M., Labadie, P., Deppong, D., Jackai, L., Jacobson, A., Kennedy, G., Ateka, E., Duffy, S., Hanley-Bowdoin, L., Carbone, I., & Ascencio-Ibáñez, J. T. (2023). Cassava begomovirus species diversity changes during plant vegetative cycles. Frontiers in Microbiology, 14.
- Chikoti, P. C., Mulenga, R. M., Tembo, M., & Sseruwagi,P. (2019). Cassava mosaic disease: a review of a threat to cassava production in Zambia. Journal of Plant Pathology, 101(3), 467–477.
- Hillocks, R. J., & Thresh, J. M. (2000). Cassava Mosaic and Cassava Brown Streak Virus Diseases in Africa: A Comparative Guide to Symptoms and Etiologies. 7(December), 1–8.
- Ndunguru, J., Legg, J., Aveling, T., Thompson, G., & Fauquet,C. (2005). Molecular biodiversity of cassava begomoviruses in Tanzania: evolution of cassava geminiviruses in Africa and evidence for East Africa being a center of diversity of cassava geminiviruses. Virology Journal, 2(1), 21.
- PATIL, B. L., & FAUQUET, C. M. (2009). Cassava mosaic geminiviruses: actual knowledge and perspectives. Molecular Plant Pathology, 10(5), 685–701.
- Zinga, I., Chiroleu, F., Legg, J., Lefeuvre, P., Komba, E. K., Semballa, S., Yandia, S. P., Mandakombo, N. B., Reynaud, B., & Lett, J.-M. (2013). Epidemiological assessment of cassava mosaic disease in Central African Republic reveals the importance of mixed viral infection and poor health of plant cuttings. Crop Protection, 44, 6–12.
- Boykin, Sseruwagi, Alicai, Ateka, Mohammed, Stanton,Kayuki, Mark, Fute, Erasto, Bachwenkizi, Muga, Mumo, Mwangi, Abidrabo, Okao-Okuja, Omuut, Akol, Apio, … Ndunguru. (2019). Tree Lab: Portable genomics for Early Detection of Plant Viruses and Pests in Sub-Saharan Africa. Genes, 10(9), 632.
- Ogbe, F. O., Atiri, G. I., Dixon, A. G. O., & Thottappilly,G. (2003). Symptom severity of cassava mosaic disease in relation to concentration of African cassava mosaic virus in different cassava genotypes. Plant Pathology, 52(1), 84–91.
- Legg, J. P., & Fauquet, C. M. (2004). Cassava mosaic geminiviruses in Africa. Plant Molecular Biology, 56(4), 585–599.
- Berry, S., & Rey, M. E. C. (2001). Molecular evidence for diverse populations of cassava-infecting begomoviruses in southern Africa. Archives of Virology, 146(9), 1795–1802.
- Sanjuán, R., & Domingo-Calap, P. (2016). Mechanisms of viral mutation. Cellular and Molecular Life Sciences, 73(23), 4433–4448.
- Sun, K., Liu, Y., Zhou, X., Yin, C., Zhang, P., Yang, Q., Mao, L., Shentu, X., & Yu, X. (2022). Nanopore sequencing technology and its application in plant virus diagnostics. Frontiers in Microbiology, 13.
- Ristaino, J. B., Anderson, P. K., Bebber, D. P., Brauman, K. A., Cunniffe, N. J., Fedoroff, N. V., Finegold, C., Garrett, K. A., Gilligan, C. A., Jones, C. M., Martin, M. D., MacDonald,G. K., Neenan, P., Records, A., Schmale, D. G., Tateosian, L., & Wei, Q. (2021). The persistent threat of emerging plant disease pandemics to global food security. Proceedings of the National Academy of Sciences, 118(23).
- Alejandro, J. A. G., Mausisa, J. H. M., & Paglinawan, C.C. (2025). Deep Learning Approach to Cassava Disease Detection Using EfficientNetB0 and Image Augmentation. 2024 IEEE 6th Eurasia Conference on IoT, Communication and Engineering, 28.
- Houngue, J. A., Zandjanakou-Tachin, M., Ngalle, H. B., Pita, J. S., Cacaï, G. H. T., Ngatat, S. E., Bell, J. M., & Ahanhanzo, C. (2019). Evaluation of resistance to cassava mosaic disease in selected African cassava cultivars using combined molecular and greenhouse grafting tools. Physiological and Molecular Plant Pathology, 105, 47–53.
- Jimenez, J., Leiva, A. M., Olaya, C., Acosta-Trujillo, D., & Cuellar, W. J. (2021). An optimized nucleic acid isolation protocol for virus diagnostics in cassava (Manihot esculenta Crantz.). MethodsX, 8, 101496.
- Hareesh, P. S., Resmi, T. R., Sheela, M. N., & Makeshkumar,T. (2023). Cassava mosaic disease in South and Southeast Asia: current status and prospects. Frontiers in Sustainable Food Systems, 7.
- Sseruwagi, P., Sserubombwe, W. S., Legg, J. P., Ndunguru, J., & Thresh, J. M. (2004). Methods of surveying the incidence and severity of cassava mosaic disease and whitefly vector populations on cassava in Africa: a review. Virus Research, 100(1), 129–142.
- Zheng, P., Zhou, C., Ding, Y., Liu, B., Lu, L., Zhu, F., & Duan, S. (2023). Nanopore sequencing technology and its applications. MedComm, 4(4).
- Fondong, V. N. (2017). The Search for Resistance to Cassava Mosaic Geminiviruses: How Much We Have Accomplished, and What Lies Ahead. Frontiers in Plant Science, 8.
- Duffy, S., & Seah, Y. M. (2010). 98% identical, 100% wrong: per cent nucleotide identity can lead plant virus epidemiology astray. Philosophical Transactions of the Royal Society B: Biological Sciences, 365(1548), 1891–1897.
- Ng, T. F. F., Duffy, S., Polston, J. E., Bixby, E., Vallad, G. E., & Breitbart, M. (2011). Exploring the Diversity of Plant DNA Viruses and Their Satellites Using Vector-Enabled Metagenomics on Whiteflies. PLoS ONE, 6(4), e19050.
- Sánchez-Campos, S., Domínguez-Huerta, G., Díaz-Martínez, L., Tomás, D. M., Navas-Castillo, J., Moriones, E., & Grande- Pérez, A. (2018). Differential Shape of Geminivirus Mutant Spectra Across Cultivated and Wild Hosts With Invariant Viral Consensus Sequences. Frontiers in Plant Science, 9.
- Legg, J. P., Ndjelassili, F., & Okao-Okuja, G. (2004). First report of cassava mosaic disease and cassava mosaic geminiviruses in Gabon. Plant Pathology, 53(2), 232–232.


