Research Article - (2023) Volume 7, Issue 1
Food Born Pathogen Contamination of Some Meat Products in Damanhur City, Egypt
2Department of Food Hygiene and Control, Faculty of Veterinary Medicine, Aswan University, Egypt
3Researcher of Bacteriology, Animal Health Research Institute, Damanhur branch, Egypt
4Department of Pharmacology and Therapeutics, Faculty of Veterinary Medicine, Damanhur University, Da, Egypt
5Universidade de Vigo, Nutrition and Bromatology Group, Department of Analytical Chemistry and Food S, Egypt
Received Date: Feb 12, 2023 / Accepted Date: Mar 10, 2023 / Published Date: Mar 27, 2023
Copyright: ©Â©2023 Amal Mohamed El-Sayed, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Abdelfattah, S. A. S., El-Sayed, A. M., Abdelgawad, M.H., Batiha, G. E., Simal-Gandara, J.S. (2023). Food Born Pathogen Contamination of Some Meat Products in Damanhur City, Egypt. Stem Cell Res Int, 7(1), 08-18.
Abstract
The study assessed the chemical and microbial quality of some meat products (Liver, Luncheon, minced meat, and Sausage) in addition to detecting some virulence genes associated with some isolated strains. Two hundred random samples of meat products including 50 samples each of (minced meat, Liver, luncheon and sausage) were randomly collected from different butchers at Damanhur city, El Behera Governorate for some chemical (TVN and TBA) and microbiological evaluation (APC, Staph. aureus, E.coli, Salmonella, Bacillus cereus, and Clostridium perfringens), in addition, detection of virulence genes in some isolated strains. The obtained results revealed that mean values of TVB-N and TBA was higher in liver than other meat products. In regard to microbiological evaluation, results revealed that incidence of staph. aureus, E.coli, Salmonella, Bacillus cereus and clostridium perfringens were (40, 24, 20 and 30%), (36, 10, 8 and 30%), (2, 4, 0, 4%), (16, 20, 24 and 34%) and (28, 10, 18, and 36%), in the examined minced meat, liver, luncheon and sausage, respectively. Salmonella could not be detected in examined luncheon samples. Regarding virulence genes (Pvl and Sea) and (invA and Stn) were detected in 100% of isolated Staph. aureus and Salmonella strains, respectively. eaeA was detected in 100% of isolated E.coli strains while Stx2 was detect- ed in only one strain. Isolated strains of Clostridium perfringens were positive for enterotoxin gene (cpe) and alpha-toxin (cpa) at percent 100%. The presence of aerobic bacteria like Staphylococci aureus, E. coli, bacillus cereus, Salmonella, and Clos- tridium perfringens may be due to poor hygienic measures during processing and handling of meat products. So, consumption of these products could be associated with possible risk of infection, suggesting the need for the institution of strict hygienic measures during handling of meat products.
Keywords
TVB-N, TBA, E. Coli, Salmonella, Staph Aureus.
Introduction
Meat and meat products over time have been considered as the heart of convenience food trend [1]. Meat products such as sau- sage, luncheon, minced meat and burger are highly demanded, be- cause they are more attractive for consumers than fresh meat due to their good taste and high nutritive value also, it is easy prepare within short time. Meat products contamination can occur during preparation, handling and storage practices [2]. Bacterial contam- ination of meat products can be measured using the aerobic plate count, total Escherichia coli. and Staphylococci. This gives a good idea on possible hygienic measures put in place during production of meat product [3]. E. coli gives reliable information on possible fecal contamination [4].
The major cause of food born disease burden in most develop- ing countries is contamination of food with Salmonella species, Staph. aureus and E. coli. These pathogens are transmitted mainly through consumption of contaminated food products with them. The presence of these pathogens in raw meat or meat products has public health implications [5]. Globally Staph. aureus is food borne pathogen due to their ability to produce extracellular toxins responsible for its virulence. These toxins cause sudden abdominal discomfort resulting in vomiting, nausea and abdominal cramps [6]. There are five major classes of Staphylococcal enterotoxins (SEs) recognized globally include SEA, SEE, SEB, SEC and SED. Another class worth noting is the toxic shock syndrome toxin (TSST-1) which is responsible for the toxic shock syndrome ex- perienced in humans during infection of Staph. aureus [7]. Some strains of Staph. aureus produced Panton-Valentine leukocidin (PVL) causes necrotizing pneumonia and associated with various kinds of soft tissue and skin reactions [8]. Staph. aureus has been recognized as an indicator of inefficiency during thermal process- ing of food products, including meat products. Staph. aureus in- dicate poor hygiene during food production or preparation. They also indicate improper cooling of food products after preparation [9].
The main sources of Salmonella infections were found in meat products so, it is very important to make surveillance for its level of contamination in meat products to prevent severe diseases [10]. The enteropathogenicity of Salmonella is dependents on different virulence genes include fimbrial related gene (spvC), the entero- toxin (stn) and invasion (invA) gene which is responsible for Sal- monella invasion and adhesion to the epithelium of the intestine and spvC is responsible for the exaggerated systemic spreading of Salmonella after inactivation of macrophage. The stn gene is re- sponsible for severe and life-threatening diarrhea associated with Salmonella infection [11]. E. coli cause foodborne diseases as it has virulence factors includes intimin (eaeA), Shiga toxin (stx)1 & 2. [12].
All strains of C. perfringens produce alpha (α) toxin and encoded by the cpa gene. Some strains have the ability to produce C. per- fringens enterotoxin encoded by cpe gene, which is responsible for food poisoning [13]. All C. perfringens strains produces alpha toxin, a necrotizing toxin causing liver damage and acute pulmo- nary (The center for food security and public health 2004). The C. perfringens ability to produce spores confer them with ability to survive adverse conditions during food processing. Contamina- tion of food products by C. perfringens spore may occur before or during food processing or following unhygienic handling of food products [14]. In Egypt, there is a lack of information about prob- lems related to the consumption of meat meals and the incidence of food-borne diseases. Therefore, the present study aimed to as- sess the microbial and chemical quality of some meat products (Luncheon, Liver, minced meat and Sausage) in addition to detec- tion of some virulence genes associated with some isolated strains.
Materials and Methods
Collection of Samples
A total of 200 random meat product samples including minced meat, Liver, luncheon and sausage (50 samples of each) were collected from different supermarkets at Damanhur city. Samples were kept for the subsequent examinations in a separate sterile plastic bag and transferred in an ice box as soon as possible to the laboratory under aseptic conditions.
Determination of Chemical Parameters
• The Determination of Total Volatile Nitrogen (TVB-N) (mg/100g) was done using Conway's test as outlined by [15].
• The Determination of Thiobarbituric acid number (TBA) (mg malonaldehyde /kg sample) as described by [16].
Bacteriological Examination of Meat Products
Sample Preparation The samples preparation was done according to 25 grams of each sample were aseptically transferred into sterile blender flask con- taining 225 ml of sterile peptone water 1% and homogenized at 14000 rpm for 2.5 minutes [17].
Aerobic Plate Count
Aerobic Plate Count (APC) was determined using the standard plate count following [18].
Staph. Aureus Determination
Staph. aureus determination was done using Baird-Parker Agar Plates according to [19]. Suspected colonies of Staph aureus were picked up onto slants of nutrient agar for further purification and morphological identification using Gram-stain; biochemically and coagulase activities according to [18].
Isolation and Identification of Enteropathogenic e. Coli
Isolation and identification of enteropathogenic E. coli was done according to [20].
Isolation and Identification of Salmonellae
Isolation and identification of Salmonellae was done according to [21]. Suspected colonies of Salmonella isolates with red and with black centers on XLD agar were identified morphologically by Gram-stain and biochemically according to [19]. All salmonella isolates were subjected to serological typing by slide agglutination test in animal health research institute, Damanhur branch, Egypt according to using standard polyvalent and monovalent Salmonel- la antisera (Seiken, Japan) [22].
Isolation and Identification of Bacillus Cereus
Isolation and identification of B. cereus was done according to us- ing PEMPA agar and incubated at 37ºC for 24 hours. Peacock blue colored colonies (3-5mm) surrounded by blue zone of egg yolk hy- drolysis against green/greenish yellow background were presumed to be B. cereus [23].
Detection of Clostridium Perfringens
Detection of C. perfringens was done according to using Tryp- tose Sulfite Cycloserine (TSC) media at 37 ± 1 â?¦C under anaerobic conditions for 20 ± 2 h. Presumptive C. perfringens appeared as black colonies surrounded by opaque white zones approximately 2–4 mm due to lecithinase activity [24].
Detection of Virulence Genes in Isolated Strains by Pcr
DNA Extraction: DNA extraction from three isolates from Staph. aureus, Salmonella species and E. coli and two isolates of C. per- fringens was performed using the QIAamp DNA Mini kit (Qiagen, Germany, GmbH) with modifications from the manufacturer’s recommendations. Briefly, 200 µl of the sample suspension was incubated with 10 µl of proteinase K and 200 µl of lysis buffer at 56OC for 10 min. After incubation, 200 µl of 100% ethanol was added to the lysate. The sample was then washed and centrifuged following the manufacturer’s recommendations. Nucleic acid was eluted with 100 µl of elution buffer provided in the kit.
Oligonucleotide Primer: Primers used were supplied from Me- tabion (Germany) are listed in table (A). PCR amplification: For PCR, primers were utilized in a 25-µl reaction containing 12.5 µl of Emerald Amp Max PCR Master Mix (Takara, Japan), 1 µl of each primer of 20 pmol concentration, 4.5 µl of water, and 6 µl of DNA template. The reaction was performed in an applied biosys- tem 2720 thermal cycler.
Analysis of the PCR Products: The products of PCR were sepa- rated by electrophoresis on 1.5% agarose gel (AppliChem, Germa- ny, GmbH) in 1x TBE buffer at room temperature using gradients of 5V/cm. For gel analysis, 20 µl of the PCR products were loaded in each gel slot. Gene ruler 100 bp ladder (Fermentas, Thermo Scientific, Germany) was used to determine the fragment sizes. The gel was photographed by a gel documentation system (Alpha Innotech, Biometra) and the data was analyzed through computer software.
Statistical Analysis
Data was expressed as mean ± SEM using Statistical data analysis was carried out using SPSS 17.0 for windows (SPSS Inc, Chicago, IL, USA).
Results
Determination of Chemical Parameters
The mean values of TVN in examined minced meat, liver, lun- cheon and sausage were 11.90±0.80, 14.23±0.81, 11.31±0.67 and 13.40±0.92 mg/100 gm; respectively while, the respective mean of TBAs were 0.56±0.04, 0.79±0.05, 0.54±0.05 and 0.40±0.02 mg MD/kg (Table 1).
Table 1: Total Volatile Nitrogen and Thiobarbituric Acid in Examined Meat Products Samples
|
Products |
TVB-N (mg/100 gm) |
TBA (mg MD/kg) |
|
Mean ±SEM |
Mean ±SEM |
|
|
Minced meat |
11.90±0.80 |
0.56±0.04 |
|
Liver |
14.23±0.81 |
0.79±0.05 |
|
Luncheon |
11.31±0.67 |
0.54±0.05 |
|
Sausage |
13.40±0.92 |
0.40±0.02 |
Total volatile nitrogen (TVB-N) and thiobarbituric acid (TBA). Value was considered significantly different at p<0.05.
Aerobic Plate Count
Aerobic mesophilic bacteria were detected in all examined meat products samples with mean values of 1.70×106±0.13×106, 5.68×105±0.42×105, 4.38×104±0.38×104 and 7.90×105±0.15×105 Cfu/g, in examined minced meat, liver, luncheon and sausage; respec- tively (Table 2).
Table 2: Statistical Analytical Results of Aerobic Plate Count in Examined Meat Products Samples
|
Products |
No. of examined samples |
Positive samples |
Min |
Max |
Mean ±SEM |
|
|
No |
% |
|||||
|
Minced meat |
50 |
50 |
100 |
1.6×105 |
2.92×106 |
1.70×106±0.13×106 |
|
Liver |
50 |
50 |
100 |
1.2×105 |
1.2×106 |
5.68×105±0.42×105 |
|
Luncheon |
50 |
50 |
100 |
6.0×103 |
9.8×104 |
4.38×104±0.38×104 |
|
Sausage |
50 |
50 |
100 |
6.0×105 |
9.5×105 |
7.90×105±0.15×105 |
Staph. aureus Determination
Staph. aureus was detected at level of 40, 24, 20 and 30% in examined minced meat, liver, luncheon and sausage with mean values of 3.15×104±0.47×104, 3.33×103±0.44×103, 5.20×102±0.41×102 and 1.85×103±0.32×103 Cfu/g; respectively (Table 3).
Table 3: Statistical Analytical Results of Staph. Aureus in Examined Meat Products Samples
|
Products |
No. of examined samples |
Positive samples |
Min |
Max |
Mean ±SEM |
|
|
No |
% |
|||||
|
Minced meat |
50 |
20 |
40 |
1.0×104 |
1.0×105 |
3.15×104±0.47×104 |
|
Liver |
50 |
12 |
24 |
1.5×103 |
6.0×103 |
3.33×103±0.44×103 |
|
Luncheon |
50 |
10 |
20 |
3.2×102 |
6.8×102 |
5.20×102±0.41×102 |
|
Sausage |
50 |
15 |
30 |
5.0×102 |
5.0×103 |
1.85×103±0.32×103 |
Staph. aureus Determination
Staph. aureus was detected at level of 40, 24, 20 and 30% in examined minced meat, liver, luncheon and sausage with mean values of 3.15×104±0.47×104, 3.33×103±0.44×103, 5.20×102±0.41×102 and 1.85×103±0.32×103 Cfu/g; respectively (Table 3).
Isolation and Identification of Enteropathogenic e. Coli
The incidences of enteropathogenic E. coli in examined minced meat, liver, luncheon and sausage were 36, 10, 8 and 30%; respectively (Table 4)
Table 4: Incidence of Enteropathogenic e. Coli in Examined Samples of Meat Products
|
Products |
No. of examined samples |
Positive samples |
|
|
No |
% |
||
|
Minced meat |
50 |
18 |
36 |
|
Liver |
50 |
5 |
10 |
|
Luncheon |
50 |
4 |
8 |
|
Sausage |
50 |
15 |
30 |
Isolation and Identification of Salmonellae
The incidence of Salmonella in minced meat, liver and sausage were 2, 4 and 4%; respectively (Table 5).
Table 5: Incidence and Serotyping Of Salmonella Species Isolated From Examined Meat Products Samples
|
Products |
No. of examined samples |
Positive samples |
Salmonella serotyping for 5 Salmonella species |
||||
|
No |
% |
S. Enteritidis |
S. typhimurium |
||||
|
No |
%+ |
No |
%+ |
||||
|
Minced meat |
50 |
1 |
2 |
1 |
20 |
0 |
0 |
|
Liver |
50 |
2 |
4 |
1 |
20 |
1 |
20 |
|
Luncheon |
50 |
0 |
0 |
0 |
0 |
0 |
0 |
|
Sausage |
50 |
2 |
4 |
1 |
20 |
1 |
20 |
%+ acc. to No of isolated strains
Isolation and identification of Bacillus cereus Bacteriological examination of meat products samples revealed that B. cereus species were found in 16, 20, 24 and 34 % of exam- ined minced meat, liver, luncheon and sausage with mean values of 3.57×103±0.54×103, 3.20×103±0.38×103, 3.08×103±0.46×103 and 1.61×104±0.11×104 Cfu/g; respectively (Table 6).
Table 6: Incidence of Bacillus Cereus in Examined Samples of Meat Products.
|
Products |
No. of examined samples |
Positive samples |
Min |
Max |
Mean ±SEM |
|
|
No |
% |
|||||
|
Minced meat |
50 |
8 |
16 |
4.0×102 |
5.0×103 |
3.57×103±0.54×103 |
|
Liver |
50 |
10 |
20 |
1.4×103 |
5.0×103 |
3.20×101±0.38×103 |
|
Luncheon |
50 |
12 |
24 |
1.0×102 |
6.0×103 |
3.08×103±0.46×103 |
|
Sausage |
50 |
17 |
34 |
8.0×103 |
2.5×104 |
1.61×104±0.11×104 |
Detection of Clostridium Perfringens
Clostridium perfringens were detected in examined minced meat, liver, luncheon and sausage at incidence rate 28, 10, 18 and 36% with mean values of 2.6×103±0.24×103, 2.16×103±0.26×103, 1.75×102±0.24×102 and 2.59×103±0.24×103 Cfu/g; respectively (Table 7).
Table 7: Incidence of Clostridium Perfringens in Examined Samples of Meat Products
|
Products |
No. of examined samples |
Positive samples |
Min |
Max |
Mean ±SEM |
|
|
No |
% |
|||||
|
Minced meat |
50 |
14 |
28 |
1.3×103 |
4.0×103 |
2.6×103±0.24×103 |
|
Liver |
50 |
5 |
10 |
1.4×103 |
2.8×103 |
2.16×103±0.26×103 |
|
Luncheon |
50 |
9 |
18 |
9.0×10 |
3.2×102 |
1.75×102±0.24×102 |
|
Sausage |
50 |
18 |
36 |
1.2×10 |
4.0×103 |
2.59×103±0.24×103 |
Detection of Virulence Genes in Isolated Strains by Pcr The present investigation showed that all three tested Staph. aureus strain 100% harbored pvl gene at 433 bp gene and sea at 102 bp by PCR (Fig 1). PCR results obtained in Fig 2 showed that eaeA gene (248 bp) detected in 100 % of all isolated E. coli Strain while Stx2 gene (779 bp) detected only in one isolated E. coli strain. stn gene and invA were detected at 617 and 284bp in three Salmonella isolates (Fig 3). The two tested C. perfringens strains possess enterotoxin gene (cpe) at 247 bp and alpha toxin (cpa) at 402 bp by PCR (Fig 4)
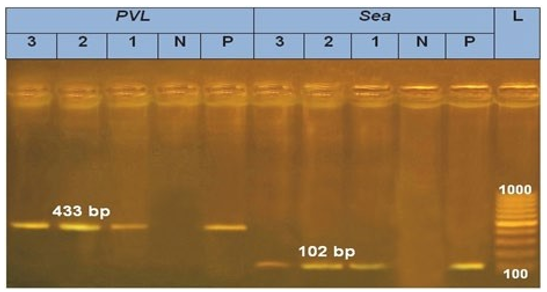
Figure 1: Agarose gel showing polymerase chain reaction amplification products of pvl (433 bp) and sea gene (102 bp) in three staph. aureus strain, l:100-1000 bp dna ladder, pos: control positive for sea and pvl, neg: control negative for sea and pvl, lane 1-3: positive strain for pvl, lane 1-3: positive strain for sea.
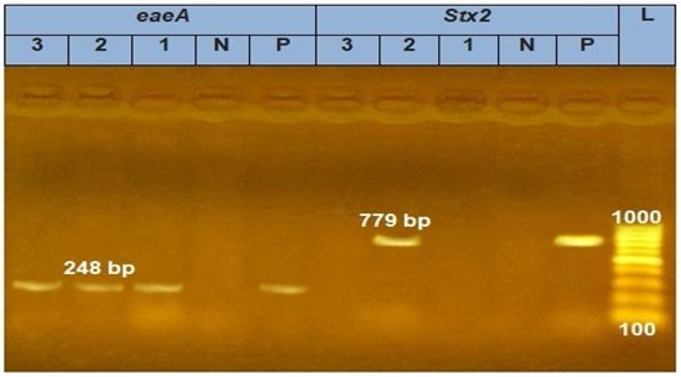
Figure 2: Agarose gel showing polymerase chain reaction amplification products of stx2 (779 bp) and eaeA gene (248 bp) in three E.coli strain, L:100-1000 bp DNA ladder, pos: control positive for eaeA and Stx2, Neg: control negative for eaeA and Stx2, lane 1-3: positive strain for eaeA , lane 2: positive strain for Stx2.
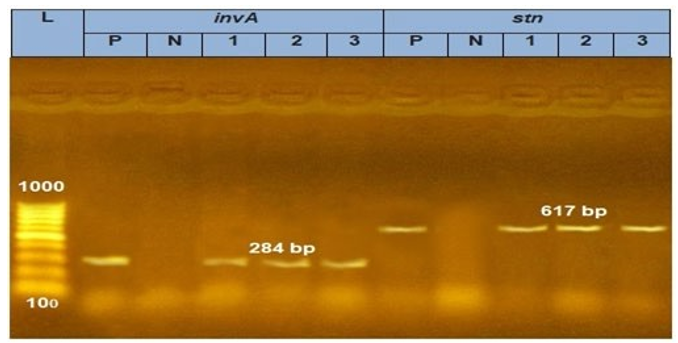
Figure 3: Agarose gel showing polymerase chain reaction amplification products of stn gene (617 bp) and invA (284) in three Salmonel- la isolates, L:100-1000 bp DNA ladder, pos: control positive for stn and invA, Neg: control negative for stn and invA, lane 1-3: positive strain for invA , lane 1-3: positive strain for stn.
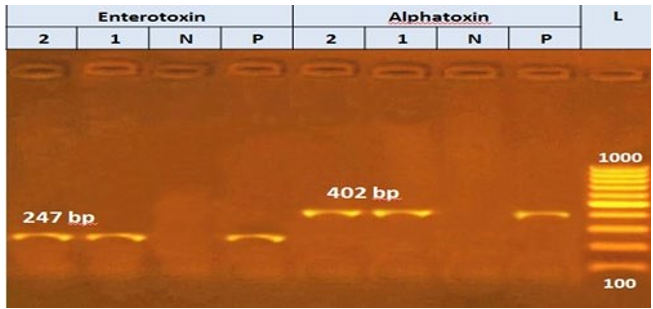
Figure 4: Agarose gel showing polymerase chain reaction amplification products of Alpha toxin (cpa) at 402 bp and enterotoxin (cpe) gene at 247 bp in two clostridium perfringens strain, L:100-1000 bp DNA ladder, pos: control positive for cpa and cpe, Neg: control negative for cpa and cpe, lane 1-2: positive strain for cpa , lane 1-2: positive strain for cpe.
Discussion
The contamination of meat products and meat with microorgan- isms from meat handlers, which may have carried the pathogenic microorganism during the processes of packing, manufacturing and marketing. Poor hygiene during production processes, refrig- eration or the retail and storage of foods or improper cooking may lead to food poisoning or meat borne illness causing increase in disease burden and consequent death in most developing countries [25]. Disease burden from improper food handling has swallowed billions of dollars in social costs and medical care [25]. TVB-N value has been used for assessing the degree of deterioration of meat. On the other, TBA is a good indicator of the quality of meat, since it is able to assess malondialdehyde concentrate on which considered as a secondary product of lipid peroxidation and a good indicator for assessing the lipid oxidation level [26].
The TVN high level in the different meat product may be due to post processing circumstances. These circumstances include fail- ure in freezing during marketing, storage and distribution [27]. Bad handling during processing and the use of old meat may have been responsible for the recorded high TBA values in the pres- ent studies. The above-mentioned factors may have resulted in the excessive oxidation of fat in the different meat products. Poor technology in the different factories handling meat products may be another possible explanation for increase in TBA level in the different meat products. Reported a mean TVN values of 13.06 ±0.04 in the liver sample they examined which was almost close to the values of TVN we observed in the present study [27]. Reported a mean TVN value of 12.60 in minced meat which was slightly higher than what we observed in the present study [28]. Higher TVN value was also reported by who reported 16.23, 24.69 and 22.1 for sausage, minced meat and luncheon; respectively [29]. Lower results of TBA (0.16 ± 0.01) in liver was reported by [27]. reported 0.17 ± 0.02 in liver samples; reported 0.45 and 0.25 in sausage and luncheon; respectively. Higher TBA in minced (0.7) was reported by and in sausage (0.68) by [28-30]. Mean values of TVN in all the examined meat products we sampled were within the safe acceptable limit (should not exceed 20 mg/100 gm) as recommended by for TVN in frozen liver, minced meat and sau- sage [31]. In addition, according to same specifications stated that TBA mean value in the examined meat products were within the acceptable limit (should not exceed 0.9 mg malonaldehyde/kg of sample) as recommended by [31].
The obtained result in Table (2) revealed that aerobic mesophil- ic bacteria were detected in all examined meat products samples with mean values of 1.70×106±0.13×106, 5.68×105±0.42×105, 4.38×104±0.38×104, and 7.90×105±0.15×105 Cfu/g, in examined minced meat, liver, luncheon and sausage; respectively. Lower results of aerobic bacterial count in examined sausage, minced meat and luncheon was mentioned by 3.3x105, 3.3x105 and 3.3x105 Cfu/g; respectively [29]. In addition, reported that mean value of aerobic plate count in examined minced meat, sausage and luncheon were 8.03×104 ±0.12×104, 6.74×104 ±0.28×104 and 5.85×104 ±0.24×104 Cfu/g; respectively [32]. The total aerobic counts reflect the bacterial contamination level and hygienic qual- ity of meat products [33].
Recorded data in table (3) showed that Staph. aureus was detected at level of 30, 24, 20 and 40% in examined sausage, liver, lun- cheon and minced meat with mean values of 1.85×103±0.32×103, 3.33×103±0.44×103, 5.20×102±0.41×102 and 3.15×104±0.47×104 Cfu/g; respectively. Lower level of Staph. aureus was reported by 0.2x102 Cfu/g in minced meat; 0.1x102 Cfu/g in luncheon and 0.3x102 Cfu/g in sausage [29]. Higher results of Staph. aureus in sausage was reported by 2.8x104 Cfu/g. Lower incidence of Staph. aureus in luncheon, sausage and minced meat 8,6, 20 and 25.7 % was reported by [32, 34]. Staph. aureus can be carried in nasal pas- sage and on the hands. Most outbreaks of food borne disease due to contamination following poor handling and production of heat stable toxins Staph. aureus. Refrigeration, proper cooking and san- itary food handling are recommended measure to prevent disease outbreak related to Staph. aureus [35].
The present investigation showed that all three tested Staph. au- reus strains 100% harbored pvl gene at 433 bp and sea at 102 bp by PCR (Fig 1). could detected Sea gene in one Staph. aureus strain isolated from meat products [36]. Results from the present study are line with the finding of who recorded detection of enterotoxi- genic Staph. aureus isolates in meat samples by using multiplex PCR [37]. The enterotoxin genes detection in given strain of Staph. Aureus by using PCR is very important for the identification of staphylococcal related food poisoning [38]. Results demonstrated in table (4) revealed that the incidences of enteropathogenic E. coli in examined minced meat, liver, luncheon and sausage were 36, 10, 8 and 30%; respectively. Presence of E. coli is a good mark- er for possible contamination of food product with fecal material [4]. Lower incidence of E. coli in minced meat (22.9 & 16 %), in sausage (17.1 & 20%) were reported by and respectively [32, 39].
In addition, cited that the incidence of E. coli in sausage was 9% [40]. Nearly similar incidence of E. coli in luncheon 8% was re- ported by while, higher incidence of E. coli in luncheon was 36 and 28% reported by and [41-43]. Lower incidence (5.7%) in luncheon was reported by [32]. The practices of manufacture and storage, handling from production sites to the consumers and effectiveness of hygienic measures may be responsible for the observed varia- tion reported by the different authors. PCR results obtained in Fig (2) showed that eaeA gene (248 bp) detected in 100 % of all iso- lated E. coli Strain while, Stx2 gene (779 bp) detected only in one isolated E. coli strain. These results agreed with who reported that eaeA gene (248 bp) was detected in 100% of isolated E. coli strains from meat products. In contrary, these results disagreed with who reported that E. coli isolates from minced meat and sausage were negative for Stx2 at 779 bp [30, 44]. Abovementioned results in table (5) showed that the incidence of Salmonella in minced meat, liver and sausage were 2, 4 and 4%; respectively while, Salmo- nella was not detected in examined luncheon samples. Salmonella species isolated from minced meat were serologically identified into Sal. enteritidis 100; from liver into Sal. enteritidis (50%) and Sal. typhimurium (50%) finally from sausage into Sal. enteritidis (50%) and Sal. typhimurium (50%).
Our finding agreed with who reported that Salmonellae failed to be detected in the examined luncheon samples [45]. the same au- thor could isolate Salmonella from sausage at incidence rate 10% and serologically identified in Sal. typhimurium (40%), Sal. typhi (40%) and Sal. enteritidis (20%) while, isolated Salmonella from minced meat at incidence rate 6% and serologically identified into Sal. typhi 33.3% and S. typhimurium 66.2%. In contrary, could iso- late Salmonella from luncheon at incidence rate 12% [39]. Nearly similar incidence of Salmonella in minced meat 2.8% was reported by while, higher incidence in minced meat 24% was reported by [39, 46]. Nearly similar incidence of Salmonella in sausage 4% was reported by while, higher results 24% were reported by [29].
In the present study PCR assay for detection of invA and Stn gene in three Salmonella isolates revealed that, two genes detected in all isolates (100%) at 617 and 284 bp PCR amplified fragment (Fig, 3). The results were agreed with who mentioned that all sev- en isolated Salmonella from meat products (100%) harbored invA gene at 284 bp. In addition, confirmed the presence of Stn gene in all identified Salmonella isolates form meat products [47, 48]. cited that, PCR is a specific, fast highly reliable and sensitive pro- cedure used for characterization of Salmonella when compared to traditional microbiological techniques [49]. Using primers of invA specific for Salmonella in PCR significantly decreases possibility of false negative results. Amplification of invA gene is considered as an international standard for detection of Salmonellae identifi- cation. It contains sequences only for Salmonella and proved to be a suitable PCR target, with an important diagnostic application [50]. The stn gene is a very important Salmonellae virulent genes which assist Salmonellae in expressing its virulence in the host and manifestation of its pathogenicity (mainly diarrhea) associated with Salmonellae infection [51].
As shown in Table (6) bacteriological examination of meat prod- ucts samples revealed that B. cereus were found in 16, 20, 24 and 34 % of examined minced meat, liver, luncheon and sausage with mean values of 3.57×103±0.54×103, 3.20×103±0.38×103, 3.08×103±0.46×103 and 1.61×104±0.11×104 Cfu/g; respectively. Nearly similar incidence of B. cereus in minced meat 17.14 % and in luncheon 20 % was reported by [52]. Higher incidence of B. cereus in minced meat 76% and luncheon 44% were reported by [53]. In contrary, could not isolate B. cereus from minced meat. B. cereus had been implicated in many disease outbreaks related to food poisoning [54]. These outbreaks are mostly characterized by abdominal pain and diarrheal syndromes however, there are many cases of food poisoning with similar signs caused by Staph. au- reus and C. perfringens [55]. B. cereus survival in meat and milk products which attributed to its spore formation ability therefore, increasing its possible viability [56]. Tabulated results in table (7) showed that C. perfringens were detected in examined minced meat, liver, luncheon and sausage at incidence rate 28, 10, 18 and 36% with mean values of 2.6×103±0.24×103, 2.16×103±0.26×103, 1.75×102±0.24×102 and 2.59×103±0.24×103 Cfu/g; respectively.
Higher incidence of Clostridia obtained by who cited that the in- cidence of Clostridia in sausage and luncheon examined samples were 60 and 52%; respectively [57]. In addition, and reported higher incidence of Clostridia in minced meat which was 93.3 and 55 %; respectively [58, 59]. Presence of large number of Clos- tridia in meat products may be responsible for lowered quantity of meat products, with consequent public health related hazards and economic losses [60]. The variation in the incidence of Clostridia in minced meat may be attribute to poor hygienic measures during processing. Clostridia spores could be separately added to meat products during cutting, wrapping and handling of meat [58]. The C. perfringens ability to produce alpha toxin (cpa) and enterotox- ins (cpe) which are responsible for possible food poisoning has remained of public health relevance [61]. PCR results in Fig (4) declared that two tested C. perfringens strains possess enterotox- in gene (cpe) at 247 bp and alpha toxin (cpa) at 402 bp by PCR. These results agree with who found that all tested C. perfringens type A (33, 100%); respectively possess alpha cpa at 204 bp and cpe at 247 bp [58].
Conclusion
The present study supported the fact meat products are of possible public health concern. The presence of aerobic bacteria like Staph. aureus, E. coli, bacillus cereus, Salmonella and C. perfringens may be due to poor hygienic measures during processing and handling of meat products. So, consumption of these products could be as- sociated with possible risk of infection. Therefore, efficient heat treatment, adequate cleaning and sanitization of utensils may assist reducing this contamination and implement hazard analysis may lead to an improvement in hygiene.
Implications for Policy and Practice
Meat products such as minced meat, sausage, luncheon and burger are highly demanded because in most cases they are considered more attractive for consumers than fresh meat due to their good taste and high nutritive value. Despite the importance of meat products in human nutrition, their contamination with various types of foodborne pathogens can occur. The presence of aero- bic bacteria like Staph. aureus, E. coli, bacillus cereus, Salmonella and C. perfringens may be due to poor hygienic measures during the processing and handling of meat products. So, consumption of these products could be associated with the possible risk of in- fection, suggesting the need for the legislation and institution of strict hygienic measures during the handling of meat products by policymakers.
References
- Leroy, F., & Degreef, F. (2015). Convenient meat and meat products. Societal and technological issues. Appetite, 94, 40- 46.
- Taulo, S., Wetlesen, A., Abrahamsen, R., Kululanga, G., Mka- kosya, R., & Grimason, A. (2008). Microbiological hazard identification and exposure assessment of food prepared and served in rural households of Lungwena, Malawi. Internation- al journal of food microbiology, 125(2), 111-116.
- Hamed, E. A., Ahmed, A. S., & Abd El-Aaty, M. F. (2015). Bacteriological hazard associated with meat and meat prod- ucts. Egypt. J. Agric. Res, 93(4), 385-393.
- Synge, B. A. (2000). Verocytotoxinâ?ÂÂproducing Escherich- ia coli: a veterinary view. Journal of applied microbiology, 88(S1), 31S-37S.
- Zafar, A., Ahmed, E., Wajiha, H., & Khan, A. B. (2016). Mi- crobiological evaluation of raw meat products available in local markets of Karachi, Pakistan: Microbial evaluation of raw meat products. Proceedings of the Pakistan Academy of Sciences: Part B (Life and Environmental Sciences), 53(2), 103-106.
- Ünal, N., & Çinar, O. D. (2012). Detection of stapylococcal enterotoxin, methicillin-resistant and Panton–Valentine leu- kocidin genes in coagulase-negative staphylococci isolated from cows and ewes with subclinical mastitis. Tropical animal health and production, 44, 369-375.
- Chiang, Y. C., Chang, L. T., Lin, C. W., Yang, C. Y., & Tsen,H. Y. (2006). PCR primers for the detection of staphylococcal enterotoxins K, L, and M and survey of staphylococcal en- terotoxin types in Staphylococcus aureus isolates from food poisoning cases in Taiwan. Journal of food protection, 69(5), 1072-1079.
- Shallcross, L. J., Fragaszy, E., Johnson, A. M., & Hayward, A.C. (2013). The role of the Panton-Valentine leucocidin toxin in staphylococcal disease: a systematic review and meta-anal- ysis. The Lancet infectious diseases, 13(1), 43-54.
- Jay, J. (2000). Modern Food Microbiology. 6th Ed., Aspen Publishers, Inc., Gaithersburg.
- Ye, Y., Wu, Q., Zhang, J., Lu, J., & Lin, L. (2011). Isolation of Salmonella from meat samples and characterization by enterobacterial repetitive intergenic consensus–polymerase chain reaction and antibiotics test. Foodborne Pathogens and Disease, 8(8), 935-937.
- Saini, K., Kaushal, A., Gupta, S., & Kumar, D. (2019). Mul- tiplexed Stn and PlcA based specific genetic marker for early detection of Salmonella enterica and Listeria monocytogenes in milk samples. The Annals of the University Dunarea de Jos of Galati. Fascicle VI-Food Technology, 43(2), 9-20.
- Kang, S. J., Ryu, S. J., Chae, J. S., Eo, S. K., Woo, G. J., &Lee, J. H. (2004). Occurrence and characteristics of entero- hemorrhagic Escherichia coli O157 in calves associated with diarrhoea. Veterinary microbiology, 98(3-4), 323-328.
- Juneja, V. K., & Sofos, J. N. (2009). Pathogens and toxins in foods: challenges and interventions. ASM Press.
- García, S., Araiza, M., Gómez, M., & Heredia, N. (2002). Inhibition of growth, enterotoxin production, and spore for- mation of Clostridium perfringens by extracts of medicinal plants. Journal of Food Protection, 65(10), 1667-1669.
- FAO "Food and Agriculture Organization "(1980). Manual of Food Quality Control. FAO, United Nation, Rome, Italy.
- Kirk, S., & Sawyer, R. (1991). Pearson's composition and analysis of foods (No. Ed. 9). Longman Group Ltd..
- ICMSF "International commission of Microbiological Speci- fication for Foods" (1996). Microorganisms in Food. I-Their Significance and methods of enumeration.3rd Ed. Univ. of Toronto, Canada.
- FDA (Food Drug Administration) (2001). Food borne patho- genic microorganisms and natural toxins. Center for food safety and applied nutrition.US, FDA, Washington, DC.
- Quinn, P. J., Markey, B. K., Carter, M. E., Donnelly, W. J. C., & Leonard, F. C. (2002). Veterinary microbiology and micro- bial disease. Blackwell science.
- ISO "International Organization of Standardization" (2001). Microbiology of food and animal feeding stuffs. Horizon- tal method for the enumeration of β- glucuronidas- Positive E-Coli. Part 2: Colony- Count technique at 44 º c using 5 bro- mo-4-chloro-3- indolyl β-D-glucuronide. 16649-2.
- ISO, International Standard Organization” (2002). Interna- tional Organization for Standardization. No.6579. Microbiol- ogy of Food and Animal Feeding Stuffs. Horizontal Methods for detection of Salmonella species.
- Grimont, P. A., & Weill, F. X. (2007). Antigenic formulae of the Salmonella serovars. WHO collaborating centre for refer- ence and research on Salmonella, 9, 1-166.
- Rasool, U., Ahmad, A., Badroo, G. A., Mudasir, M., Fayaz, S., & Mustafa, R. (2017). Isolation and identification of Ba- cillus cereus from fish and their handlers from Jammu, India. International Journal Current Microbiology Applied Science, 6, 441-447.
- ISO, E. (2004). 7937. Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of Clostridium Perfringens—Colony-Count Technique. Interna- tional Organization for Standardization: Geneva, Switzerland.
- Food and Drug Administration "FDA" (2012). Bad bug book Food borne Pathogenic Microorganisms and Natural Toxins Handbook, 2nd ed. Staphylococcus aureus.
- Hemmat, M. I., Reham, A. A., Omima, A. S., & El Shafay, M.S. (2013). Quality of beef and edible offal at abattoir level.
- Benha Veterinary Medical Journal, 25(2), 254-263.Hemmat, M. I., Reham, A. A., Omima, A. S., & El Shafay, M.S. (2013). Quality of beef and edible offal at abattoir level.Benha Veterinary Medical Journal, 25(2), 254-263.
- Kortoma, S. (2015). Chemical composition of some Egyptian meat products. Ph.D. thesis, Alexandria University.
- Hassanien, E. S., Fahim, S. A., Mohammed, H. F., Lotfy, L. M., & Hatem, E. M. (2018). Quality assurance of some meat products. J Dairy Vet Anim Res, 7(4), 171-174.
- Hassan, M. A., & Amin, R. (2020). Assessment of chemical quality of local frozen cattle's liver. Benha Veterinary Medical Journal, 38(2), 97-100.
- ES “Egyptian Organization for Standardization and Quality Control” (2005). Egyptian Organization for Standardization and Quality Control. No. 1473 for frozen liver, 1694 for fro- zen minced meat and 1972 for frozen sausage.
- Shaltout, F. A., Maarouf, A. A., El-Kewaiey, I., & Heweidy,Y. (2016). Prevalence of some foodborne microorganisms in meat and meat products. Benha Veterinary Medical Journal, 31(2), 213-219.
- Paulsen, P., Schopf, E., & Smulders, F. J. M. (2006). Enumer- ation of total aerobic bacteria and Escherichia coli in minced meat and on carcass surface samples with an automated most-probable-number method compared with colony count protocols. Journal of food protection, 69(10), 2500-2503.
- El-Tawab, M. M. (2014). Studies on mycotoxins in some meat products. MV Sc. Meat Hygiene), Fac. Vet. Med., Benha Univ. Egypt.
- Food Safety and Inspection service "FSIS" (2003). United states Department of Agriculture; Meat preparation: Beef from farm to table, Washington. DC.
- Saif, M., Saad, S., Shaltout, F., Hassanin, F. S., & Zaghloul,M. (2019). Molecular detection of enterotoxigenic Staphylo- coccus aureus in ready to eat beef products. Benha Veterinary Medical Journal, 37(1), 7-11.
- Morshdy, A. E., Hussein, M. A., Tharwat, A. E., & Fakhry,A. (2018). PREVALENCE OF ENTEROTOXIGENICAND MULTIDRUG-RESISTANT Staphylococcus aureusIN READY TO EAT MEAT SANDWICHES. Slovenian Vet-erinary Research, 55.
- Argudín, M. Á., Mendoza, M. C., & Rodicio, M. R. (2010). Food poisoning and Staphylococcus aureus enterotoxins. Tox- ins, 2(7), 1751-1773.
- Eltanani, G. S., & Arab, W. S. (2021). Quality Assurance of Some Meat Products. Alexandria Journal for Veterinary Sci- ences, 69(1).
- Abd El Tawab, A. A., El-Hofy, F. I., Maarouf, A. A., & El- Said, A. A. (2015). Bacteriological studies on some food borne bacteria isolated from Chicken meat and meat products in Kaliobia Governorate. Benha Veterinary Medical Journal, 29(2), 47-59.
- El-Shabrawy, H. (2015). Bacteriological and chemical eval- uation of some locally manufactured beef burger. M. V. SC. Thesis Meat Hygiene, Fac. Vet. Med., Benha Univ., Egypt.
- Ashraf, A. (2016). Microbial evaluation of heat treated meat products. MV Sc (Doctoral dissertation, Thesis, meat hygiene, Fac. Vet. Med., Alexandria University).
- Ramadan, A. M. (2015). Contamination of meat products with human pathogens. MV Sc. Alex. Univ., Egypt.
- Ali Elsayed, A. (2019). Bacteriological and molecular stud- ies on antibiotic resistant Escherichia coli isolated from meat and its products in Kaliobia, Egypt. Benha Veterinary Medical Journal, 36(2), 335-344.
- Edris, A. M., Shaltout, F., Salem, G. H., & El-Toukhy, E. I. (2011). Incidence and isolation of salmonellae from some meat products. Food Science and Technology, 1, 194-200.
- Cohen, N., Filliol, I., Karraouan, B., Badri, S., Carle, I., En- naji, H., ... & Karib, H. (2008). Microbial quality control of raw ground beef and fresh sausage in Casablanca (Morocco). Journal of Environmental Health, 71(4), 51-55.
- Elbakry, H. G. A. E. M., Ezzat, M., & Helal, I. (2020). Prev- alence of virulent genes in Salmonella isolated from some raw meat products. Suez Canal Veterinary Medical Journal. SCVMJ, 25(2), 339-355.
- Sallam, K. I., Mohammed, M. A., Hassan, M. A., & Tamura,T. (2014). Prevalence, molecular identification and antimicro- bial resistance profile of Salmonella serovars isolated from retail beef products in Mansoura, Egypt. Food control, 38, 209-214.
- Oliveira, S. D., Rodenbusch, C. R., Ce, M. C., Rocha, S. L. S., & Canal, C. W. (2003). Evaluation of selective and non-se- lective enrichment PCR procedures for Salmonella detection. Letters in Applied Microbiology, 36(4), 217-221.
- Malorny, B., Hoorfar, J., Bunge, C., & Helmuth, R. (2003). Multicenter validation of the analytical accuracy of Salmonel- la PCR: towards an international standard. Applied and envi- ronmental microbiology, 69(1), 290-296.
- Nayak, R., Stewart, T., Wang, R. F., Lin, J., Cerniglia, C. E., & Kenney, P. B. (2004). Genetic diversity and virulence gene determinants of antibiotic-resistant Salmonella isolated from preharvest turkey production sources. International journal of food microbiology, 91(1), 51-62.
- Gharib, A., El-Aziz, A., & Allam, M. (2020). Multiplex poly-merase chain reaction for detection of toxin genes of Bacillus cereus group isolated from meat and chicken products. Zaga- zig Veterinary Journal, 48(4), 457-470.
- Tharwat, A. E., Eleiwa, N. Z., Ali, N. S. M., & Merwad, A. M.A. (2020). Prevalence and distribution of enterotoxin genes among Bacillus cereus isolated from meat and meat products in Egypt. Advances in Animal and Veterinary Sciences, 8(s1), 41-46.
- Milojevic, L., Velebit, B., Djordjevic, V., Jankovic, V., La- kicevic, B., Bajcic, A., & Betic, N. (2019, September). Screening of Bacillus cereus presence in minced meat and meat products originating from Serbian retail facilities. In IOP Conference Series: Earth and Environmental Science (Vol. 333, No. 1, p. 012079). IOP Publishing.
- Bennett, S. D., Walsh, K. A., & Gould, L. H. (2013). Food- borne disease outbreaks caused by Bacillus cereus, Clostrid- ium perfringens, and Staphylococcus aureus—United States, 1998–2008. Clinical infectious diseases, 57(3), 425-433.
- Christiansson, A., Bertilsson, J., & Svensson, B. (1999). Ba- cillus cereus spores in raw milk: factors affecting the contam- ination of milk during the grazing period. Journal of dairy science, 82(2), 305-314.
- Abdel-Atty, N. S., Khalafalla, F. A., & Barakat, D. A. (2020). Bacteriological Quality of Canned Meat Marketed in Egypt. Benha Veterinary Medical Journal, 39(1), 154-158.
- Younis, G. A., Radwan, M. M., Elkenany, R. M., & Elderieny,M. H. (2018). Detection of alpha toxin and enterotoxins of Clostridium perfringens isolated from minced meat by real time polymerase chain reaction (PCR). African Journal of Mi- crobiology Research, 12(23), 546-555.
- Kamber, U., Gokce, H. I., & Elmali, M. (2007). Clostridium perfringens and its toxins in minced meat from Kars, Turkey. Food additives and contaminants, 24(7), 673-678.
- Shaltout, F. A., Zakaria, I. M., & Nabil, M. E. (2017). De- tection and typing of Clostridium perfringens in some retail chicken meat products. BMVJ, 33, 283-291.
- Schalch, B., Sperner, B., Eisgruber, H., & Stolle, A. (1999). Molecular methods for the analysis of Clostridium perfrin- gens relevant to food hygiene. FEMS Immunology & Medical Microbiology, 24(3), 281-286.
- Bisi-Johnson, M. A., Obi, C. L., Vasaikar, S. D., Baba, K. A., & Hattori, T. (2011). Molecular basis of virulence in clinical isolates of Escherichia coli and Salmonella species from a ter-tiary hospital in the Eastern Cape, South Africa. Gut patho- gens, 3, 1-8.
- Dipineto, L., Santaniello, A., Fontanella, M., Lagos, K., Fio- retti, A., & Menna, L. F. (2006). Presence of Shiga toxinâ?ÂÂpro- ducing Escherichia coli O157: H7 in living layer hens. Letters in applied microbiology, 43(3), 293-295.
- Marrakchi, A. E., Bennour, M., Bouchriti, N., Hamama, A., & Tagafait, H. (1990). Sensory, chemical, and microbiological assessments of Moroccan sardines (Sardina pilchardus) stored in ice. Journal of food protection, 53(7), 600-605.
- Kaneko, I., Miyamoto, K., Mimura, K., Yumine, N., Utsu- nomiya, H., Akimoto, S., & McClane, B. A. (2011). Detection of enterotoxigenic Clostridium perfringens in meat samples by using molecular methods. Applied and Environmental Mi- crobiology, 77(21), 7526-7532.
- Mehrotra, M., Wang, G., & Johnson, W. M. (2000). Multiplex PCR for detection of genes for Staphylococcus aureus entero- toxins, exfoliative toxins, toxic shock syndrome toxin 1, and methicillin resistance. Journal of clinical microbiology, 38(3), 1032-1035.
- Murugkar, H. V., Rahman, H., & Dutta, P. K. (2003). Distribu- tion of virulence genes in Salmonella serovars isolated from man & animals. Indian Journal of Medical Research, 117, 66.
- Park, H. K., Woo, S. Y., Jung, Y. J., Lee, E. O., Cha, J. E.,Park, H. S., & Lee, S. J. (2008). Detection of virulence genes of Staphyloccus aureus and Staphylococcus epidermidis iso- lated from suprapubic urine from infants with fever. Journal of Bacteriology and Virology, 38(4), 189-196.
- Ranjbar, R., Masoudimanesh, M., Dehkordi, F. S., Jonaidi-Ja- fari, N., & Rahimi, E. (2017). Shiga (Vero)-toxin producing Escherichia coli isolated from the hospital foods; virulence factors, o-serogroups and antimicrobial resistance properties. Antimicrobial Resistance & Infection Control, 6, 1-11.
- The center for food security and public health (2004). “Epsi- lon Toxin of Clostridium perfringens”. Institute for interna- tional cooperation in animal biologics, Fac. Vet. Med. Iowa Univ. USA, pp. 1-4.
- Yoo, H. S., Lee, S. U., Park, K. Y., & Park, Y. H. (1997). Mo-lecular typing and epidemiological survey of prevalence of Clostridium perfringens types by multiplex PCR. Journal of clinical microbiology, 35(1), 228-232.


