Research Article - (2020) Volume 1, Issue 1
Evocation of DNA from the Blood Sample Mixed with Road Concrete- A Forensic Review
2Forensic Science Laboratory, Home Department, GNCT of Delhi, Rohini- Delhi, 110085, India
Received Date: Nov 11, 2020 / Accepted Date: Nov 18, 2020 / Published Date: Nov 23, 2020
Copyright: ©Copyright: �2020 Dr Amit Chauhan. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Several types of biological samples are recovered in different conditions in various types of crimes. Sometimes, due to mishandling of the evidence they do not produce good results. Blood is a common type of biological sample found at crime scene involving hitand-run, murder and various sexual assaults, etc. In the cases such as hit and run, the blood gets mixed up with the soil and act as resistance in obtaining proper DNA profiling of a victim or a suspect. The advanced technologies for DNA profiling like STR analysis help in the identification of a criminal even if there is not much quantity of biological sample from the crime scene. But non-scientific procedure of blood collection and preservation reduces the chances of amplification of DNA. Additionally, in many outdoor cases blood sample contaminated with soil becomes problematic because of the presence of humic acid in soil. Humic acid inhibits the amplification of DNA and leads to unsuccessful profiling of DNA. The inhibitors in the samples act as obstacle in the cases where blood is lifted from the surface of earth and lead to the unsuccessful DNA analysis. PCR artifacts like partial profiles, multi-peaks, or complete failure of DNA profile can be seen in STR profiles obtained from contaminated samples. In this study, we reviewed the blood samples recovered from different surfaces (wall of plaster, cemented floor pieces, black road concrete) over a period of 5 to 36 months. This study concludes that the sample containing less soil particles yield DNA higher than the samples containing high amount of soil particles.
Keywords
Extraction of DNA, Blood, Concrete, Plaster, Cement, Inhibitors, STR Profile etc
Introduction
Mainly the forensic scientists apply scientific and analytical techniques for the examination of evidences found from different sources such as victim, suspect, or crime scene itself. After analyzing they reach to the conclusion and prepare their legal statement in which they provide the summary of the results obtained from analysis. There are various crime cases such as murder, sexual assaults, hit and run, etc. where the body of the victim is found on the road-side and biological fluid evidences are mixed with road concrete containing cement, sand and gravels [1].
If we observe it legally, it is necessary that crime scene should be barricaded as soon as possible by the investigating agency. At the crime scene, where biological fluid evidences such as blood is found, DNA can be obtained and identification of a suspect or of the anonymous body found at the crime scene can be done which helps the Investigating Officers in their further Investigation [2]. Most of the time forensic experts are called upon to provide their support and to analyze the various biological samples and give their report which links the suspect or victim to the scene of crime. The analysis becomes challenging for the forensic scientist when the biological fluid evidences obtained from the crime scene gets contaminated or collected in unscientific manner due to improper handling as these are generally collected and preserved by the police personnel, present at the crime scene [3].
The biological fluid evidence such as blood or a pool of blood which is mixed with road concretes should be collected scientifically using sterile piece of cotton cloth or a gauze pad. The sample should be completely air dried before packaging it in a paper bag and then it should be kept in a labeled brown paper bag or a box. It should not be kept in a plastic bag as even the slightly moist soil present in a road concrete can cause the growth of microorganisms that will eventually destroy the evidence. In this case autolysis occurs which leads to destruction of cell by bacteria [4].
It is important to remove the contaminants from the biological fluid sample found at the road side mixed with concrete before processing the amplification of DNA as they might act as a PCR inhibitor. There should be proper amplicon concentration for the complete DNA profiling process otherwise the profile will contain false peaks, split peaks or off ladder alleles. The process of amplification gets obliterated if the inhibitors present in the sample binds themselves to the DNA [5]. The magnetic beads are used for analyzing the sample containing inhibitors. If we use the degraded sample directly the results of the DNA profile would not be complete or successful and the result can be false negative as it will affect the autosomal STR analysis. In such cases blotting paper can be used for soaking of blood and it can be used for DNA profiling.
Materials and Methods
Samples recovered from various surfaces were taken by the police officers or for forensic expert during the crime scene investigations that were preserved for laboratory examination. A total 45 samples were taken in which 15 included ‘blood from wall plaster’ other 15 included ‘blood from black road concrete’ and remaining 15 ‘blood-stained cemented floor pieces’. These were taken on cloth gauze and dried in an incubator for around 2 to 4 hours at 40°C [6-8]. The exhibit ‘blood from the wall plaster’ which was lifted by officers already from the spot, was directly taken for examination purposes. Organic extraction method was used for all the sample processing. Since, the soil is most common inhibitor; the first step was to remove the soil at the isolation stage. The column provided with the Qiagen kit was used to remove the soil particles from blood.
DNA Extraction and Quantitation
Phenol chloroform extraction method was applied for the isolation of DNA. The solvent used for separation of DNA from protein and cell debris was buffered tris-phenol. The gauze was prepared from ‘blood from wall plaster’ and from the forensic exhibit of ‘blood stain cemented floor pieces’ and ‘blood stained black road concrete’, blood stain gauze cloth pieces were prepared. The small pieces were cut and collected in 1.5 ml tubes. It is believed that if we use direct lysis method sheering of DNA will occur and impurities like humic acid will not be removed which brings the need of extra purification process but it might lead to less yield of DNA [8]. Samples having soil content were mixed with normal saline and the process of incubation was preceded for around 3 hours at 60°C. The solution of soil obtained was transferred to elution micro tubes containing filter membrane and column tube. The filtration of sample which was lysated in elution micro tube was done by the process of centrifugation at 10,000 rpm. The sample lysate was then moved in normal 1.5 ml tubes. To each tube containing a small gauze pieces and sample lysate 500 microliter forensic buffer, 40 microliter SDS 20% and 25 microliter PK was added for the process of incubation it was kept overnight at 37°C in thermo mixer. The next day, we used organic extraction method with phenol, chloroform and Isoamyl alcohol. For the precipitation of DNA, sodium acetate and isopropanol was used leading to formation of pellet. With 70% alcohol the DNA pellet was washed twice and for the last time by 100% alcohol [8]. Then, DNA was dissolved in 40 microliter TE. After the process of isolation, the determination of DNA concentration in each sample was done using Quantifiler Duo Quantification Kit (Applied Bio system) with 7500 Real Time PCR machine in accordance with the protocols provided by the manufacturer [6]. The quantity of DNA was determined in each sample using quantification standards. The kit used for the process contains three components- Taq Man, SRY gene and IPC (International position control).
Results & Discussion
By an implementation of 7500 Real Time PCR machine, the DNA quantity obtained from different samples is shown in the table number 1. Percentage of success rate can be observed from the data. It was a potential challenge to examine all the samples. The quantity of DNA extracted was highest in case of ‘blood stained from cemented floor piece’ (94.90 g/ microliter) and least in case of ‘blood from the wall plaster’ (0.16 g/ microliter) [7]. It was noticeable that the peak height in the profiles is affected by the quality and quantity of DNA. Complete balanced STR profiles with the absence of any PCR artifacts were produced by the samples from ‘blood stain cemented floor piece’, ‘blood from wall plaster’ and ‘blood stain black road concrete’. In isolating high-quality genomic DNA, removal of inhibitors was effective (with some exceptions) as shown by the peak heights obtained from STR profiles as they were equivalent to the input amount [9, 10]. It was noted that the sample containing less soil particle yield DNA higher than the samples containing high amount of soil particles.
Comparative chart showing total DNA yield in ÃÃÂ???g/µl obtained from different samples.
Table: Quantity of DNA obtained from the blood collected from various surfaces.
|
S. No. |
Blood from The Wall Plaster |
Blood Stained Black Road Concrete |
Blood Stained Cemented Floor Pieces |
|
(DNA in ng./ µl.) |
(DNA in ng./ µl.) |
(DNA in ng./ µl.) |
|
|
1 |
0.00 |
1.92 |
16.11 |
|
2 |
0.00 |
2.68 |
7.63 |
|
3 |
2.69 |
2.18 |
0.24 |
|
4 |
0.00 |
9.32 |
33.9 |
|
5 |
0.00 |
6.07 |
14.94 |
|
6 |
1.03 |
0.16 |
1.83 |
|
7 |
0.96 |
1.34 |
4.01 |
|
8 |
0.94 |
6.52 |
23.95 |
|
9 |
0.86 |
0.00 |
14.07 |
|
10 |
0.00 |
3.68 |
1.69 |
|
11 |
1.11 |
2.19 |
8.08 |
|
12 |
0.00 |
4.20 |
0.00 |
|
13 |
0.16 |
1.06 |
33.32 |
|
14 |
0.00 |
3.87 |
0.94 |
|
15 |
1.29 |
0.00 |
2.19 |
|
Avg. |
0.60 |
3.01 |
10.86 |
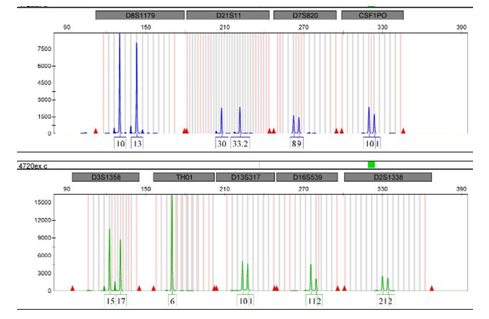
Graph 1: DNA profile from blood taken from smooth earthy surfaces.
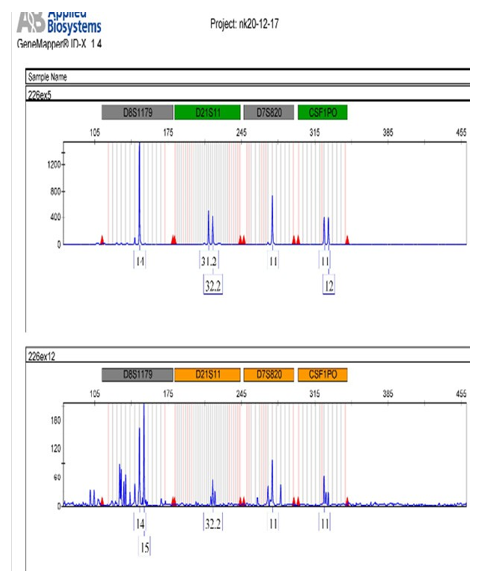
Graph 2: Showing the comparative profile of four markers of blue dye (FAM) lower sample is degraded.
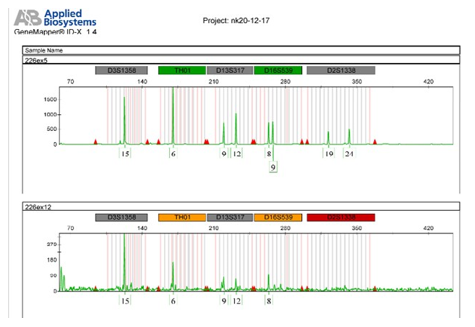
Graph 3: Showing the comparative profile of four markers of green dye (VIC) lower profile is affected by the soil inhibition while upper one is accurate.
Conclusion
Sometimes, it is challenging for the forensic scientists to extract DNA from samples mixed with the soil and road concrete because such samples contain contaminants and PCR inhibitors and there is variation observed in STR profiles and yield of DNA. The sample containing higher soil contaminants produces multiple peaks on low size markers less than 200 base pairs and no peaks on big size markers8. Exhibits having higher content of soil and degradation show allelic dropout and incomplete or partial profiles. Lastly, it can be concluded that some of the sample generated complete profiles, remaining were partial or incomplete because of the presence of soil in the sample. To obtain this, the sample should contain minimum number of contaminants and inhibitors and if the sample is damply preserved, there is reduction in chances of amplification of DNA.
References
- Kastle J, St Roberts N (1909) Tests for pus and blood. From the Division of Chemistry, Hygieni Laboratory, U.S. Public Health and Marine Hospital Service, Washington. Journal of Biological Chemistry 6: 650-652.
- Thornton JI (1997) The general assumptions and rationale of forensic identification. In: Modern Scientific Evidence: The Law and Science of Expert Testimony. West Publishing Co., St. Paul 2
- WG Wang, P Kumar, M Schwarz, G Malone, A Haworth, et al. (1994) P C R Amplification of 40-year paraffin-embedded Tumour-tissues-comparison of 4 different DNA extraction and purification methods. Int J Oncol 5: 453-457.
- Shahzad MS (2009) Effect of blood-stained soils and Time period on DNA and allele drop out using Promega 16Powerplex Kit. Forensic Science International Genetics Supplement Series 2: 161-162.x
- JM Butler (2005) Forensic DNA typing, biology, Technology and genetics of STR markers. 2nd Ed. New York: Elsvier Acedemic Press.
- YL Tsai BHO (1992) Detection of low numbers of Bacterial cells in soils & sediments by Polymerase Chain Reaction. Appl Environ Microbiol 58: 754-757.
- YL Tsai BHO (1992) Rapid method for separation of Bacterial DNA from humic substances in Sediments for polymerase chain reaction. Appl Environ Microbiol 58: 2292-2295.
- Naresh Kumar (2017) Effect of inhibitor on the blood taken from earthy surfaces and DNA profiling in forensic cases. Journal of Punjab academy of Forensic Medicine and toxicology 17: 46-52.
- C Ladd, MT Bourke, CA Scherczinger, EM Pagliaro, RE Gaensslen, et al. (1996) A PCR-based strategy for ABO genotype determination. Forensic Sci 41: 134-137.
- Gilmore TE, Howe C (1959) An Aerobic Soil Microorganism Which Decomposes Blood Group Substance Cell Free Extracts on Blood Group Substances. J Bacteriol 78: 814-820.


