Research Article - (2023) Volume 4, Issue 1
DNA Profiling of Dry Umbilical Cord Tissue Unfolds the Mystery of Disowned Dead Female Infant
Received Date: Oct 14, 2023 / Accepted Date: Nov 03, 2023 / Published Date: Nov 15, 2023
Copyright: ©©2023 S.R. Khairkar, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Shah, B.T., Parulekar, V.B. Khairkar, S.R., Shinde S.A., Ghumatkar, S.V. (2023). DNA profiling of dry umbilical cord tissue unfolds the mystery of disowned dead female infant. In J Fore Res. 4(1), 188-191.
Abstract
DNA profiling or DNA fingerprinting, has revolutionized forensic science and greatly enhanced the ability of law enforcement to match perpetrators with crime scenes. DNA typing has become an invaluable tool in criminal investigations and has led to the resolution of numerous cases. Forensic DNA profiling plays an important role in paternity testing. Using this advanced vital tool, one can provide accurate scientific evidence of the biological parents. In present case, DNA profiling proved to be an inevitable tool to establish the paternity of a piece of small dry umbilical cord tissue of disowned female infant and helps to reach the real culprit and hence the justice prevails in this case.
Keywords
DNA Profiling, Disowned Female Infant, Umbilical Cord Tissue, Paternity
Introduction
One of the most significant advancements in DNA profiling came in the late 1980s with the introduction of the polymerase chain reaction (PCR) technique. PCR enabled scientists to amplify small quantities of DNA, making it possible to generate profiles from even trace amounts of biological material, such as a single hair or a drop of blood.
The identification of a newborn's genetic profile can have various applications. One important use is in verifying the identity of the baby, which can be crucial for legal and medical purposes, such as ensuring the right baby is matched to the right mother in hospitals or confirming parentage in cases of legal disputes.
Furthermore, as DNA technology advances, there is the potential for umbilical cord blood to play an even more significant role in healthcare. Genetic and metabolic screening can be performed on cord blood samples to identify potential disorders and conditions that the infant might be at risk for. This early identification can allow healthcare professionals to initiate treatment or interventions before the onset of symptoms, potentially improving outcomes and quality of life for the child.
As the technology improved, more STR markers were identified, allowing for even greater discrimination between individuals. Fo- rensic DNA databases were established to store DNA profiles from convicted offenders, providing a valuable resource for matching crime scene evidence to known individuals.
The accuracy and reliability of DNA profiling have led to the suc- cessful resolution of countless criminal cases. It has been instrumen- tal in exonerating wrongfully convicted individuals and identifying perpetrators who might have otherwise gone undetected. DNA evi- dence has become a powerful tool in the courtroom, often serving as a definitive link between a suspect and a crime scene [1].
Over the years, advancements in DNA technology, such as the in- troduction of short tandem repeat analysis and the development of automated DNA sequencing methods, have further improved the accuracy and efficiency of DNA profiling. Today, DNA typing is a standard procedure in forensic investigations worldwide. Its impact on law enforcement and the criminal justice system cannot be over- stated. The ability to link perpetrators to crime scenes with a high degree of accuracy has not only helped bring justice to victims but has also acted as a powerful deterrent against future crimes.
DNA fingerprinting, or forensic DNA profiling, has indeed revolu- tionized forensic investigations. It is widely used in various medi- co-legal cases to establish identity, determine parentage, and solve crimes. The technique compares the DNA in a person's cells with DNA found at a crime scene or with the DNA of other individuals to establish identification or exclusion. The applications of DNA fingerprinting in forensic science are numerous. Some of the ma- jor applications include: Paternity and Maternity Disputes: DNA testing can determine biological relationships between individuals, such as establishing paternity or maternity in legal cases.
DNA fingerprinting relies on specific genetic markers called short tandem repeats (STRs). STR loci are regions of DNA where short sequences of nucleotides are repeated. The choice of STR loci for forensic analysis is based on factors such as discrete and distin- guishable alleles, robust amplification, high power of discrimina- tion, absence of genetic linkage with other loci, low artifact forma- tion during amplification, and the ability to be used in multiplex PCR (Polymerase Chain Reaction) [2, 3].
Case History
Early morning on 04/08/2022 at around 7.30 am a Pantnagar po- lice station received phone call from unknown person that a new born infant baby is lying in an unconscious state near a building. Upon reaching the crime scene, they found one day old female child naked and unconscious. Near that female infant dry umbili- cal cord tissue pieces mixed with garbage and mud was also lying.
Upon medical examination, the female infant was declared dead. The sternum and dry umbilical cord tissue pieces from crime scene were sent to FSL, Mumbai for DNA examination along with sus- pected mother and father’s blood.
Herein, we attempted extraction and quantification of sternum bone did not yield sufficient DNA to generate DNA profile. So the only source of DNA was the umbilical cord tissue of infant. As the sample was mixed with garbage and mud, it became very difficult to extract the DNA. We have optimized the above said DNA extraction protocol for such samples; it gives good quality and quantity of amplifiable DNA.
Experimental
Materials and Methods
The forensic community has selected STR loci to incorporate into multiplex reactions based on several features including 1) Discrete and distinguishable alleles 2) amplification of the locus should be robust 3) a high power of discrimination 4) an absence of genet- ic linkage with other loci being analysed 5) low levels of artifact formation during the amplification 6) the ability to be amplified as part of a multiplex PCR. An essential feature of any STR used in forensic analysis is that biological material should give an iden- tical profile regardless of the individual or laboratory that carries out the analysis [4, 5].
The AmpFISTR Identifiler analyzes 15 STR along with amelogenin locus. This kit is used widely worldwide particularly for kinship testing. The Amp FISTR Identifiler kit is one of the widely used kits for DNA profiling. It analyzes 15 STR loci, along with the amelogenin locus, which determines the sex of an individual. It's important to note that advancements in DNA profiling techniques and the use of DNA databases have significantly improved the accuracy and reliability of forensic investigations, leading to the identification of culprits and the exoneration of innocent individuals. DNA finger- printing has become an invaluable tool in modern forensic science.
1) Investigator kit (Qiagen)
2) Identifiler kit (Applied Biosystems)
3) Lysis buffer
4) 1M DTT
5) Proteinase K
6) PBS buffer
A) DNA extraction from sternum bone:
50 mg of sternum bone piece taken in an eppendorf tube to which add (400ul) of Forensic buffer, (40ul) 1mM DTT, (20ul) of Pro- teinase K. Vortex the contents and short spin. Incubate tube at 560C with shaking overnight. DNA extraction is carried out with EZ1 Advanced (Qiagen) magnetic bead based liquid handling system.
B) DNA extraction from piece of dry umbilical cord tissue
:- 50 mg of umbilical cord tissue taken in an eppendorf tube to which add (1 mL) of PBS. Keep it for 30 mins. Drain the PBS and add 1 ml D/W. After washing and cleaning, drain the D/W. Add (400ul) Forensic buffer, (25ul) of Proteinase K. Vortex the contents and short spin. Incubate tube at 560 C with shaking overnight. Next day kept it at 950 C for 5 minutes and then DNA extraction was car- ried out with EZ1 Advanced (Qiagen) magnetic bead based liquid handling system.
C) DNA extraction of all blood samples is carried out using EZ1 Advanced (Qiagen) magnetic bead based liquid handling system.
Extracted DNA from the above samples were quantified using In- vestigator Quantiplex pro kit and then amplified at 15 STR loci by using identifiler PCR amplification kit. Amplified PCR products were genotyped on 3500-XL Genetic analyzer.
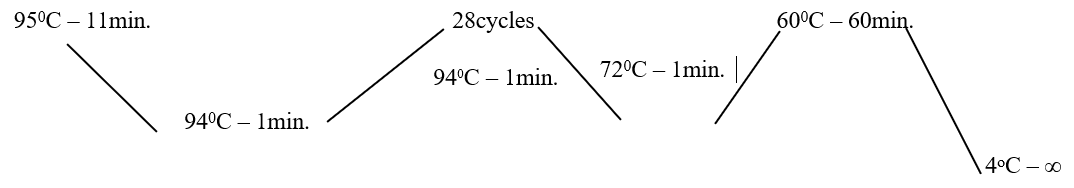
PCR PROTOCOL:- STR genotyping was carried out using the AmpFISTR identifiler PCR Amplification kit (Applied Biosyste- ms, Foster City)
AmpFISTR PCR reaction mix: 10.5ul
AmpliTaq Gold DNA polymerase: 0.5ul
AmpFISTR Primer set: 5.5ul
DNA Sample: 10ul
Genotyping
STR genotyping is detected and analysed on 3500-XL Genetic An- alyzer (Applied Biosystems) instrument by capillary electrophore- sis of single stranded amplified DNA fragments.
Results of Analysis
The DNA of extracted samples was typed at 15 STR LOCI and gender specific Amelogenin locus using PCR Amplification tech- nique.
The results of DNA typing are summarized as follows:-
|
STR LOCUS |
GENOTYPE |
||
|
DNA OF BLOOD OF SUSPECTED MOTHER |
DNA OF PIECE OF DRY UMBILI- CAL CORD TISSUE |
DNA OF BLOOD OF SUSPECTED FATHER |
|
|
D8S1179 |
10,15 |
10,15,16 |
10,16 |
|
D21S11 |
30,30 |
30,32.2 |
30,33.2 |
|
D7S820 |
10,11 |
10, 11,12 |
11,12 |
|
CSF1PO |
11,11 |
10,11 |
12,13 |
|
D3S1358 |
15,18 |
15,16,18 |
15,16 |
|
THO1 |
6,6 |
6,6 |
9,9.3 |
|
D13S317 |
9,11 |
9,10,11 |
8,12 |
|
D16S539 |
9,12 |
9,12,13 |
8,9 |
|
D2S1338 |
21,22 |
16,21,22 |
18,18 |
|
D19S433 |
13,15 |
12,13,15 |
13,15 |
|
vWA |
15,16 |
15,16,17 |
14,16 |
|
TPOX |
11,11 |
9,11 |
9,11 |
|
D18S51 |
14,17 |
14,17,18 |
14,19 |
|
AMELOGENIN |
X,X |
X,X |
X,Y |
|
D5S818 |
12,12 |
12,12 |
10,11 |
|
FGA |
19,20 |
19,20,22 |
19,19 |
Observation:
1) No amplifiable DNA is obtained from sternum bone of female infant.
2) Piece of dry umbilical cord tissue is a mixture of umbilical cord of infant and some maternal blood from suspected mother.
Interpretation:
- Interpretation of STR genotyping involves comparing the genotypes of different individuals to determine their relatedness or to establish individual identity. Here are some key points in the interpretation process:
Allele Calling: The first step is to identify the specific alleles (number of repeats) present at each STR locus for an individual. This is done by comparing the sizes of the DNA fragments obtained from the electrophoresis analysis to a known set of reference alleles.
Genotype Comparison: Once the alleles are identified, the genotypes of different individuals are compared. A genotype refers to the combination of alleles observed at multiple STR loci for an individual. By comparing the genotypes, one can determine if two individuals share the same alleles at multiple loci, suggesting a potential familial relationship or individual identity.
Statistical Analysis: Statistical calculations are performed to determine the frequency of observed genotypes in the population. This information is used to estimate the rarity of a particular genotype and calculate the likelihood of randomly encountering the same genotype in unrelated individuals.
DNA Profiling: In forensic applications, STR genotyping is used for DNA profiling, where an individual's genotype is compared to a database of known profiles to identify or exclude potential matches. The more loci that are analyzed, the higher the discriminatory power of the DNA profile.
It's important to note that STR genotyping provides a statistical estimate of the likelihood of a particular genotype occurring in a population, rather than absolute certainty. The interpretation of the results requires expertise in forensic genetics and should be done by qualified professionals.
Collection of DNA Samples: DNA samples are collected from the child and the alleged parents. The samples can be obtained through non-invasive methods such as buccal swabs or by using other biological samples like blood or saliva.
Selection of STR Loci: Specific STR loci are chosen for analysis. These loci are highly variable among individuals, making them ideal for determining parentage. The commonly used loci include those recommended by forensic DNA databases, such as the Combined DNA Index System (CODIS).
1) For all the 15 different genetic systems analyzed with the PCR, suspected mother matched the obligate maternal alleles present in piece of dry umbilical cord tissue at all loci.
2) Out of 15 different genetic systems analyzed with the PCR, suspected father did not match the obligate paternal alleles present in piece of dry umbilical cord tissue at 11 loci.
Analytical finding-
1) Suspected mother is concluded to be the biological mother of piece of dry umbilical cord tissue.
2) Suspected father is excluded to be the biological father of piece of dry umbilical cord tissue.
Discussion
The collected DNA samples are subjected to STR genotyping, where the targeted loci are amplified using PCR and analyzed using techniques like capillary electrophoresis. The resulting genotypes for each individual are determined by identifying the number of repeats at each locus.
Genotype Comparison: The genotypes of the child and the alleged parents are compared to assess the genetic similarities. In parentage cases, the focus is on finding shared alleles between the child and the biological parents.
It's important to note that DNA profiling for parentage determination is highly accurate when properly conducted by accredited laboratories using recognized protocols and quality control measures. The power of DNA profiling lies in its ability to detect genetic similarities and exclusions, providing reliable evidence for establishing or confirming biological parentage.
Conclusion
In the future, umbilical cord blood may be used for genetic and metabolic screening to identify disorders and begin treatment prior to the onset of symptoms. As DNA technology advances, forensic professionals can help ensure that even the tiniest patients benefit from ongoing research.
In present case, repeated extraction and quantification of sternum bone did not yield sufficient DNA to generate DNA profile. So the only source of DNA was the umbilical cord tissue of infant. As the sample was mixed with garbage and mud, it became very difficult to extract the DNA. We have optimized the above said DNA extraction protocol for such samples; it gives good quality and quantity of amplifiable DNA.
References
- Roewer, L. (2013). DNA fingerprinting in forensics: past, present, future. Investigative genetics, 4(1), 1-10.
- Scientific working Group on DNA Analysis Method (SWGDAM) (200) short tandem repeats (STR) interpretation guidelines. Forensic Science Communication 2(3).
- AJS BHANWER*, ARVIND RUP SINGH** and JAI RUPSINGH. DNA Fingerprinting in forensic and medico-legal applications, JPAFMAT, 2001, Vol.: 1.
- Scientific working Group on DNA Analysis Method (SWGDAM) (200) short tandem repeats (STR) interpretation guidelines. Forensic Science Communication 2(3).
- Reynolds, R., Sensabaugh, G., & Blake, E. (1991). Analysis of genetic markers in forensic DNA samples using the polymerase chain reaction. Analytical chemistry, 63(1), 2-15.


