Research Article - (2023) Volume 6, Issue 1
Cytosine Quadruplexes in Duplex DNA may be Non-transient and have Epigenetic Relevance
Received Date: Sep 05, 2023 / Accepted Date: Sep 24, 2023 / Published Date: Oct 03, 2023
Copyright: ©Â©2023 Simon Chapman. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Chapman, S., (2023). Cytosine Quadruplexes in Duplex DNA may be Non-transient and have Epigenetic Relevance. Toxi App Pharma Insights, 6(1), 37-53.
Abstract
Guanine quadruplexes folding in single-strand DNA have been extensively studied for some time. But far less attention has been paid to systems arising in duplex DNA. Furthermore, the literature shows an apparent absence of quadruplex systems generated exclusively by other nucleic acid bases. This study investigates biplanar and triplanar complexes folding in double-strand DNA when derived from cytosine (C4s). The role of various co-ordinating cations was also investigated. Molecular dynamics simulations determined triplanar models as relatively fragile systems, but all biplanar systems were found to be stable into biologically relevant times. Cationic co-ordination in biplanar systems appears to be much less contributive to stability than in triplanar equivalents. Uniquely biplanar or triplanar C4s have ostensibly not been previously described. It is suggested here that these structures folding in the human genome may have physiological relevance even if relatively short-lived. A non-transient folding / unfolding cycle might persist and influence epigenetic processes.
Keywords
DNA Quadruplex; Cytosine; Computational Chemistry and Molecular Modelling; Molecular Dynamics.
Introduction
A survey and analysis of guanine quadruplexes (G4s) in the litera- ture shows very little consideration of the complementary strand of duplex DNA. The human duplex telomeric sequence d (TTAGGG) n is necessarily rich in cytosine having the complementary profile of d (AATCCC)n. An in vitro supramolecular structure predicated on protonated C-C base pairing was initially reported some time ago [1]. The stability of this conformation, characterised as an i-motif, is critically contingent on local pH [2, 3]. This particular supramolecular structure is especially prevalent in cytosine-rich sequences [4]. The structure is more stable in pH conditions below physiological levels and is consolidated by π-π stacking (figure 1)
Figure 1: for pH values < 7.0, cytosines can be protonated to associate into a dimer.
(A): The paired bases share three H-bonds. (B): Dimers can stack vertically with consecutive planes rotated through 900. (C): Alternate planes (light green and dark green) are linked by the phosphate-sugar backbone (light green and dark green vertical lines).
The cytosine dimer system may have other topologies [5]. The low values for local pH of the in vitro models discussed in the earli- est papers would seem to preclude an in vivo telomeric cytosine i-motif. However, recent research concludes the structure may be relatively stable at levels close to physiological pH [6]. Moreover, the sequence required for potentially forming a cytosine i-motif appears ubiquitous in the human genome and particularly in the centromeric environment [7].
G4s have been promising targets for anti-cancer intervention [8]. But paradoxically, some complexes have been linked inter alia to neurodegeneration [9, 10]. It is suggested here that anti-sense cy- tosines may also self-associate into a quadruplex structure. Either uniquely, or in combination with guanine, these may compromise sense-strand beneficial G4 activity. Bio-informatics identify G4s populate specific sites such as immunoglobulin switch regions, promoter regions and recombination sites [11, 12]. A gene reg- ulatory role has been proposed for G4s adjacent to transcription start sites [13]. Exploring the possibility of cytosine quadruplex- es located complementary to beneficial guanine equivalents with such crucial roles, might underpin some aspects of the paradoxical pathogenicity. The proposal that anti-sense G4s may also fold ap- pears not to have been investigated. Accounting for these propos- als in the C-rich anti-sense strand suggested an initial assessment of cytosine dimerisation for this study.
Cytosine dimers
A series of cytosine-cytosine dimers cytosine were designed and are shown in figure 2. A pre-requisite for a feasible structure to incorporate into a nucleic acid strand was cytosine-N1 availability (circled) for covalent bonding to the phosphate backbone
Figure 2: cytosine dimers. (a), (b) cytosine-cytosine (c) dimer co-ordinated by metal cation, M+ (d) cytosine, protonated dimer.
Structure 2d was considered feasible based on the early research into the i-motif structure. However, the caveat was noted that protonated cytosine is unlikely at physiological pH. Figure 2a ap- peared viable in principle and showed some commonality with a structure recently designed in silico on a graphene surface [14]. However, the Graphene paper discusses the relevance of dimeri- sation to nanoparticle technology and does not extend to raising implications for a biological context. Amino hydrogens of one paired dimer have recently been suggested as capable of H-bond- ing to carbonyl oxygen of the other [15]. Further structures were developed from the structure described in figure 2a to explore the possibility of forming a cytosine quartet. Both cytosines of the model were marginally re-aligned. This manipulation suggested a dimer was possible by H-bonding between H atoms on N4 of one cytosine with O2 and N3 of the other (figure 2b). Amino hydro- gens of one paired dimer have recently been suggested as capable of H-bonding to carbonyl oxygen of the other [16, 17]. Proposed here is the formation of a cytosine quartet where two such re-ori- entated dimers might further H-bond (figure 3). Significantly, N1 hydrogens are available in this model for covalent bonding to the DNA chain
Figure 3: possible alignment of a cytosine dimer pair (from figure 2b). Both molecules are planar and the arrangement appears viable
except that carbonyl oxygens have unfavourable angles for H-bonding with opposing amino groups.
The bonding angle O2-H1-N4 between associating cytosines is far from 180o. Within a small tolerance, this value is required for orthodox H-bonds and to be sustainable, this point should be ad- dressed [18].
Cytosine quartets
Cytosine quartet co-ordinated with non-physiologically relevant ions: Ba2+ A hybrid guanine-cytosine quartet may fold in C-rich DNA and stack vertically between guanine quartets [19]. The paper does not describe the molecular structure of a cytosine quartet and has no proposal of progression to a quadruplex. A cytosine quartet has been identified in a guanine quadruplex co-ordinated with Ba2+ mediated by water molecules (figure 4) [20]. Although the struc- ture is planar and appears well-served by H-bonds, barium is toxic in human physiology [21].
Figure 4: Water molecules mediating Ba++ co-ordinating a cytosine quartet. Adapted from Zhang, 2014. Four self-associating cytosines, mediated by water molecules and co-ordinated with Ba2+. Cytosine N1 of all four bases appears too available for covalent bonding ex- ternal to the quartet.
Cytosine quartets have previously only been observed in vitro DNA solutions [22]. Protonation of cytosine N3 and the conse- quent delocalised positive charge on the quartet were suggested as a mechanism for interaction with a cation mediated by a molecule of water [22]. Distances between cytosines were too large to co-or- dinate a cation directly, hence waters were required to mediate the electrostatic bonding. Cytosine quartets co-ordinated with non-physiologically relevant ions: Pb2+, Tl+, Rb+, Hg2+ Quartets have recently been identified in the crystal structure of a DNA quadruplex comprising tetrads of various base combinations including cytosine-thymine pairs [23, 24]. However, a majority of these complexes are co-ordinated by non-physiologically relevant ions such as Pb2+, Tl+, Rb+ Hg2+ [25, 26]. The NMR structure of a non-hybrid uniquely cytosine quartet located between guanine quartets has been reported [27]. The folding sequence derives from the Simian Virus 40 genome. No co-ordinating cation is noted, but there appears to be sufficient intramolecular space in the central channel to accommodate ions mediated by waters. The model dis- plays single H-bonds between each cytosine O2 and adjacent ami- no-H, a total of four H-bonds. Cytosine quartets co-ordinated with physiologically relevant ions: Li+, Na+ or K+ A cytosine quartet derived from two re-orientated dimers (figure 2c: cytosine III), might further H-bond. Significantly, such a struc- ture retains N1 hydrogen availability for non-covalent deoxyribo- phosphate bonding. Each cytosine has four H-bonds to adjacent molecules suggesting a degree of stability. In contrast, the cytosine quartet constructed on a graphene surface can only be assembled if one of the cytosines is inverted (figure 5, circled in orange) [14]. Moreover, the complex does not lend itself to linkage with a DNA strand since one cytosine N1 (circled in red), is not available.
Figure 5: Potential cytosine quartet formed on graphene, adapted from Saikia (2018).
Given these constraints, the structure was rejected as invalid. Accordingly, focus returned to the paired dimer in figure 3(B). This model was slightly re-aligned to allow favourable N-H…O bonding between carbonyl oxygens and amino hydrogens (figure 6):
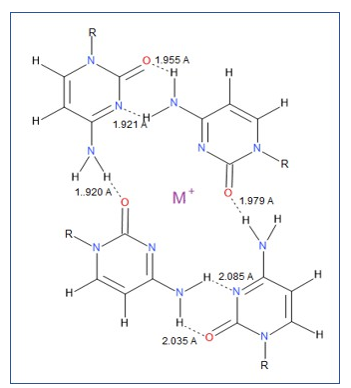
Figure 6: Re-aligned paired cytosine dimers to allow H-bond angles (O2-H1-N4) converging on 180o. The electronegative central space is sufficient to accommodate a metal cation (M+). Distances between ion and each cytosine-O2 = 3.381 Å; between M+ and each cytosine-N3 = 3.683 Å. Six H-bonds are predicted for this structure.
Overall, these observations mitigate the plausibility of cytosine quartets outlined in figure 6 stacking vertically into a quadruplex. The system is predicted to be planar and has N1 hydrogen avail- ability for non-covalent bonding to loops in a phosphate chain. There is also sufficient interplanar space for cationic co-ordination.
Guanine-cytosine hybrid quartets
Cytosine in combination with guanine may associate into a qua- druplex structure. Antisense cytosine-rich regions have recently been suggested as allowing a GCGC quartet to fold between a pair of guanine quartets [20]. The possibility of guanine-cytosine combinations in quadruplexes was proposed as underpinning some forms of neurodegeneration , and also in the regulatory region of the gene PLEKHG3 strongly associated with autism [28, 29]. Some epigenetic processes might be affected by such a guanine-cyto- sine-guanine quadruplex [20]. Developing a G-C-G-C quartet to a C-C-C-C structure is a rational next step. Such a cytosine quartet located between two guanine quartets could derive from: 5’…d (TT) AGCGTTAGCGTTAGCGTTAGCG…3’ Which is characteristic of the single strand telomeric sequence folding into a G4, but with four G-C substitutions. Other guanines could also undergo G-C substitution. Although the sequence has loops comprising three bases, it is as yet unclear if a maximum number has been validated. Selection pressures have favoured short loops as stable and optimised folding characteristics [30]. However, investigation of G-C-G hybrids is currently the subject of a separate study here and results will be published in due course
Cytosine quadruplexes
Cytosine dimers pairing into a quartet, then three such quartets stacking into a cytosine quadruplex (C4) is a logical sequential progression. As noted in a recent study, there are conflicting ob- servations regarding quadruplex stability with or without co-or- dinating ions [31]. For example, the stability of G4s co-ordinated with Li+ has been regarded as contentious [23, 25]. A triplanar guanine quadruplex acted as a scaffold for substitution by cytosine (described in Methods). Since this system is co-ordinated by K+, this cation, together with Na+ and Li+, were included in models for MD simulations. The general model for triplanar sytems is ex- emplified in figure 7:
Figure 7: Cytosine substituted for all guanine. The 22-mer guanine quadruplex 1KF1 folds from the sequence AGGG (TTAGGG)3, hence the cytosine structure has ACCC(TTACCC)3. Quartets derive from x2-x8-x14-x20, x3-x9-x15-x21 and x4-x10-x16-x22 where x= cytosine. Biplanar models comply with the same pattern except quartets derive from x2-x8-x14-x20 and x3-x9-x15-x21 only.
Methods
1KF1 was accessed from the Protein Data Bank (PDB) and adopt- ed as a scaffold for triplanar models. (PDB) 2KF8 was similarly accessed for biplanar systems; this system has no cationic co-or- dination. Cytosine was substituted for all guanines in the quartets but loops remained unaltered (refer to figure 1.2). Nine models were constructed for triplanar and biplanar quadruplexes in UCSF Chimera 1.12 (build_41623) by vertically stacking cytosine quar- tets linked by AAT loops using the swapna command. Triplanar quadruplexes are referred to as CDN12 and biplanar as CDN8. CDN12 and CDN8 models were co-ordinated by K+, Na+ or Li+ but the latter also with no ionic co-ordination.
Triplanar models retained K+ ions co-ordination by virtue of their 1KF1 derivation. The ions were replaced in turn with Li+ or Na+ by manual reformat of the .psf and .pdb files. Since 2KF8 has no co-ordinating cations, the derived cytosine model endorsed this factor. However, for comparison with biplanar guanine equiva- lents, K+, Na+ or Li+ ions were imposed on biplanar cytosine mod- els by the same method. Octagonal co-ordination was noted for K+ and Na+ systems. Models were transferred from Chimera to Dis- covery Studio Visualiser (Client) v16.1.0.15350 (DSV) for con- firmation of structural changes and to note modified residue num- bers. Monitoring triplanar models for K+ or Na+ systems confirmed co-ordinating ions have octagonal co-ordination. This observation is in close agreement with a 2016 review paper [32]. However, Li+ models adopted a tetrahedral geometry. Models showed π-H- bond donor interactions between amino nitrogens and the hetero- cyclic aromatic ring of cytosine, resonating with in silico cytosine π-stacking interactions noted in earlier research [33].
Molecular dynamics protocols.
Scripts were drafted for Molecular Dynamics (MD) analyses of models for simulation times of 500ns. Several Linux servers were accessed: 256-core allocation at the ARCHER2 National Super- computer, PowerEdge T640 96-core, 16-core and 8-core personal Desktops. All servers were supported by Linux OS. Data were col- lated with a Microsoft OS 8-core personal Desktop. Nano Molec- ular Dynamics NAMD_2.13_Linux-x86_64-multicore (NAMD) and Visual Molecular Dynamics (VMD 1.9.3) were the primary research tools, supported by DSV.
The water model used was AMBER TIP3P with an explicit solvent [34]. PBCs were applied and systems were neutralised by addition of an appropriate number of Na+ counter-ions via VMD Auto-ion- ize Plug-in v.1.5. A median salt concentration of 150mM was im- plemented using the Tool Command Language (Tcl) interface of NAMD to reflect physiological conditions and pH retained at the default setting of 7.0. Periodic boundary conditions (PBCs) were applied.
Systems were minimised by 50 000 x 0.01 Å steps of steepest de- scent and 5 000 steps of equilibration. Initial and target tempera- tures were 0.0 K and 310 K respectively. The default setting of zero was used for the pressure variable. The Langevin algorithm was used as the barostat. A constant volume simulation (NVT) for 100 ps was run at 310 K with the barostat off to thermalise the system. To provide good equilibration, the models were then run in a constant pressure simulation (NPT) for 100 ps at 310 K with the barostat on. Initial and target temperatures for equilibration were 0 K and 310 K respectively, the latter value more accurately reflecting a physiological environment. The default setting of zero was used for the pressure variable. AMBER force field ffSB14 was applied to the standard residues. The Particle Mesh Ewald summa- tion with a cut-off set at 8 Å was used to compute wider electro- static effects. The CHARMM27 force field was applied with 5 000 steps of equilibration.
Parameters for Periodic Boundary Conditions were calculated at (49.48 0.0 0.0), (0.0 44.79 0.0) and (0.0 0.0 54.75) for the three spatial co-ordinates, and models centred at (0.0 -0.86 0.66). All co-ordinates are in Ångstrom units.
Analytical metrics were: Root Mean Square Deviation (RMSD), ellipsoidal Root Mean Square Deviation (eRMSD), Radius of Gyration (Rg) and frequency of Hydrogen-bond formation (HB- form).
ERMSD uniquely analyses the geometry of nucleobases in a mol- ecule and evaluates differences in their relative interaction. Mea- surement is continuous, symmetric and provides a more detailed analysis of MD trajectories than RMSD. Units are Ångstrom. Molecular modelling and in silico analysis of RNA is generally contingent on only a few intrinsic features: backbone dihedrals, hydrogen-bond networks, and stacking interactions [35, 36]. Ra- dius of gyration parameterises configuration for an entire system at equilibrium and consequently determines change in molecular structure during MD simulations. It is calculated in NAMD as the sum of the root mean square distances of an atomic set from the molecular centre of mass (Eq.1.2)
<img src="https://www.opastpublishers.com/scholarly-images/6266-698ad4f775a90-cytosine-quadruplexes-in-duplex-dna-may-be-nontransient-and-.png" width="500" height="90">
Where N defines the atom set, mi is the mass of the atom i, ri is the distance of atom i from the molecular centre of mass [37]. Analy- sis of radius of gyration reinforces data from RMSD/eRMSD by quantifying fluctuations in Cartesian dimensions of the molecule. In the course of MD simulation, variations in molecular compact- ness are rendered. This parameter describes the ratio of molecular surface area to that of a perfect sphere with comparable volume. Higher values indicate elevated dynamic activity at a given point in the simulation and implicit less stability. Correspondingly, low- er values suggest greater molecular integrity at a given point. The calculation is weighted for mass but not charge and hydrogens are excluded. Units are Ångstrom.
The morphology and function of biomolecules is partially de- termined by hydrogen-bonding. Although an individual bond is empirically weaker than covalent or ionic bonding, a network of H-bonds collectively influences molecular integrity. Accordingly, interplanar H-bonding in quadruplexes contributes significantly to stability of the system. Frequency of H-bond formation analysis can elicit further insight into dynamic activity within a given com- plex. This factor was monitored per residue for each model.
Results
Table 1 summarises minima /maxima molecular dynamics data for biplanar and triplanar cytosine-based quadruplexes when co-ordi- nated by K+, Na+, Li+ or (for the biplanar model only) with no ion. Each result is described briefly in the text and shown graphically in Supplementary Material (eg.S1).
Triplanar cytosine quadruplex, CDN12 The RMSD trajectory for CDN12 K+ (S1), shows a strong rising trend between t = 0 ns and t = ~60ns from ~2.2 Å to ~4.2 Å which drops suddenly to ~3.2 Å and thereafter continues with a slight ris- ing trend to ~4.3 Å at the simulation end. There is some enhanced dynamic activity between ~180 Å and ~250 Å. The eRMSD tra- jectory (S2) is marginally dynamic but maintains a steady trend for the entire simulation with eRMSD values ranging between ~1.7 Å and ~2.8 Å. Radius of gyration for the system (S3) shows two rap- id cycles of expansion and contraction before returning to a much less dynamic trajectory to the simulation end. Maxima and minima are 0.5 Å and 1.5 Å. Time dependent data (S4) show an inconsis- tent pattern of H-bond formation. A histogram (S5) indicates the population mean ranges between seven and eleven, but values for zero or twenty are comparable.
Although the RMSD trajectory for CDN12 Na+ (S6), has a dis- tinctly dynamic rising trend for most of the simulation, there are two briefly level dynamic regions between t = 0ns and t = ~40 ns, and t = ~460 ns and 500 ns. A sharp peak is noticeable between t = ~190 ns and t = ~230 ns to ~4.7 Å. The eRMSD plot (S7), is very similar to the trajectory for the CDN12 K+ co-ordinated model. Although more dynamic activity is indicated, the values maintain a steady range between ~1.6Å and ~2.8 Å. There may be a poorly-defined cyclical trend to the data. Data returns for radius of gyration (S8), indicate the system is very dynamic with con- tinual cycles of contraction and expansion. There is a sudden and rapid reduction in Rg at ~210 ns. Time dependent data (S9) show an inconsistent pattern of H-bond formation similar to data for the K+ model except low single-figure values are more frequent. The histogram (S10) shows a poorly defined maximum between seven and fifteen.
The RMSD trajectory for CDN12 Li+ (S11), is marginally dynamic but noticeably level with values ranging between ~1.6 Å and ~2.7 Å. There is a brief peak centred on t = ~150 ns. Although some dynamic activity is indicated, the eRMSD trajectory (S12), main- tains a fairly constant path to t = ~280 ns. Thereafter, a slight rise is noticed to the simulation end. The plot is less erratic than either the K+ or Na+ models, with values centred on ~2.0 Å. The Radius of gyration trajectory (S13), shows very little activity for the duration of the simulation. There are no obvious changes in Rg with values ranging between 0.6 Å and 0.9 Å. Frequency of H-bond formation time dependent data (S14) show a marginally less erratic frequen- cy than for K+ or Na+ models but slightly greater number for zero and higher value populations. The histogram (S15) calculates a very poorly-defined maximum ranging between five and eighteen.
Biplanar cytosine quadruplex, CDN8
The RMSD trajectory for CDN8 without cationic co-ordination (S16), shows noticeable dynamic activity between t = 0 ns and t = ~70 ns with RMSD ranging from ~2.1 Å to ~4.5 Å and a brief smaller peak centred on t = ~200 ns. Thereafter the trajectory is steady with minimal activity. There is some dynamic activity re- corded in the eRMSD trajectory (S17), but no obvious continued rising or descending trend is indicated. Values range between ~1.3 Å and 2.0 Å. The radius of gyration (S18), is dynamic but displays only minor expansions and contractions. There is a slight reduc- tion at ~90 ns before a return at ~ 120 ns. Values range between 0.6 Å and 0.8 Å. Frequency of H-bond formation (S19), is relatively constant for the duration of the simulation. A large majority of the population centred on a frequency of two. A marginally lower pop- ulation frequency centred on four is predicted and a third stratum with a much lower population is centred on a frequency of eight. The histogram (S20) supports this observation.
The RMSD trajectory for CDN8 K+ (S21), is noticeably dynamic and generally follows a rising trend with RMSD values ranging between ~1.6 Å and ~4.9 Å. A less dynamic region occurs between t = 0ns and t = ~60 ns. There are two steep rising trends between t = ~220 ns and t = ~300 ns, and t = ~400 ns and 500 ns linked by a complementary descending trend. The eRMSD trajectory (S22), has a brief approximately level region to t = ~90 ns, after which there is noticeable activity. A sharp rising trend to ~2 .9 Å is indi- cated at t = ~180 ns which levels to an erratic plot ranging between ~1.5 Å and ~2.5 Å to the simulation end. The radius of gyration trajectory (S23), is dynamic with no obvious cyclical expansion or contraction. The model appears to develop more instability after ~ 300 ns. For frequency of H-bond formation, time-dependent data (S24) show the majority of values calculated at five or less are not apparent as observed in the strata of 3.6.1.4. Maxima shown in the histogram (S25) centre on five H-bonds, but range between four and seven.
RMSD values for CDN8 Na+ (S26), closely resemble the trajec- tory for CDN8 co-ordinated by K+ and overall show similar dy- namic activity. The eRMSD trajectory (S27) is highly erratic for the entire simulation with values similar to the CDN8 K+ co-or- dinated model. For Radius of gyration data (S28), there is a slight downward trend in values to ~200 ns after which the system de- velops noticeable variation. There is no obvious cyclical character to the trajectory. There appear to be minor peaks in the frequency of H-bond formation at three, five and six H-bonds (S29) although local maxima appear across most of the histogram (S30).
RMSD data for CDN8 Li+ (S31) indicate a sharp rising trend be- tween t = 0 ns and t = ~70 ns from ~1.5 Å to ~3.0 ns, after which the trajectory is dynamic around ~2.8 Å to 500 ns. A noticeable drop in RMSD occurs between t = ~220 ns and t = ~350 ns. The eRMSD trajectory (S32) is dynamic with a steady rising trend after t = ~100 ns. The range of eRMSD values is similar to the K+ and Na+ co-ordinated models. Radius of gyration values (S33) centre on ~0.8 Å for the duration of the simulation with very little vari- ation. There may be a minor expansion and reduction cycle. Fre- quency of H-bond formation analysis shows local maxima at four, five and six H-bonds (S34) with all values predicted to be less than ten (S35). Occupancy of waters in CDN12 and CDN8.
Introduction and methodology
Arrays of bridging water molecules are known to be imperative in binding ligands to parallel G4s; hydrogen bonding mediates interaction between the two species [38]. Interplanar loops are populated with waters and regarded as reinforcing stability of their conformation [39]. However, such waters are superficial or peripheral to a quadruplex; little is known of waters internalised into the system. Ostensibly, molecular dynamics simulations ap- pear to be unsuited in resolving this problem. A hydration shell neutralised with counterions and enclosing any given quadruplex model may typically contain between 15 000 and 20 000 water molecules. Consequently, manipulating the model to ascertain the number of waters located in the centre of the structure over time is highly impractical. To address this challenge, a script was written to determine by calculation occupancy of waters rather than by inspection (Supplementary Material, script 1).
Predicted interplanar water occupancy of biplanar and triplanar cytosine quadruplexes was calculated. For a closer comparison with triplanar models, biplanar equivalents were analysed with co-ordinating ions K+, Na+, or Li+ imposed into the systems in addition to the archetype model exclusive of ions. CDN12 was assessed with K+, Na+, or Li+ co-ordination to reflect its derivation from (PDB) 1KF1.
Empirical values (S36-S38) and histograms (figures 8 and 9) were plotted for each system.
Results: Occupancy of waters
Figure 8: Histogram of interplanar water occupancy in triplanar cytosine quadruplex models co-ordinated by K+, Na+ or Li+. Time-de- pendent data for this histogram are plotted in S36-S38.
Water occupancy for K+ co-ordinated models is higher overall than for the Na+ and Li+ systems, but all three indicate an average pop- ulation between four and six waters. The histogram implies high- er numbers of waters are slightly better tolerated than for K+ and Na+ models. This finding compares with a population approaching zero in Li+ co-ordinated triplanar cytosine quadruplexes. Figures S36-S38 reinforce this finding: time-dependent data show Li+ systems have the most consistent water occupancies of the three models.
Figure 9: Histogram of interplanar water occupancy in biplanar cytosine quadruplex models with no cationic co-ordination or co-ordi- nated by K+, Na+ or Li+. Time-dependent data for this histogram are plotted in S39, S40, S41 and S42.
Biplanar DNA systems with monovalent cations artificially im- posed contrast with triplanar models where co-ordinants are typ- ically present. A higher population of waters is generally tolerat- ed better in biplanar systems without cationic co-ordination than models with co-ordinants. This observation is particularly relevant for higher occupancies. Li+ co-ordinated models are predicted to consistently allow a slightly higher population of waters than K+ or Na+ systems. Time dependent data (figures S40, S41 and S42) indi- cate a wider variation of numbers in cationic co-ordinated models. Deoxyribose conformations in CDN12 and CDN8.
Introduction and methodology
The non-planarity of ribose as a sugar ring in nucleic acids is pred- icated on the four carbon atoms and one oxygen comprising the structure. Accordingly, the exact non-planar conformation derives from the five interior angles of the pentagonal ring. Implicitly, the most energetically favourable conformation would show the max- imum possible interatomic distances between the principal com- ponents. The basic substituents of DNA and RNA sugar rings are shown in figure 10.
Figure 10: comparison of deoxyribose (A) and ribose (B) and atom numeration. The C2’ carbon of ribose has a single bonded oxygen
unlike deoxyribose. The additional oxygen allows greater reactivity with consequent biological ramifications (discussed in text).
Unequal charge distribution in the molecule is contingent on these atomic components. The lone pair of O4’ could prompt dipole in- teractions therefore subsequent differences in non-planarity, an in- teraction colloquially referred to as the Rabbit Ear Effect. Interac- tions in DNA and RNA are susceptible to the electronegativity of the sugar-phosphate backbone [40]. Reciprocity arises from re-ori- entation of substituents triggering changes in planarity and caus- ing significant alterations in the backbone [41, 42]. Consequently, there are ostensible links within a biological framework [43].
The displacement vector away from the plane determines termi- nology. Carbon facing the plane of the base is defined as an -endo configuration; facing away from the base defines an -exo configu- ration. However, naïve conformers are affected by substituent dif- ferentials in the sugar ring. Hence there is some latitude in an exact angular definition for both endo forms; there is a range of values for internal angles within which the endo form can arise. C2’-en- do structures are ubiquitous in purines, but pyrimidines favour the C3’-endo form. The prevalence in RNA and C3’-endo conformers is contingent on the marginally greater interatomic distance be- tween C2’ and C3’ oxygens in ribonucleosides. Fluctuations in the internal angles of the sugar component result in various conforma- tions. Two in particular were analysed in this study: C2’-endo and C3’-endo conformations (figure 11).
Figure 11: The C2′-endo conformation is uniquely observed in β-form DNA (A) but C3′-endo is prevalent in RNA and α-form DNA (B). Distance d between consecutive phosphate atoms is greater for a C2′-endo conformation as a direct result of the molecular re-alignment.
Since the change from C2’-endo to C3’-endo takes place via a series of marginally different intermediates, the progression of non-planar conformers may be presenting as a continuum of variation [44]. Nevertheless, the structures are defined by energy barriers. The conversion of C2’-endo to C3’-endo is mediated by O4’-endo.
The energy barrier ranges from 2 kcal mol-1 to 5 kcal mol-1 with an overall potential energy of 1.5 kcal mol-1 [45, 46]. The C3′-en- do pucker produces a significantly shorter phosphate-phosphate distance in the backbone, resulting in a more compact helical conformation. C2’-endo and C3’-endo conformations (figure 11) of cytosine models were analysed because the C3′-endo pucker produces a significantly shorter phosphate-phosphate distance in the backbone. A more compact helical conformation results with consequent implications for stability. Although pyrimidines favour the C3’-endo form, the C2’-endo configuration tends to dominate in DNA systems. Although 2KF8 is characterised by absence of a co-ordinant, CDN8 models also had K+, Na+ or Li+ imposed as co-ordinat- ing ions. Triplanar systems were co-ordinated by K+, Na+ or Li+. Mean distances monitored were very similar for both systems: 5.9Å for C3’-endo and 7.2Å for C2’-endo DNA configurations. Configurations are bounded by an approximate range of dihedral angles; C2’endo: 0o to 36o, C3’-endo: 144o to 180o [47]. The spe- cific dihedral describing these configurations is C’1-C2’-C3’-C4’ and calculated with the VMD command: measure dihed {a b c d} frame all where a–d are atom numbers for C1’-C2’-C3’-C4’. These atoms are listed in the .pdb file as: 156 155 152 151. This C1’-C2’-C3’-C4’ dihedral was calculated for three sugar rings of residues biplanar guanine quadruplexes. Residue 7 was selected as a typical quartet constituent.
Results: deoxyribose conformations
Trajectories of triplanar DNA cytosine quadruplexes ribose C2’ / C3’ fluctuations are typified by data shown for residue seven with different co-ordinating ions (figures 12a, 12b and 12c):
Figures 12a, 12b and 12c: trajectories for C1’-C2’-C3’-C4’ dihedrals in DNA triplanar cytosine models in residue 7 co-ordinated with K+ (left), Na+ (centre) and Li+ (right).
The majority of systems appear to favour the C2’-endo configuration, but all show notable dynamic activity. Models with Na+ co-ordi- nation typically the most erratic trajectories with poor definition. Li+ co-ordinated systems are also dynamic but generally show clear changes between C2’-endo and C3’-endo configurations. K+ models appear to show brief periods of stable C2’-endo forms alternating with highly erratic and poorly-defined trajectories (Table 2).
<img src="https://www.opastpublishers.com/scholarly-images/6266-698ad840ad024-cytosine-quadruplexes-in-duplex-dna-may-be-nontransient-and-.png" width="900" height="500">
Figures S43 and S44 show data for models co-ordinated with Na+ and Li+.
CDN8 in the absence of ionic co-ordination shows an overall endency to the C3’-endo configuration. However, models with K+, Na+ or Li+ co-ordination are notably erratic with poor definition for either configuration (figures 13a, 13b, S45 and S46):
Figures 13a and 13b, trajectories for C1’-C2’-C3’-C4’ dihedrals in DNA biplanar cytosine models in residue 7 with no ionic co-or- dination (left) or co-ordinated with K+ (right). Trajectories for systems co-ordinated with Na+ or Li+ are shown in figures S45 and S46.
<img src="https://www.opastpublishers.com/scholarly-images/6266-698ad8db453e3-cytosine-quadruplexes-in-duplex-dna-may-be-nontransient-and-.png" width="500" height="400">
Figure 14: distribution histograms from figures 13a and 13b data for CDN8-N1 no ionic co-ordination (blue) and CDN12-N1 K+ co-ordination dihedrals (blue). C2’-endo and C3’-endo configurations are evenly represented for the duration of the simulation for both models. The wide range of conformations indicate sugars in each system fluctuate frequently during the simulation.
Biplanar quadruplexes characterised by absence of a co-ordinant consistently show configurations in the C3’-endo region (Table 3).
<img src="https://www.opastpublishers.com/scholarly-images/6266-698ad9424d811-cytosine-quadruplexes-in-duplex-dna-may-be-nontransient-and-.png" width="800" height="300">
Discussion of C2’-endo and C3’endo configurations
Conformational re-alignment from C2’-endo to C3’endo may in- fluence nucleotide polymerisation whereas C3’-endo to C2’-endo may be crucial in the catalytic capacity of RNA [47, 48]. These two observations imply that flexibility of configuration is an im- portant biological requirement. A contribution to stability of the C2’-endo conformer might derive from reduced hydration waters. Consequently, there is an overall reduction in the enthalpy of hy- dration as this configuration would require fewer water molecules. Conversely, this implies there should be an entropic cost for the reconfiguration to the C3’-endo conformer as a reduced population of waters would need to off-set the energy differential. In turn, it follows that the C3’-endo structure must be biologically imperative if the transition occurs regularly despite being thermodynamically unfavourable. Data for C1’-C2’-C3’-C4’ dihedrals in the biplanar quadruplexes investigated here, strongly indicate compromise by the entropic tariff of differential hydration between C2’-endo and C3’endo configurations.
Discussion of cytosine quadruplexes
The concept of cytosine dimers pairing into a quartet, then three quartets stacking vertically into a cytosine quadruplex (C4), was addressed in this study. The quartets were linked with AAT loops, and physiologically relevant K+, Na+ or Li+ were selected as co-ordinating ions. Contrasting with guanine quadruplexes co-or- dinated with Li+ as contentious, a biplanar cytosine equivalent structure with no cationic co-ordination is found in this study to be non-transient [49, 50]. However, the greater dynamic activity predicted for Li+, K+ or Na+ co-ordinated biplanar models indi- cates triplanar systems may be transient. All triplanar models are characterised by greater dynamic activity and therefore strongly suggestive of reduced stability. The results indicate the stability of cytosine quadruplexes varies not only between triplanar and bipla- nar structures, but also the presence or absence of a co-ordinating ion. For triplanar systems, cations remain within the complex and underpin quadruplex stability. This observation contrasts notably with dynamic activity within biplanar systems where presence of co-ordinating cation reduces overall integrity of a complex.
RMSD and eRMSD data for triplanar models coincide with a se- quence of higher values and more erratic trajectories in the se- quence Li+ > K+ > Na+. Less dynamic trajectories are evidenced in biplanar models, but with lower deviation. However, all biplanar models with cationic co-ordination are notably more dynamic than a system without an ion. Radius of gyration data generally confirm this trend with some exceptions. In a triplanar system, Li+ and K+ have similar compactness and which is less than for Na+-co-or- dinated models. All biplanar models with cationic co-ordination consistently show greater expansion than for a system with no cat- ion.
Frequency of H-bond formation appears to be generally consistent amongst all models whilst noting that biplanar systems have lower values overall. This would accord with only two cytosine planes available for interaction as opposed to three in triplanar quadru- plexes. As a general observation, frequency of H-bond formation in Na+-co-ordinated models is slightly higher than for other sys- tems including a biplanar quadruplex without cationic co-ordi- nation. This may be an artefact of hydrated sodium ion clusters which arise more readily than other Group 1 metals [51]. If such clusters were to arise in Na+-co-ordinated models even transiently, H-bonding may occur at a slightly higher frequency overall.
For triplanar quadruplexes, interplanar occupancy of waters varies for K+ and Na+-co-ordinated models more than for Li+ system. Whilst this observation may underpin some of the enhanced fre- quency of H-bond formation in Na+-co-ordinated models, it might plausibly apply to K+ systems. This trend is not displaced by water occupancy being lower in biplanar models, although less available interplanar access must be a major factor, if not solely responsible.
Although the data for ribose configuration are generally inconsis- tent for all models, some trends are noted. Trajectories for tripla- nar Na+-co-ordinated systems have no discernable patterns and this observation applies to both K+ and Na+-co-ordinated biplanar quadruplexes. The trajectory for a biplanar system with no cationic co-ordination is an exception to these observations. The configura- tion is calculated as consistently C3’-endo. There may be some pe- riodicity to a triplanar K+-co-ordinated system and an Li+-co-or- dinated models generally show a shift from C2’-endo C3’-endo at an approximate mid-point of the simulation.
Conclusion
Overall, this study implies a biplanar cytosine quadruplex is sta- ble to 500 ns in the absence of cationic co-ordination, but models co-ordinated by Li+, K+ or Na+ may be less stable. Triplanar cy- tosine quadruplexes retain their co-ordinating ions but generally exhibit more erratic trajectories across most metrics.
If biplanar cytosine quadruplexes do fold in single-strand DNA, their viability is credible but may be less stable than triplanar gua- nine systems. The latter are characterised by three H-bonds be- tween bases contrasting with only two in cytosine-cytosine base pairing. Cationic co-ordination is also a factor for both species of theoretical cytosine quadruplexes: paradoxically, absence stabilis- es biplanar models, but presence reduces overall integrity of tripla- nar systems. However, two points arise: firstly, both show a robust network of inter-planar π-stacking. Secondly, although there is an H-bond differential between guanine and cytosine models, geome- try may influence the formation. The combination may be tending towards a lower energy configuration.
Executive summary
• Cytosine is known to dimerise
• Two dimers are shown in this study to have the potential for self-associating non-covalently into a planar quartet
• Given a specific sequence of bases, biplanar and triplanar qua- druplexes could develop from cytosine quartets
• Stability of such supramolecular structures depends on sever- al factors and varies widely as a consequence
• Cytosines dimers participating in the central core of a theoret- ical quadruplex are mutually ligated with two hydrogen bonds but paired dimers have only one, a total of six per quartet. Guanine equivalents have two hydrogen bonds between each base, a total of eight per quartet.
• Cationic co-ordination marginally reduces stability in tripla- nar models, but inclusion of a cation in biplanar systems is very disruptive. • Some enhancement of structural integrity for cytosine and systems may derive from a lowest energy geometry and no- ticeable interplanar π-bonding networks.
Future work
Plausibly, cytosine quadruplexes may have limited physiological relevance predicated on relatively poor stability compared to gua- nine equivalents. If these structures were to fold genomically, per- haps any significance lies with intermittent appearance. Although short-lived, a folding / unfolding cycle may persist into biolog- ically relevant times and influence epigenetic processes. Such mechanisms could be amenable to molecular modelling and ther- modynamic analysis. Methylation at C5 or C6 of cytosine would probably further reduce the molecular lifetime of a particular quadruplex. However, methylation may also tag the structure for downstream modification: docking protocols with small molecules or enzymes should adumbrate further perspectives. For instance, an antisense cytosine quadruplex might compromise the benefi- cial effects of a complementary sense-strand guanine equivalent. The novel suggestion of an anti-sense guanine quadruplex is also raised. Thymine substitution in quadruplex structures may further qualify the data for guanine and cytosine systems.
References
- Gehring, K., Leroy, J. L., & Guéron, M. (1993). A tetrameric DNA structure with protonated cytosine-cytosine base pairs. Nature, 363(6429), 561-565.
- Leroy, J. L., Gueron, M., Mergny, J. L., & Helene, C. (1994). Intramolecular folding of a fragment of the cytosine-rich strand of telomeric DNA into an i-motif. Nucleic acids re- search, 22(9), 1600-1606.
- Mergny, J. L., Lacroix, L., Han, X., Leroy, J. L., & Helene,C. (1995). Intramolecular folding of pyrimidine oligodeoxy- nucleotides into an i-DNA motif. Journal of the American Chemical Society, 117(35), 8887-8898.
- Day, H. A., Pavlou, P., & Waller, Z. A. (2014). i-Motif DNA: structure, stability and targeting with ligands. Bioorganic & medicinal chemistry, 22(16), 4407-4418.
- Phan, A. T., & Leroy, J. L. (2000). Intramolecular i-motif structures of telomeric DNA. Journal of Biomolecular Struc- ture and Dynamics, 17(sup1), 245-251.
- Choi, J., Kim, S., Tachikawa, T., Fujitsuka, M., & Majima, T. (2011). pH-induced intramolecular folding dynamics of i-mo- tif DNA. Journal of the American Chemical Society, 133(40), 16146-16153.
- Gallego, J., Chou, S. H., & Reid, B. R. (1997). Centromericpyrimidine strands fold into an intercalated motif by forming a double hairpin with a novel T: G: G: T tetrad: solution struc- ture of the d (TCCCGTTTCCA) dimer. Journal of molecular biology, 273(4), 840-856.
- Balasubramanian, S., Hurley, L. H., & Neidle, S. (2011). Tar- geting G-quadruplexes in gene promoters: a novel anticancer strategy?. Nature reviews Drug discovery, 10(4), 261-275.
- Wang, E., Thombre, R., Shah, Y., Latanich, R., & Wang, J. (2021). G-Quadruplexes as pathogenic drivers in neurodegen- erative disorders. Nucleic acids research, 49(9), 4816-4830.
- Simone, R., Fratta, P., Neidle, S., Parkinson, G. N., & Isaacs,A. M. (2015). G-quadruplexes: Emerging roles in neurode- generative diseases and the non-coding transcriptome. FEBS letters, 589(14), 1653-1668.
- Huppert, J. L., & Balasubramanian, S. (2007). G-quadruplex- es in promoters throughout the human genome. Nucleic acids research, 35(2), 406-413.
- Verma, A., Yadav, V. K., Basundra, R., Kumar, A., & Chow- dhury, S. (2009). Evidence of genome-wide G4 DNA-medi- ated gene expression in human cancer cells. Nucleic acids research, 37(13), 4194-4204.
- Smestad, J. A., & Maher, L. J. (2015). Relationships between putative G-quadruplex-forming sequences, RecQ helicases, and transcription. BMC medical genetics, 16, 1-14.
- Saikia, N., Johnson, F., Waters, K., & Pandey, R. (2018). Dy- namics of self-assembled cytosine nucleobases on graphene. Nanotechnology, 29(19), 195601.
- Sun, Z. Y., Wang, X. N., Cheng, S. Q., Su, X. X., & Ou, T. M.(2019). Developing novel G-quadruplex ligands: From inter- action with nucleic acids to interfering with nucleic acid–pro- tein interaction. Molecules, 24(3), 396.
- Jawiczuk, M. (2018). A theoretical study on the hydrogen bond and stability of cytosine and thymine dimers. Computa- tional and Theoretical Chemistry, 1123, 26-34.
- Cheng, R., Loire, E., & Fridgen, T. D. (2019). Hydrogen bonding in alkali metal cation-bound i-motif-like dimers of 1-methyl cytosine: An IRMPD spectroscopic and computa- tional study. Physical Chemistry Chemical Physics, 21(21), 11103-11110.
- Boese, A. D. (2015). Density functional theory and hydrogen bonds: are we there yet?. ChemPhysChem, 16(5), 978-985.
- Barman, P., Reddy, D., & Bhaumik, S. R. (2019). Mecha- nisms of antisense transcription initiation with implications in gene expression, genomic integrity and disease pathogenesis. Non-coding RNA, 5(1), 11.
- Zamiri, B., Mirceta, M., Bomsztyk, K., Macgregor Jr, R. B., & Pearson, C. E. (2015). Quadruplex formation by both G-rich and C-rich DNA strands of the C9orf72 (GGGGCC) 8•(GG- CCCC) 8 repeat: effect of CpG methylation. Nucleic acids re- search, 43(20), 10055-10064.
- Kravchenko, J., Darrah, T. H., Miller, R. K., Lyerly, H. K., & Vengosh, A. (2014). A review of the health impacts of bari- um from natural and anthropogenic exposure. Environmental geochemistry and health, 36, 797-814.
- Patel, P. K., Bhavesh, N. S., & Hosur, R. V. (2000). Cation-de-pendent conformational switches in d-TGGCGGC containing two triplet repeats of Fragile X Syndrome: NMR observa- tions. Biochemical and Biophysical Research Communica- tions, 278(3), 833-838.
- Kocman, V., & Plavec, J. (2017). Tetrahelical structural fam- ily adopted by AGCGA-rich regulatory DNA regions. Nature Communications, 8(1), 15355.
- Cáceres, C., Wright, G., Gouyette, C., Parkinson, G., & Sub- irana, J. A. (2004). A thymine tetrad in d (TGGGGT) quadru- plexes stabilized with Tl+/Na+ ions. Nucleic acids research, 32(3), 1097-1102.
- Liu, H., Wang, R., Yu, X., Shen, F., Lan, W., et al. (2018).High-resolution DNA quadruplex structure containing all the A-, G-, C-, T-tetrads. Nucleic Acids Research, 46(21), 11627- 11638.
- Li, T., Li, B., Wang, E., & Dong, S. (2009). G-quadru- plex-based DNAzyme for sensitive mercury detection with the naked eye. Chemical Communications, (24), 3551-3553.
- Patel, P. K., Bhavesh, N. S., & Hosur, R. V. (2000). NMR observation of a novel C-tetrad in the structure of the SV40 repeat sequence GGGCGG. Biochemical and biophysical re- search communications, 270(3), 967-971.
- Asamitsu, S., Obata, S., Yu, Z., Bando, T., & Sugiyama, H. (2019). Recent progress of targeted G-quadruplex-preferred ligands toward cancer therapy. Molecules, 24(3), 429.
- Kumari, S., Bugaut, A., Huppert, J. L., & Balasubramanian,S. (2007). An RNA G-quadruplex in the 5′ UTR of the NRAS proto-oncogene modulates translation. Nature chemical biol- ogy, 3(4), 218-221.
- Puig Lombardi, E., Holmes, A., Verga, D., Teulade-Fichou,M. P., Nicolas, A., et al. (2019). Thermodynamically stable and genetically unstable G-quadruplexes are depleted in ge- nomes across species. Nucleic Acids Research, 47(12), 6098- 6113.
- RebicÌ?, M., Laaksonen, A., SÌ?poner, J., UlicÌ?nyÌ, J., & Mocci,F. (2016). Molecular dynamics simulation study of parallel telomeric DNA quadruplexes at different ionic strengths: Evaluation of water and ion models. The Journal of Physical Chemistry B, 120(30), 7380-7391.
- Bhattacharyya, D., Mirihana Arachchilage, G., & Basu, S. (2016). Metal cations in G-quadruplex folding and stability. Frontiers in chemistry, 4, 38.
- Mignon, P., Loverix, S., Steyaert, J., & Geerlings, P. (2005). Influence of the π–π interaction on the hydrogen bonding ca- pacity of stacked DNA/RNA bases. Nucleic Acids Research, 33(6), 1779-1789.
- Jorgensen, W. L. (1981). Quantum and statistical mechanical studies of liquids. 10. Transferable intermolecular potential functions for water, alcohols, and ethers. Application to liq- uid water. Journal of the American Chemical Society, 103(2), 335-340.
- Hershkovitz, E., Sapiro, G., Tannenbaum, A., & Williams, L.D. (2006). Statistical analysis of RNA backbone. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 3(1), 33-46
- Leontis, N. B., & Westhof, E. (2001). Geometric nomencla-ture and classification of RNA base pairs. Rna, 7(4), 499-512.
- Arnittali, M., Rissanou, A. N., & Harmandaris, V. (2019). Structure of biomolecules through molecular dynamics simu- lations. Procedia Computer Science, 156, 69-78.
- Neidle, S. (2021). Structured waters mediate small molecule binding to G-quadruplex nucleic acids. Pharmaceuticals, 15(1), 7.
- Li, K., Yatsunyk, L., & Neidle, S. (2021). Water spines and networks in G-quadruplex structures. Nucleic Acids Re- search, 49(1), 519-528.
- Stasyuk, O. A., Jakubec, D., Vondrasek, J., & Hobza, P. (2017). Noncovalent interactions in specific recognition mo- tifs of protein–DNA complexes. Journal of Chemical Theory and Computation, 13(2), 877-885.
- Bryce, D. L., Grishaev, A., & Bax, A. (2005). Measurement of ribose carbon chemical shift tensors for A-form RNA by liquid crystal NMR spectroscopy. Journal of the American Chemical Society, 127(20), 7387-7396.
- Potaman, V. N., & Sinden, R. R. (2013). DNA: Alternative conformations and biology. In Madame Curie Bioscience Da- tabase [Internet]. Landes Bioscience.
- Saragi, R. T., Juanes, M., Abad, J. L., Pinacho, R., Rubio, J. E., et al. (2022). Chirality-Puckering correlation and intermo- lecular interactions in Sphingosines: Rotational spectroscopy of jaspine B3 and its monohydrate. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy, 267, 120531.
- Olson, W. K., & Sussman, J. L. (1982). How flexible is the fu- ranose ring? 1. A comparison of experimental and theoretical studies. Journal of the American Chemical Society, 104(1), 270-278.
- Harvey, S. C., & Prabhakaran, M. (1986). Ribose puckering: structure, dynamics, energetics, and the pseudorotation cycle. Journal of the American Chemical Society, 108(20), 6128- 6136.
- Foloppe, N., Nilsson, L., & MacKerell Jr, A. D. (2001). Ab initio conformational analysis of nucleic acid components: intrinsic energetic contributions to nucleic acid structure and dynamics. Biopolymers: Original Research on Biomolecules, 61(1), 61-76.
- Julien, K. R., Sumita, M., Chen, P. H., Laird-Offringa, I. A., & Hoogstraten, C. G. (2008). Conformationally restricted nucle- otides as a probe of structure–function relationships in RNA. RNA, 14(8), 1632-1643.
- Zhang, N., Zhang, S., & Szostak, J. W. (2012). Activated ri- bonucleotides undergo a sugar pucker switch upon binding to a single-stranded RNA template. Journal of the American Chemical Society, 134(8), 3691-3694.
- Sacca, B., Lacroix, L., & Mergny, J. L. (2005). The effect of chemical modifications on the thermal stability of different G-quadruplex-forming oligonucleotides. Nucleic acids re- search, 33(4), 1182-1192.
- Risitano, A., & Fox, K. R. (2003). Stability of intramolecular DNA quadruplexes: comparison with DNA duplexes. Bio- chemistry, 42(21), 6507-6513
- Wang, P., Shi, R., Su, Y., Tang, L., Huang, X., et al. (2019). Hydrated Sodium Ion Clusters [Na+ (H2O) n (n= 1–6)]: An ab initio Study on Structures and Non-covalent Interaction. Frontiers in chemistry, 7, 624.


