Research Article - (2025) Volume 10, Issue 3
Changing Sensitivity Pattern of Pseudomonas Species- A 5-year Study at a Tertiary Care Hospital
2Department of Internal Medicine, Foundation University Medical College, Islamabad, Pakistan
3Department of Internal Medicine, Foundation University Medical College, Islamabad, Pakistan
4Department of Internal Medicine, Foundation University Medical College, Islamabad, Pakistan
5Department of Internal Medicine, Foundation University Medical College, Islamabad, Pakistan
6Department of Internal Medicine, Foundation University Medical College, Islamabad, Pakistan
7Department of Pathology, Foundation University School of Health Sciences (FUSH), Islamabad, Pakistan
8Department of Pathology, Fauji Foundation Hospital Rawalpindi, Foundation University School of Healt, Pakistan
Received Date: Jun 20, 2025 / Accepted Date: Jul 21, 2025 / Published Date: Jul 28, 2025
Copyright: ©�2025 Hafiz Muhammad Hamz, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Ali, M. M., Hamza, H. M., Ullah, A. I., Tariq, M. D., Manzoor, M. I. A., et al. (2025). Changing Sensitivity Pattern of Pseudomonas Species- A 5-year Study at a Tertiary Care Hospital. J Nur Healthcare, 10(3), 01-09.
Abstract
Purpose: Pseudomonas aeruginosa is a Gram-negative opportunistic organism known for its ability to cause serious infections, particularly in immunocompromised individuals. The development and dissemination of antimicrobial resistance in Pseudomonas aeruginosa have become a significant global health concern. The study aims to investigate the antimicrobial sensitivity patterns of Pseudomonas aeruginosa over the past five years in a tertiary care setting.
Methods: A comprehensive retrospective analysis was conducted using clinical data from a tertiary healthcare setting spanning five years (2018-2022). Isolates of Pseudomonas aeruginosa obtained from various clinical specimens were included in the study. Standardized procedures were used to perform the antimicrobial susceptibility tests, and the relevant statistical techniques were applied to analyze the data.
Results: A total of 1888 Pseudomonas aeruginosa isolates were included in the study. The study revealed an increasing trend in antimicrobial resistance among Pseudomonas aeruginosa isolates over five years. The most significant resistance was observed against commonly used antibiotics such as carbapenems, fluoroquinolones, and aminoglycosides.
Conclusion: This comprehensive analysis of Pseudomonas aeruginosa antimicrobial susceptibility patterns over the past five years emphasizes the increasing concern about antibiotic resistance in clinical settings. The rising resistance rates, particularly against essential antibiotics, underscore the urgent need for effective antimicrobial stewardship programs and the development of alternative therapeutic approaches to combat Pseudomonas aeruginosa infections. This study offers significant perspectives on the dynamic landscape of antibiotic resistance, empowering medical practitioners to make knowledgeable choices about the most suitable treatment plans for Pseudomonas aeruginosa infections. Keywords: Pseudomonas aeruginosa, antibiotic resistance, nosocomial infections, changing pattern, opportunistic pathogen.
Keywords
Pseudomonas Aeruginosa, Antibiotic Resistance, Sensitivity Pattern, Hospital Acquired Infection, Five Years
Introduction
Pseudomonas aeruginosa is opportunistic human bacteria that frequently causes infections that are challenging to treat because of its exceptional capacity to develop resistance to therapy and poor intrinsic sensitivity to medications [1]. The Gram-negative bacterial pathogen Pseudomonas aeruginosa is a frequent cause of nosocomial infections, including pneumonia, infection in immunocompromised hosts, and infection in persons with structural lung illnesses such as cystic fibrosis [2]. Depending on the area and the particular strain of the bacteria, Pseudomonas aeruginosa's sensitivity pattern to different antibiotics can vary. Most European nations report more than 10% resistance rates for all antibiotic classes under observation. In 2015, the European Centres for Disease Prevention and Control reported that 13.7% of Pseudomonas aeruginosa isolates exhibited resistance to at least three antimicrobial groups, and 5.5% of isolates were resistant to all five antimicrobial groups under observation (EARS-Net).
Multidrug resistant (MDR) Pseudomonas aeruginosa is the cause of 13% of healthcare-associated severe infections, according to data from a study conducted in the United States in 2019 [3]. According to published studies, Pseudomonas aeruginosa is one of ICUs' most frequently isolated infections. Pseudomonas aeruginosa was reported to have the highest level of antibiotic resistance against meropenem, followed by ticarcillin, imipenem, and piperacillin; 13% of the isolates exhibited multi-drug resistance [4].
According to a research carried out in KMCTH (Kathmandu Medical College Teaching Hospital, Nepal) in 2018, Pseudomonas aeruginosa isolates were resistant to 65.21% of cefixime, 63.04% of ceftazidime, 56.52% of ceftriaxone, 56.52% of cefotaxime and 56.52% of piperacillin [5]. According to a study done in Pakistan, ofloxacin had the highest level of quinolone activity (51% sensitivity) against the resistant bacterium, followed by ciprofloxacin (50%), sparfloxacin (49.8%) and enoxacin (49.6%) [6]. However, a Pakistani study found that 25% of patients were resistant to ciprofloxacin. According to the same survey, nalidixic and pipemidic acid resistance was 87% and 25%, respectively. In another investigation, the resistance of these two antimicrobials was found to be 98.8% and 98.6%, respectively, verifying the quinolones' restrained therapeutic activity due to their continuous use [6]. According to the findings, Pseudomonas aeruginosa acquires various genetic variants depending on whether it is exposed to tobramycin, ceftazidime, or ceftazidime-avibactam when it is present in urine, synthetic sputum, or laboratory-rich medium [1].
The virulence factor that Pseudomonas aeruginosa employs to enter the bloodstream during infection may cause an excessive and improperly controlled immune response to bacteria, resulting in sepsis [7]. Amoxicillin, oxytetracycline, and doxycycline were all utterly ineffective against any isolates (93.3 and 86.7 percent, respectively). All clinical isolates showed resistance to oxytetracycline, doxycycline, and amoxicillin [8]. The Pakistan Journal of Medical Sciences published a 2018 study that revealed Pseudomonas aeruginosa to be highly resistant to imipenem, ciprofloxacin, and ceftriaxone; however, it did exhibit some susceptibility to amikacin and piperacillin-tazobactam [9].
It is significant to remember that antibiotic resistance patterns can alter over time, and local surveillance data should be routinely checked to help determine the best course of antibiotic treatment for infections brought on by Pseudomonas aeruginosa. This study aims to investigate the susceptibility patterns of Pseudomonas aeruginosa over the most recent years in Pakistan to update our local empirical therapy accordingly.
Material and methods
A retrospective cross-sectional study was conducted by the Department of Microbiology at the Fauji Foundation Hospital (FFH) in Rawalpindi. The Institutional Ethical Review Board provided the necessary ethical clearance for the study. The research encompassed all Pseudomonas aeruginosa samples collected in the microbiology laboratory between January 2018 and December 2022. During the course of this study, a total of 1888 samples of Pseudomonas aeruginosa isolates were received. However, to ensure the integrity and accuracy of the data, certain exclusions were made. Specifically, any duplicates or samples from patients who were already receiving broad-spectrum antibiotics were not included in the analysis. It is worth noting that all other samples were included in the study, and great care was taken to maintain the highest standards of quality control throughout the process. In the context of routine phenotypic tests regularly performed in clinical laboratories, we conducted inoculation of samples on various selective media to identify Pseudomonas aeruginosa. The media employed for this purpose included blood agar and MacConkey's agar, and the samples underwent incubation at a temperature of 37°C for a period of 24-48 hours.
The isolates were identified based on a series of routine tests, including colony morphology, positive oxidase test, glucose fermentation, hydrolysis of gelatin, and growth at a temperature of 42°C. [10]. Subsequent to the microorganism isolation, an antimicrobial sensitivity test was performed using a modified Kirby Bauer disc diffusion method on Muller Hinton agar (Oxoid). The antimicrobial disks, with their respective strengths, were applied to the isolates and incubated for a period of 18 to 24 hours at 370°C.
The aforementioned procedure was conducted to determine the sensitivity of the microorganisms to the antimicrobials. Below is a list of the antimicrobial discs that were used throughout the process, along with an explanation of their respective strengths: Penicillin (10µg), Amikacin (30µg), Ampicillin (10µg), Augmentin (30µg), Azithromycin, Ceftazidime (30), Ceftriaxone (30µg), Chloramphenicol (30µg), Ciprofloxacin (5µg), Colistin, Cotrimoxazole (25µg), Gentamicin (10µg), Imipenem (10µg), Augmentin, Meropenem (10µg) and Cephradine (10µg). Antibiotic sensitivity data was analyzed using Clinical Laboratory Standards Institute (CLSI 2022) standards [11]. Microsoft Excel version 2019 was utilized to calculate dependent variables such as the frequency of bacterial isolates and the pattern of their antimicrobial sensitivity, as well as independent variables such as the socio-demographic data of patients.
Results
Different age groups were investigated; however, the patients in the (33–64) age group had the highest spike in Pseudomonas aeruginosa infections, as shown in Figure 1.
Figure 1: Demographic Presentation of Different Age Groups with Sample Sizes
A total of 1888 Pseudomonas aeruginosa were isolated from the sample in which pus culture had 1138 (60%) isolates of Pseudomonas aeruginosa, followed by sputum culture 584 (31%), blood culture 63(3.3%), CSF 52 (3%) and the HVS 51 (3%) which are shown in Figure 2 and Table 1
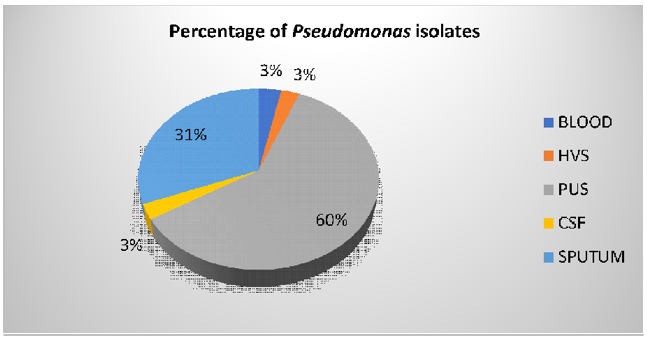
Pseudomonas aeruginosa isolates showed varying resistance trends to colistin. In pus cultures, colistin exhibited a susceptibility rate of 56.2%. However, in Cerebrospinal Fluid CSF cultures, a decrease in sensitivity was observed, with only 17.3% of isolates being susceptible, indicating an increased resistance.
In sputum and HVS cultures, the susceptibility rates were 67% and 47%, respectively, suggesting a slightly decreased sensitivity to colistin in these sample types (Table 2) The susceptibility of Pseudomonas aeruginosa isolates to meropenem demonstrated decreased sensitivity. In the current study, the susceptibility rates were 50.2% in pus cultures, 68.3% in sputum cultures, and 49% in HVS cultures. Like meropenem, Pseudomonas aeruginosa isolates exhibited increased resistance to ceftazidime in the current study. The susceptibility rates were 34.1% in pus cultures, 60.4% in sputum cultures, and 50.4% in HVS cultures (Table 3). The present research study highlights an alarming trend of increased resistance and decreased sensitivity of Pseudomonas aeruginosa isolates to amikacin, colistin, meropenem, and ceftazidime.
Table 1 presents the antibiotic sensitivity of Pseudomonas species isolates obtained from pus and sputum cultures, highlighting significant variations in resistance patterns between these sample types. Table 2 focuses on blood and CSF culture isolates, revealing distinct sensitivity profiles that are crucial for understanding the treatment challenges in these critical infections. Table 3 summarizes the antibiotic susceptibility of isolates from high vaginal swab (HVS) samples, providing insights into the resistance trends in genital tract infections. These tables collectively illustrate the dynamic and diverse nature of antibiotic resistance in Pseudomonas species across different clinical samples, underscoring the importance of tailored antimicrobial therapy.
|
Antibiotics |
Pus Culture Isolates (n = 1138) |
Sputum Culture Isolates (n = 584) |
|
Amikacin 30µg |
572 (50.2%) |
456 (78%) |
|
Aztreonam 30µg |
406 (35.6%) |
311 (53.2%) |
|
Ceftazidime 10µg |
389 (34.1%) |
353 (60.4%) |
|
Ciprofloxacin 5µg |
502 (44.1%) |
410 (70.2%) |
|
Colistin |
640 (56.2%) |
333 (67%) |
|
Carbapenems 10µg |
572 (50.2%) |
399 (68.3%) |
|
Sulzone |
473 (40.1%) |
360 (61.6%) |
|
Gentamicin 10µg |
403 (35.4%) |
369 (63.1%) |
|
Polymyxin B 300µg |
931 (81.8%) |
440 (75.3%) |
Table 1: Pseudomonas Species Isolates from Pus and Sputum Cultures
|
Antibiotics |
Blood Culture Isolates (n = 63) |
CSF Culture Isolates (n = 52) |
|
Amikacin 30µg |
34 (53.9%) |
30 (57.6%) |
|
Aztreonam 30µg |
19 (30.1%) |
13 (25%) |
|
Ceftazidime 10µg |
29 (46%) |
13 (25%) |
|
Ciprofloxacin 5µg |
40 (63.4%) |
32 (61.5%) |
|
Colistin |
35 (55%) |
9 (17.3%) |
|
Carbapenems 10µg |
34 (53.9%) |
29 (55.7%) |
|
Sulzone |
26 (41%) |
2 (3.84%) |
|
Gentamicin 10µg |
26 (41%) |
27 (51.9%) |
|
Polymyxin B 300µg |
39 (61.9%) |
45 (86.5%) |
Table 2: Pseudomonas Species Isolates from Blood and CSF Cultures
|
Antibiotics |
HVS Sample Isolates (n = 51) |
|
Amikacin 30µg |
37 (72.5%) |
|
Aztreonam 30µg |
33 (64.7%) |
|
Ceftazidime 10µg |
26 (50.4%) |
|
Ciprofloxacin 5µg |
30 (58.8%) |
|
Colistin |
24 (47%) |
|
Carbapenems 10µg |
25 (49%) |
|
Sulzone |
26 (50.9%) |
|
Gentamicin 10µg |
26 (50.9%) |
|
Polymyxin B 300µg |
35 (68.6%) |
Table 3: Pseudomonas Species Isolates from HVS Samples
To illustrate the trends in antibiotic resistance over the five-year period, we have plotted line graphs for each type of isolate. Figure 1 depicts the sensitivity patterns of Pseudomonas species isolated from pus cultures, showing fluctuations in resistance rates to key antibiotics. Figure 2 focuses on sputum culture isolates, highlighting changes in antibiotic susceptibility that are critical for respiratory infection management. Figure 3 presents data for blood culture isolates, emphasizing the evolving resistance trends in bloodstream infections. Figure 4 shows the sensitivity profiles for CSF culture isolates, providing insights into the treatment challenges in central nervous system infections. Finally, Figure 5 illustrates the resistance patterns in isolates from HVS samples, shedding light on the trends in genital tract infections. These figures collectively underscore the dynamic nature of antibiotic resistance in Pseudomonas species across various clinical specimens.
Figure 3: Sensitivity of Antibiotics in Pus Culture Isolates
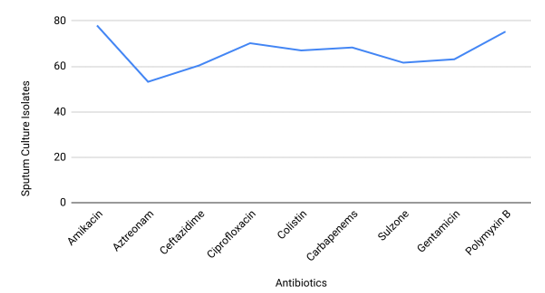
Figure 4: Sensitivity of Antibiotics in Pus Culture Isolates
Figure 5: Sensitivity of Antibiotics in Blood Culture Isolates
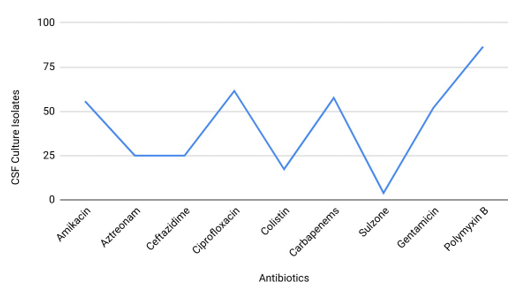
Figure 6: Sensitivity of Antibiotics in CSF Culture Isolates
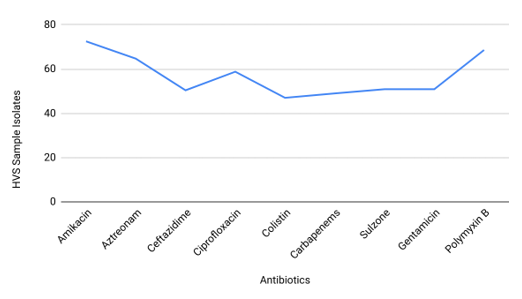
Figure 7: Sensitivity of Antibiotics in HSV Sample Isolates
Discussion
Multidrug-resistant (MDR) opportunistic pathogens, such as Pseudomonas aeruginosa, pose a significant threat to immunocompromised patients. These pathogens are capable of causing both acute and long-term infections, which can be challenging to treat and may lead to serious complications. Since Pseudomonas aeruginosa mutates quickly and adapts to become resistant to medicines, treating an infection with it can be quite challenging. Additionally, because Pseudomonas aeruginosa prefers to grow on moist surfaces, it is among the top-ranked organisms that cause hospital-acquired infections. These infections are frequently discovered in medical devices (ventilation). As per WHO data, it is one of the "critical" pathogen families that immediately need new antibiotics in clinics [12]. Based on a review of published papers, the prevalence of Pseudomonas aeruginosa bloodstream infections is thought to range from 1% to 4% of all bloodstream infections [12].
The incidence, however, might be higher in some patient populations, such as those undergoing invasive procedures or those with weakened immune systems. Data analysis of this study shows the prevalence of aeruginosa isolates to be 3% of all positive blood cultures, which is practically identical to a recent study conducted in Brazil by M.C. Melo in 2021, which reported that Pseudomonas aeruginosa was found in 4.60% of positive blood culture isolates [12]. The same study also focused on the sensitivity patterns of Pseudomonas aeruginosa towards common antibiotic groups like cephalosporins, monobactams, aminoglycosides, and fluoroquinolones.
In light of the analysis conducted by Dana Carmen Zaha in 2019, the antibiotic sensitivity pattern of Pseudomonas aeruginosa isolates obtained from the ICU came out to be as follows: amikacin (56.8%), colistin (99%), meropenems (55.1%) and gentamicin (43%) [13]. However, this study revealed the decreasing sensitivity of Pseudomonas aeruginosa towards amikacin (53.9%), colistin (55%), meropenems (53.9%), and gentamicin (41%) when isolated from blood cultures. Research conducted in Bangladesh from 2019 to 2020 revealed that gram-positive and gram-negative bloodborne bacteria showed higher susceptibility to cefepime and meropenem, contrary to the claim of this study analysis, decreased sensitivity to these antibiotics over time [14]. A total of 1888 cases from distinct clinical specimens tested positive for Pseudomonas aeruginosa. There were 1138 Pus isolates out of a total of 1888 clinical isolates, or 60 %, which is almost equal to the study done in Peshawar, Pakistan, where they showed isolation from pus samples to be 68.6% [15].
The results of this study showed that Pseudomonas aeruginosa isolated from pus cultures showed a decreased antibiotic sensitivity pattern as amikacin (50.2%), ciprofloxacin (44.1%), gentamicin (35.4%), colistin (56.2%) and ceftazidime (34.1%). Compared to a study conducted in Nepal in 2021, where the sensitivity pattern of Pseudomonas aeruginosa towards antibiotics from pus cultures was amikacin (92.85%), ciprofloxacin (7.14%), gentamicin (90.2%), colistin (100%), and ceftazidime (43%) [16]. The sensitivity of Pseudomonas aeruginosa isolates to amikacin (74%), gentamicin (34%), ciprofloxacin (47%), ceftazidime (49%), and imipenem (87%) was also reported in a 2017 publication [17].
The above findings contrasted with the sensitivity found in this study, which involved amikacin (50.2%), ciprofloxacin (44.1%), gentamicin (35.4%), colistin (56.2%), and ceftazidime (34.1%), suggesting an increase in antibiotic resistance. Pseudomonas aeruginosa was found in 38% of sputum samples in a study conducted in Saudi Arabia in 2016 [18]. The Pseudomonas aeruginosa isolates from sputum samples were found to be 46.5% in a study conducted in Nepal. The same survey also showed Pseudomonas aeruginosa sensitivity patterns to standard antibiotics such as amikacin (91.5%), meropenem (78.5), ceftazidime (51.5%), and ciprofloxacin (69.50%) [19]. Comparatively, in this study, Pseudomonas aeruginosa in sputum culture has a positivity rate of 33%. The sensitivity pattern, however, was markedly reduced for amikacin (78%), meropenem (68.3%), ceftazidime (60.4%), and ciprofloxacin (70.2%). In accordance with the study done in Peshawar, the sensitivity of Pseudomonas aeruginosa isolates found in sputum samples towards antibiotics is amikacin (92.96%), meropenems (91.55%), ceftazidime (71.01%), ciprofloxacin (66.2%) and gentamicin (74.65%) in comparison to the sensitivity outcome in this study where for amikacin (78%), meropenem (68.3%), ceftazidime (60.4%) and ciprofloxacin (70.2%) showing a decreased effectiveness of Pseudomonas aeruginosa to these antibiotics [20].
In light of a study conducted by M.S. Raza in a tertiary care hospital in India, the prevalence of Pseudomonas aeruginosa infections in patients suffering from meningitis accounts for 8.33%, which is higher than the incidence found in this study (3%). The same research shows that colistin (100%) and meropenems (60%) are the most effective antibiotics against Pseudomonas aeruginosa infections in meningitis [21]. An investigation of patient’s data in China in 2021 showed Pseudomonas aeruginosa susceptibility pattern to be amikacin (93.8%), meropenems (84.6%), ceftazidime (68.8%), aztreonam (50%) and gentamicin (84.6%) , whereas the analysis of this srudy showed low sensitivity of Pseudomonas aeruginosa to the following antibiotics; Amikacin (55.7%), meropenems (57.6%), ceftazidime (25%), aztreonam (25%) and gentamicin (51.9%). In 2021, an investigation was carried out in India that validates the results of this study which dictates the decreasing sensitivity of Pseudomonas aeruginosa to various antibiotics with sensitivity percentages as amikacin (84.4%), imipenem (93.8%), ceftazidime (74.4%), aztreonam (87.5%) and gentamicin (73.1%) 22,23.
The frequency of Pseudomonas aeruginosa in high vaginal swabs (HVS) in this study accounts for 3%, which is superior to the HVS samples (0.67%) isolated in another study conducted in a tertiary care hospital in southwest Nigeria in 2021 [24]. Analysis of data from a tertiary clinical setting in Bangalore, India, in 2015 shows amikacin, meropenems, tigecycline, ceftazidime, and ciprofloxacin are 40%, 82.65%, 20%, 58%, and 42% effective respectively [25]. In contrast, this study shows a changing sensitivity pattern of Pseudomonas aeruginosa to amikacin (72.5%), meropenems (68.3%), tigecycline (9.8%), ceftazidime (50.4%) and ciprofloxacin (58.8%). The medications mentioned above are no longer effective in treating Pseudomonas aeruginosa infections due to their development of resistance. Multidrug- resistant Pseudomonas aeruginosa infections (MDR-PA) comprise about 13% of all cases.
Even healthy people can have minor infections like rashes and ear infections, but critically ill patients are more likely to get significant infections. Historically, MDR/XDR-Pseudomonas aeruginosa has been treated empirically with conventional antipseudomonal- lactam medicines. These conventional treatments are frequently ineffective due to Pseudomonas aeruginosa's extraordinary ability to transmit resistance via several routes [26]. The extensive use of antibiotics creates problems for sustainability, underscoring the demand for long-term research to track antimicrobial sensitivity and develop local treatment standards. The proper prescription of antibiotics, rapid monitoring of patient responses, and implementation of appropriate medication substitution or decrease are all essential components of responsible antibiotic stewardship. Further research is advised to improve antibiotic resistance surveillance and prompt reporting.
Conclusion
This study investigated patterns in Pseudomonas species to isolate antibiotic sensitivity across various sample types, revealing alarming resistance to amikacin, colistin, meropenem, and ceftazidime compared to prior studies. Based on the results presented in the tables and figures, we observed that Pseudomonas species isolates from pus cultures showed significant antibiotic resistance, particularly low sensitivity to ceftazidime and colistin. CSF culture isolates demonstrated a marked increase in resistance, particularly to colistin, highlighting the treatment challenges for central nervous system infections. Isolates from HVS samples showed moderate sensitivity to most antibiotics, with notable resistance to colistin and carbapenems, indicating a need for targeted infection control measures. These findings highlight the requirement for thorough monitoring and treatment strategy adjustment to treat Pseudomonas species infections successfully. The declining susceptibility to amikacin, ceftazidime, meropenem, and colistin emphasizes the necessity of careful pharmaceutical administration.
Limitations
This study may be subjected to some limitations. Access to this hospital is restricted to hospital staff, retired military members, and their families. As a result, patients are more likely to be women than men. The study will be biased because more females and kids than men are participating. We have data going back two and five years. There can be variations in the findings due to the limited sample size.
References
- Laborda, P., Martinez, J. L., & Hernando-Amado, S. (2022). Evolution of habitat-dependent antibiotic resistance in Pseudomonas aeruginosa. Microbiology Spectrum, 10(4), e00247-22.
- Reynolds, D., & Kollef, M. (2021). The epidemiology and pathogenesis and treatment of Pseudomonas aeruginosa infections: an update. Drugs, 81(18), 2117-2131.
- Horcajada, J. P., Montero, M., Oliver, A., Sorlí, L., Luque, S., Gómez-Zorrilla, S., ... & Grau, S. (2019). Epidemiology and treatment of multidrug-resistant and extensively drug-resistant Pseudomonas aeruginosa infections. Clinical microbiology reviews, 32(4), 10-1128.
- Alhussain, F. A., Yenugadhati, N., Al Eidan, F. A., Al Johani, S., & Badri, M. (2021). Risk factors, antimicrobial susceptibility pattern and patient outcomes of Pseudomonas aeruginosa infection: A matched case-control study. Journal of infection and public health, 14(1), 152-157.
- Pokharel, K., Dawadi, B. R., Bhatt, C. P., & Gupte, S. (2019). Prevalence of Pseudomonas aeruginosa and its antibiotic sensitivity pattern. Journal of Nepal Health Research Council, 17(01), 109-113.
- Shah, D. A., Wasim, S., & Abdullah, F. E. (2015). Antibiotic resistance pattern of Pseudomonas aeruginosa isolated from urine samples of Urinary Tract Infections patients in Karachi, Pakistan. Pakistan journal of medical sciences, 31(2), 341.
- Shah, D. A., Wasim, S., & Abdullah, F. E. (2015). Antibiotic resistance pattern of Pseudomonas aeruginosa isolated from urine samples of Urinary Tract Infections patients in Karachi, Pakistan. Pakistan journal of medical sciences, 31(2), 341.
- Tarazi, Y. H., Abu-Basha, E., Ismail, Z. B., & Al-Jawasreh, S.(2021). Antimicrobial susceptibility of multidrug-resistant Pseudomonas aeruginosa isolated from drinking water and hospitalized patients in Jordan. Acta tropica, 217, 105859.
- Khan, M. I., Xu, S., Ali, M. M., Ali, R., Kazmi, A., Akhtar, N., ... & Li, F. (2020). Assessment of multidrug resistance in bacterial isolates from urinary tract-infected patients. Journal of Radiation Research and Applied Sciences, 13(1), 267-275.
- Al-Ahmadi, G. J., & Roodsari, R. Z. (2016). Fast and specific detection of Pseudomonas Aeruginosa from other pseudomonas species by PCR. Annals of burns and fire disasters, 29(4), 264.
- Khan, A., Pettaway, C. H., Dien Bard, J., Arias, C. A., Bhatti, M. M., & Humphries, R. M. (2021). Evaluation of the performance of manual antimicrobial susceptibility testing methods and disk breakpoints for Stenotrophomonas maltophilia. Antimicrobial agents and chemotherapy, 65(5), 10-1128.
- Melo, M. C., Carvalho Neto, A. P. M., Maranhão, T. L. G. Q.,Costa, E. S., Nascimento, C. M. A., Cavalcanti, M. G. S., ... & Rocha, T. J. M. (2021). Microbiological characteristics of bloodstream infections in a reference hospital in northeastern Brazil. Brazilian Journal of Biology, 84, e253065.
- Zaha, D. C., Bungau, S., Aleya, S., Tit, D. M., Vesa, C. M., Popa, A. R., ... & Aleya, L. (2019). What antibiotics for what pathogens? The sensitivity spectrum of isolated strains in an intensive care unit. Science of the total environment, 687, 118- 127.
- Zerin, Tamanna & Islam, Ariful & Gulnahar, Sabera & Farjana, Noor & Begum, Mosa.Asia. (2021). Identification and Antibiotic Susceptibility of Blood Culture Isolates from Rajshahi, Bangladesh.
- Abbas, S. H., Naeem, M., Adil, M., Naz, S. M., Khan, A., & Khan, M. U. (2015). Sensitivity patterns of Pseudomonas aeruginosa isolates obtained from clinical specimens in Peshawar. Journal of Ayub Medical College Abbottabad, 27(2), 329-332.
- Paudel, P., Shrestha, S., Poudel, S., & Tiwari, B. R. (2021). Multi-drug resistant bacteria isolated from pus and phenotypic detection of metallo β-lactamase activity of Pseudomonas aeruginosa. Nepal Journal of Health Sciences, 1(1), 8-15.
- Rai, S., Yadav, U. N., Pant, N. D., Yakha, J. K., Tripathi, P. P., Poudel, A., & Lekhak, B. (2017). Bacteriological profile and antimicrobial susceptibility patterns of bacteria isolated from pus/wound swab samples from children attending a tertiary care hospital in Kathmandu, Nepal. International journal of microbiology, 2017(1), 2529085.
- Ahmed, O. B. (2016). Incidence and antibiotic susceptibility pattern of pseudomonas aeruginosa isolated from inpatients in two Tertiary Hospitals. Clin Microbiol, 5(2), 2-4.
- Bhatta, S., Pradhan, M., Singh, A., Chaudhary, R., & Singh, Y.(2020). Antimicrobial Sensitivity Pattern of Pseudomonas Aeruginosa Isolated from a Tertiary Care Hospital. Medical Journal of Shree Birendra Hospital, 19(2), 70-74.
- Samad, A., Ahmed, T., Rahim, A., Khalil, A., & Ali, I. (2017). Antimicrobial susceptibility patterns of clinical isolates of Pseudomonas aeruginosa isolated from patients of respiratory tract infections in a Tertiary Care Hospital, Peshawar. Pakistan journal of medical sciences, 33(3), 670.
- Raza, M. S., Das, B. K., Goyal, V., Lodha, R., Chaudhry, R., Sood, S., ... & Kapil, A. (2019). Emerging multidrug resistance isolates of hospital-acquired bacterial meningitis in a tertiary care centre in North India. Journal of medical microbiology, 68(11), 1585-1590.
- Peng, X., Zhu, Q., Liu, J., Zeng, M., Qiu, Y., Zhu, C., ... &Collaborative Working Group of the Pediatric Subgroup of the China Society of Infectious Diseases. (2021). Prevalence and antimicrobial resistance patterns of bacteria isolated from cerebrospinal fluid among children with bacterial meningitis in China from 2016 to 2018: a multicenter retrospective study. Antimicrobial Resistance & Infection Control, 10(1), 24.
- Ali, M. M., Kaleem, F., Farwa, U., Ali, H., Ishtiaq, S., Javed, S., ... & Abbasi, S. A. (2023). Bacterial Isolates and their Sensitivity Profile from CSF Samples–A 5 Year Study at a Tertiary Care Hospital. medRxiv, 2023-03.
- Adejobi, A., Ojo, O., Alaka, O., Odetoyin, B., & Onipede,A. (2021). Antibiotic resistance pattern of Pseudomonas spp. from patients in a tertiary hospital in South-West Nigeria. Germs, 11(2), 238.
- Narayana-Swamy, N., Ramalingappa, P., & Bhatara, U. (2015). Antimicrobial sensitivity pattern of microorganisms isolated from vaginal infections at a tertiary hospital in Bangalore, India. International Journal of Medical Students, 3(1), 34-39.
- Kunz Coyne, A. J., El Ghali, A., Holger, D., Rebold, N., & Rybak, M. J. (2022). Therapeutic strategies for emerging multidrug-resistant Pseudomonas aeruginosa. Infectious diseases and therapy, 11(2), 661-682.



