Research Article - (2024) Volume 2, Issue 2
Antimicrobial Susceptibility Profile of Salmonella, and Escherichia Coli K99 from Diarrheic Calves in and Around Awash Fentale District of Afar Region, Eastern Ethiopia
2Department of Microbiology, Immunology and Veterinary Public Health, Addis Ababa University, Ethiopia
3Department of Microbiology and Veterinary Public Health, Jigjiga University, Ethiopia
Received Date: Jan 08, 2024 / Accepted Date: Feb 01, 2024 / Published Date: Feb 09, 2024
Copyright: ©Â©2024 Dek Kahin Yosef, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Yosef, D. K., Tegenaw, H. G., Adem, A. A. (2024). Antimicrobial Susceptibility Profile of Salmonella, and Escherichia Coli K99 from Diarrheic Calves in and Around Awash Fentale District of Afar Region, Eastern Ethiopia. Int Internal Med J, 2(2), 01-24.
Abstract
Background: The global community is confronted with a significant challenge in the form of antimicrobial resistance, which affects both public and animal health. This phenomenon is greatly influenced by the widespread use of antimicrobial agents in various domains, including human medicine, veterinary practices, and other sectors. The increasing prevalence of drug resistance in E. coli and Salmonella spp. strains isolated from humans and animals has sparked a vigorous debate concerning the potential risks to public health associated with the utilization of antibiotics in animal production. Hence, the objective of this research was to assess the susceptibility of Salmonella enterica to antimicrobial agents and to detect the presence of Escherichia Coli K99 in diarrheic calves within the Awash Fentale District of the Afar Region, in Eastern Ethiopia.
Objectives: To assess antimicrobial susceptibility profile of Salmonella enterica, and to identify Escherichia Coli K99 from Diarrheic Calves in and Around Awash Fentale District of Afar Region, Eastern Ethiopia.
Methods: The current cross-sectional study was conducted from December 2022 to November 2023 with the aim to isolate and identify E. coli K99 and Salmonella enterica and associated risk factors for occurrence of these two pathogens. Fecal samples were collected from a total of 188 diarrheic calves aged up to 6 months from 150 households in Awash Fentale district Afar region, eastern Ethiopia. The samples were transported in ice box to Aklilu Lemma Institute of Pathobiology laboratory, microbiology department for isolation and identification of bacteria. Feces were stored at 4 °C until the time of processing. These samples were processed during the study period for the isolation and identification of E.coli K99 and Salmonella enterica using culture media, biochemical tests, Pathasure kit and Biolog GEN III microplate. Data after laboratory test result was recorded in Microsoft excel® 2021 from prepared result recording format paper. The data was then imported to the software STATA windows version 14 (StataCorp LP, College Station, Texas, USA) for appropriate statistical analysis. Pearson Chi-square (X2) test followed by multivariable logistic regression analysis was utilized to assess the degree of association between each risk factor and calf diarrhea causing E.coli K99 and Salmonella enterica. An association was regarded as significant if the p-value is <0.05.
Result: Out of 188 samples, 20.74% and 5.32% were found to be positive for E.coli K99 and Salmonella enterica respectively. Antimicrobial sensitivity patterns of Salmonella enterica were also investigated using 12 commonly used antibiotics. All isolates from calves were susceptible (100%) to amikacin, chloramphenicol, ampicillin, amoxicillin+clavulanic acid, ceftriaxone, cephalothin ciprofloxacin, tetracycline, gentamycin and trimethoprim+sulfamethoxazole. Multivariable logistic regression analysis was done to identify significant risk factors associated with examined bacteria. Thus, study showed that the occurrence of E. coli K99 in male calves was higher (25.27%) than in female calves (16.49%). This might be the reason that more care is given to female calves than male calves. Because female calves are needed by the pastoralist were compared with male calves for milk production. On the other hand, age and colostrum feeding within 24 hours were significantly associated with Salmonella enterica shedding. Calf age was significantly associated with Salmonella enterica (P=0.014, OR=13.15, 95% CI=1.677-103.127). The highest occurrence of Salmonella enterica was recorded in the 61-120 days of age (13.79%) than 1-60 days of age (2.86%) and 121-180 days of age (10.53%) diarrheic calves.
Conclusion: The isolation and identification of bacteria, as well as the assessment of risk factors linked to the transmission of bacteria causing diarrhea, can prove beneficial in developing effective strategies for prevention and control.
Keywords
Calf Diarrhea, E. Coli K99, Salmonella Enterica, Culture, Isolation, Risk Factors, Awash Fentale.
Background
Antimicrobial agents are essential drugs for human and animal health and welfare. The World Organization for Animal Health (OIE) recognizes the need for access to antimicrobial agents in veterinary medicine. Antimicrobial agents are essential for treating and controlling infectious diseases in animals [1]. Antimicrobial agents are considered popular to fight diarrhea in calves. Nevertheless, their wide spectrum of activity, the emergence of microbial tolerance of different antimicrobial agents has become a well-known phenomenon, which represents a major concern [2].
Antimicrobial resistance is a global public and animal health concern that is influenced by the usage of antimicrobial agents in humans, animals and elsewhere [1]. The historical and growing emergence of drug resistance among E. coli and Salmonella spp. strains isolates from humans and animals has increased the debate on public health hazard associated with the use of antibiotics in animal production [3].
Antimicrobial resistance is a global problem in general, but it might be more severe in Ethiopia where there is lack of antimicrobial resistance assessments of Salmonella and lack of rigorous regulations, but there is easy access of antimicrobials for purchase of people without prescription and incomplete treatment courses as the result of patient non-compliance [4]. The historical and growing emergence of drug resistance among E. coli and Salmonella spp. strains isolates from humans and animals has increased the debate on public health hazard associated with the use of antibiotics in animal production [5].
The spread of multiple antimicrobial-resistant pathogenic bacteria has been recognized by the World Organization for Animal Health (OIE), the Food and Agriculture Organization (FAO) and the World Health Organization (WHO) as a serious global human and animal health problem. The development of bacterial antimicrobial resistance is neither an unexpected nor a new phenomenon. It is, however, an increasingly troublesome situation because of the frequency with which new emerging resistance phenotypes are occurring among many bacterial pathogens and even commensal organisms [1].
In Ethiopia, a 30% pre-weaning calf mortality rate was reported in mixed crop- livestock production systems in Amhara Region and an 18% mortality rate was found in market-oriented dairy farms in Central Ethiopia [6,7]. According to Asmare and Kiros, 2016; the overall morbidity and mortality recorded are 66.7 % and 20 %, respectively.
Diarrhea is one of the most important causes of morbidity and mortality, leading to economic losses due to the cost of treatment, prophylaxis, increased susceptibility to other infections, reduced growth rates, and death of calves Worldwide [8]. The main causes of calf diarrhea are grouped into two categories: non-infectious cause and infectious causes. Infectious diarrhea is the most significant cause of morbidity and mortality in neonatal dairy calves throughout the world The important viral infectious agents include bovine rotavirus group A, bovine corona virus, and bovine viral diarrhea virus.
Among bacterial causes, Salmonella Typhyimurium, Escherichia coli K99+ as “white scour”, and Clostridium perfringens type C and among protozoa, Cryptosporidium parvum and coccidiosis either singly or in combination are affected young calves [9].
In Ethiopia, S. Typhimurium appears to be more important than others, because it was one of the dominant isolates in cattle (17.4%) (10). Several risk factors for calf diseases, particularly diarrhoea, have been identified, including farm size, presence of a calving pen and hygiene, the quality of colostrum and the route of colostrum feeding as well as type of calf housing [11]. The applications of false drugs and feed additives also play a very important role in predisposition of the etiology in adult calf diseases.
the World Health Organization (WHO), diarrheal diseases are the second leading cause of death (~760,000 per year) in children less than 5 years of age [12]. Diarrhea may be infectious caused by bacteria, viruses or parasites [13].
Human salmonellosis is one of the most common and economically important zoonotic diseases. The Centers for Disease Control and Prevention (CDC) estimates that salmonellosis causes more than 1.2 million illnesses each year in the United States of America with more than 23,000 hospitalizations and 450 deaths [14]. The most common cause of infection with Salmonella is eating contaminated foods including raw or under cooked eggs or egg products, meat, poultry, contaminated fresh fruit and vegetables and soft cheeses made from unpasteurized milk [15]. In developing countries including Ethiopia up to 2 million people die per year due to disease of foodborne pathogens. Over the last 20 years, the emergence of major foodborne pathogens such as Salmonella and E. coli have persisted as a major public health concern and provide clear examples of the persistence of foodborne pathogens despite considerable efforts aimed at prevention and control [16].
Ethiopia ranks second after Nigeria in the health burden of zoonotic diseases in Africa. This country is located in a sub-region that experiences the second highest foodborne disease burden in the world, where E. coli O157 is one of the leading causes of foodborne disease disability adjusted life years. In Ethiopia, years of life lost due to diarrheal diseases was 2.6 million and diarrheal diseases are the second leading cause of premature death after lower respiratory infections. A study conducted on diarrheic children under 5 years in northern part of Ethiopia, E. coli O157:H7 was 28.9% [17].
Although there are some research works carried out in certain parts of the country, on problem of calf mortality and its economic significance, information on calf morbidity and mortality as result of diarrhea caused by the two important enteric bacteria, E.coli K99 and Salmonella enterica is limited in study area, in and around Awash Fentale district. Hence antimicrobial susceptibility profile of salmonella, and Escherichia Coli K99 from diarrheic calves is needed for planning a proper control and preventive measure in the area as well as in country.
Study Area
The study was conducted in Awash Fentale district, zone 3, Afar Regional State of Ethiopia from December 2022 to November 2023. The zone is located in the part of the middle Awash basin about 225 km north east of Addis Ababa. Geographically, the study area Awash Fentale located between 8052â??to 9024â??North latitude and 39040â??to 400 20â??East longitudes (Figure 1). The altitude range is from 743 – 1387m above sea level and the majority of the land is rocky with annual rain fall range 150–500mm per year. The average daily ranges of minimum and maximum temperature are 250C and 390C, respectively.
Awash Fentale is bordered on the south by the Oromia Region, on the west by the Amhara Region, on the north by Dulecha, and on the east by Amibara. The district has 5 kebeles with total population 29,076 within 5101 households [18]. Towns in Awash Fentale include Awash Sebat Kilo and Sabure. Rivers in this woreda include the Awash and its tributary the Germama. A large portion of this woreda is occupied by the Awash National Park. The soil texture of the area is characterized under sandy loam which accounts 55% sand, 40% silt (mud) and 5% of clay. The water holding capacity of such soil is very poor [19]. The dry season is from May to June while the main rainy season extends from July to September accounts for above 60% of the annual total rainfall. The livestock populations of the Awash Fentale district are composed of cattle 86085, sheep 35389, goat 83691, horses, asses 45260, mules 4355, camels 41245 and poultry 4355 .
Figure 1: Map of Study Area, Diduba, Diho and Kebena in Awash Fentale district (Map Constructed Using Arc Version 10.4 GIS Software, 2015)
Study Animals
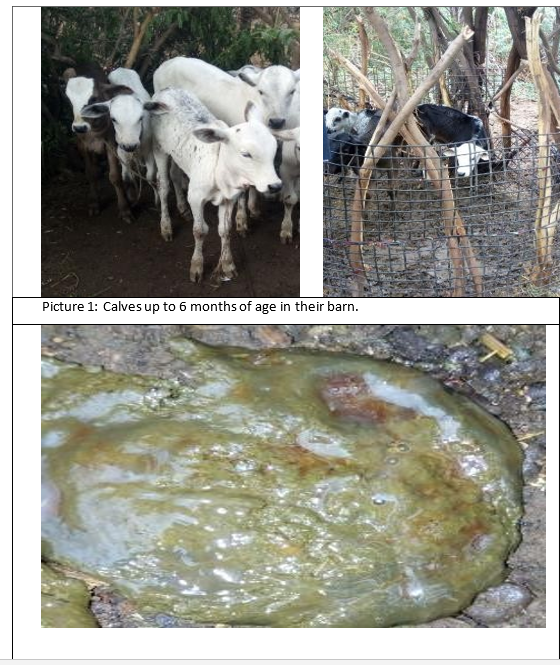
Calves of both sexes up to 6 months old affected with diarrhea and exhibiting clinical signs were included in the study. These calves showed, but had not been treated with antibiotics before. The study calves were local breeds, kept under traditional extensive management system from three selected Kebele namely Diho, Diduba and Kebena.
Study Design and Sampling Method
A cross-sectional study design was undertaken to isolate and identify E.coli K99 and Salmonella enterica from calvesâ?? diarrhea and assess associated risk factors. Individual animals were carefully identified and sex, age, species were recorded. Purposive type sampling method was conducted to select households and calf sample. The sample size was determined based on the availability of diarrheic calf (case). 50 households from each three Kebele were selected purposively by the “The project Addressing causes of young stock mortality in Ethiopia” based by considering on accessibility to vehicle or proximity to road and distribution of study animal population. Accordingly, a total of 188 fecal samples from diarrheic calves were collected from 150 households in 3 Kebeles namely Diho, Diduba and Kebena in Awash Fentale district. Then collected samples were confirmed on the basis of their, culture, pathasure kit, biochemical tests, Biolog and antimicrobial testing using standard bacteriological procedures described by [15,20].
For the questioner survey, questionnaire was administered and about 50 households were interviewed as calves taken for sample collection to gather information about risk factors to exposure of E.coli K99 and Salmonella enterica.
Sample Collection and Transportation
Fecal samples were aseptically collected from the rectum of each the calves with sterile hand glove those showing clinical manifestations of calf diarrhea by rectal stimulation while avoiding environmental contamination. Approximately 10 ml of fecal samples were collected from non-treated diarrheic calves directly from the rectum by using disposable plastic gloves and transferred immediately to sterile plastic container. Gloves were changed between calves to reduce the risk of contamination during sample collection which will affect the result after laboratory diagnosis. The containers were labeled legibly using permanent marker with individual identifier and placed in ice box. The samples were protected from light, extreme temperatures and desiccation. The samples were transported in ice box to Aklilu Lemma Institute of Pathobiology laboratory, microbiology department for isolation and identification of bacteria. Feces were stored at 4 °C until the time of processing [21].
Questionnaire Survey
A structured questionnaire was administered to calvesâ?? owners to assess the general calf husbandry practices of the households. Management as well as herd and calf-level risk factors associated with diarrhea causing bacteria, E.coli K99 and Salmonella enterica were observed and assessed during sample collection. Questionnaire formats and checklists were developed. The checklist was contained household ID, young stock ID (code number of the calf), sex, age and diarrhea type. Thus, the information that was included in the questionnaires were colostrum feeding, age of dam at parturition, dam gestation number (parity), area where the animal lies down look clean and dry, animal housed on bedding, general health care, herd size, occurrence of calf diarrhea as well as disease control measures practiced in the herd. The format was filled directly by face-to-face interviewing (Annex I).
Laboratory Investigations
Isolation and Identification of E.coli K99
Antigen Test Kit (ELISA) for the Detection of Escherichia coli K99
Pathasure Enteritis 4 is a sensitive and specific diagnostic kit based on the ELISA technology. The presence of infectious agents in tested samples is detected using highly specific antibodies and is revealed by means of a colour reaction easily detected by eye. Homogenizing the concentrated wash solution, it was diluted at 1/10 with purified water. Fecal samples were diluted in 1X wash solution at 1/10 and each dilution was properly mixed before being distributed into the wells. Then diluted fecal samples were incubated in wells coated with a mixture of 4 antibodies (Abs) specific to rotavirus, coronavirus, E. coli K99 and Cryptosporidium parvum at 23 ± 2°C for 30 minutes. After washing each well 5 times with wash solution to eliminate unbound substances, a conjugate (an antibody coupled to an enzyme) targeted at rotavirus, coronavirus, E. coli K99 and Cryptosporidium parvum was added. After incubation at 23 ± 2°C for 30 minutes, excess of this conjugate was eliminated by a second wash and its attachment to the specific pathogen was revealed with a chromogenous substrate. Following incubation period at 23 ± 2°C for 10 minutes, the enzyme reacted with the substrate and a color was developed. The intensity of the color allowed the determination of the type of sample tested (Annex II). A negative sample was displayed a weak reaction (colorless, pale blue) whereas a positive sample showed a reaction stronger than the one for the negative control (red, darker blue, green and violet than negative control).
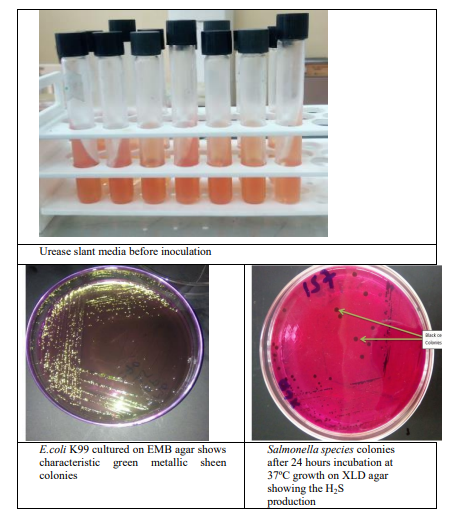
Formerly positive fecal samples in ELISA test were plated directly on to buffered peptone water. Then incubated feces to minimize overgrowth by other organisms were 6 hours at 37°C. Enrichment broths were pre-warmed to prevent cold-shocking the organisms and slowing their initial growth. Selective culture was done by inoculating on MacConkey agar containing 1% D-sorbitol. MacConkey agar with sorbitol was used because inexpensive medium on which no sorbitol fermenting E. coli grow as small, round greyish-white colonies[15]. Medium size and bright ink to red with flat or elevated surface and complete weight edge colonies was seen on MacConkey agar [15,20]. Media were inoculated in a logical order from least selective to most selective to avoid the inhibition of organisms to the selective agent. First indicator media MacConkey and second selective media Eosin methylene blue agar were used and bright pink, green metallic sheen colonies were observed respectively (Annex X, Figure 2).
Isolation and Identification of Salmonella Enterica
The conventional bacteriological methods were used to isolate Salmonella enterica from faecal samples. All media were prepared according to manufacturerâ??s direction.
• Non-selective pre-enrichment: The fecal samples at ratio of 1:10 were inoculated in 0.1% buffered peptone solution and incubated at 370C for 18-20 hours for pre- enrichment.
• First selective enrichment: 0.1 ml and 1 ml of the culture obtained from incubated buffered peptone water broth were transferred to a tube containing 10 ml of the RVS broth and 10 ml MKTTn broth respectively. Then inoculated RVS broth and MKTTn broth were incubated at 41.5 °C, 37 °C for 24 and 48 hours respectively.
Second Selective Enrichment and Plating Out and Identi- fication
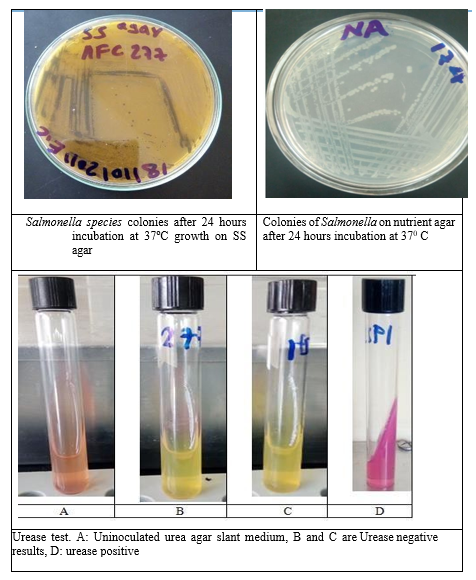
A loop full from the inoculated and incubated RVS and Muller- Kauffmann tetrathionate broth was streaked on XLD and on SSA agar plates and incubated at 37oC for 24 hours. After development of characteristic black centered colony the positives were selected and inoculated onto nutrient agar plates to allow well-isolated colonies at 37 °C for 24 hours (Annex X, Figure 11). Presumptive Salmonella colonies from nutrient agar were phenotypically confirmed by biochemical properties in differential agars, such as TSI, LIA and Simmons citrate after incubation at 37 ºC for 16-24 hours [20].
development of characteristic black centered colony the positives were selected and inoculated onto nutrient agar plates to allow well-isolated colonies at 37 °C for 24 hours (Annex X, Figure 3). Presumptive Salmonella colonies from nutrient agar were phenotypically confirmed by biochemical properties in differential agars, such as TSI, LIA and Simmons citrate after incubation at 37 ºC for 16-24 hours [15,20].
Biochemical Confirmation of Salmonella Enterica
Urease Test: A portion of well isolated colony was streaked on the surface of a urea agar slant to detect the ability of an organism to produce an enzyme urease which splits urea to carbon dioxide and ammonia. Then it was left the cap on loosely and incubated the test tube at 370c in ambient air for up to 24 hours.
Triple Sugar Iron Agar: A straight inoculating needle was used to pick up isolates from culture of isolated colony. The TSI slant was inoculated by stabbing the butt down to the bottom, and then streaked over the surface of the slant back and forth. The TSI slant was then incubated for 16 hours and 24 hours at temperature of 370C. The positive result for Salmonella was detected based on the properties blackening of the medium provided by hydrogen sulphide gas production. The typical reaction for Salmonella in TSI agar, a red (alkaline) slant, yellow(acid) butt and superimposed (black) H2S production (R/Y/H2S+) was observed (Table 3).
Lysine Iron Agar (LIA) Test: Lysine iron agar was inoculated with a straight needle by stabbing to the base of the butt and streaking the slant. The caps of the tubes were replaced loosely so that aerobic conditions prevail on the slant. Then incubated at 37°C in ambient air for 24 hours. After 24 hours incubation period purple slant/black butt, color was observed (Figure 4).
Figure 2: Lysine Iron Agar:-Both A and B are Positive for Salmonella Species
Citrate Utilization Test: Simmons citrate solid slant agar medium was inoculated with Salmonella isolated colony to test for ability to use citrate as carbon source. Then, the Simmons citrate slant was incubated at 350C to 370C for 18 to 24 hours. After 24 hours incubation blue color was observed (Figure 5).
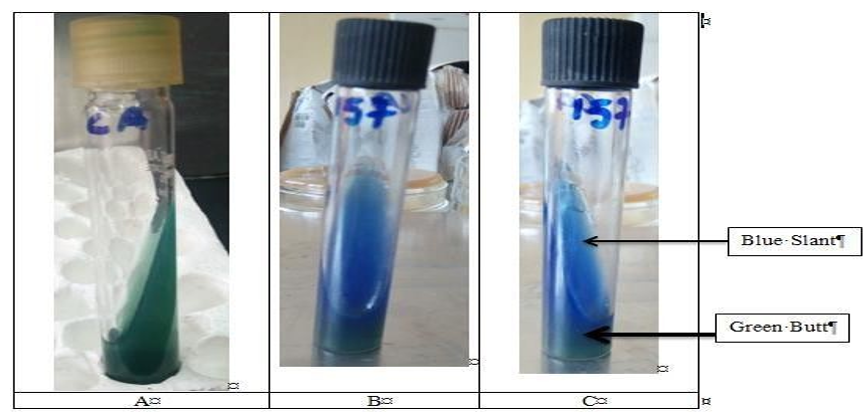

Figure 3: Citrate Utilization Test:-A: Control, B And C Are Salmonella Species
|
S/No. |
Biochemical test |
|
Positive reaction |
|
|
|
|
Color change |
H2S production |
|
|
|
|
Slant |
Butt |
|
|
1. |
TSI |
Red |
Yellow |
Yes, Black color |
|
2. |
LIA |
Purple |
Black |
Yes, Black color |
|
3. |
Citrate utilization test |
Blue |
Green |
Yes, Black color |
|
4. |
Ureases test |
Pale yellow color |
Pale yellow color |
No change in black color |
Table 1: Summary of Biochemical Tests Results used to Identify Salmonella Enterica
Identification of Salmonella Enterica by Using Biolog GEN III Microplate
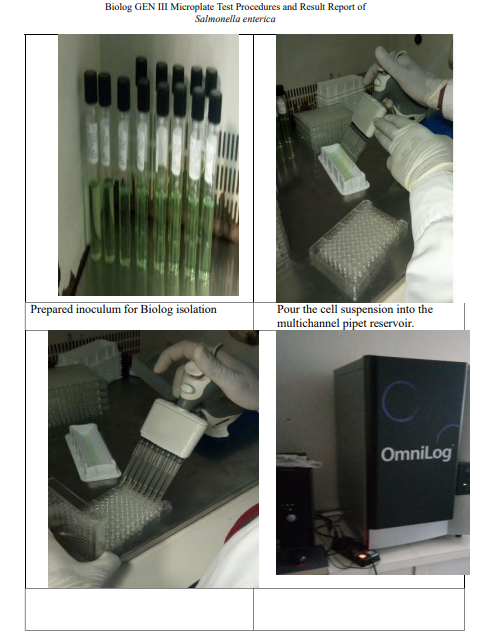
Biolog is powerful carbon source utilization technology accurately identifies environmental and pathogenic microorganisms by producing a characteristic pattern or “metabolic fingerprint” from discrete test reactions performed within a 96 well microplate. The scope of the 96 assay reactions, coupled with sophisticated interpretation software, delivers a high level of accuracy that is comparable to molecular methods. The one minute per sample set up is much simpler and faster than DNA sequencing and the automated pattern matching eliminates the need for training and expertise in gene sequence interpretation. 13 Salmonella isolates were confirmed using biochemical tests from a total of 188 fecal samples. Then these 13 isolates were sent to Ethiopian Biodiversity institute for identification Salmonella enterica by using Biolog GEN III microplate. Agar plates was streaked using correct techniques on Biolog Universal growth (BUG) agar medium to generate well isolated colonies. A single colony grown on BUG agar medium was selected and emulsified into inoculating fluid A. According to the manufacturerâ??s instructions, cell density of the bacterial inoculum was measured for a specified transmittance (90 to 98%) using a turbidimeter, as specified in the user guide. For each isolate, 100 μl of the cell suspension was inoculated in to each of the 96 well coated micro plates, using automatic multichannel pipette and incubated aerobically at 33°C for 22 hours . The OmniLog identification system automatically read each micro plate and provided identification called species/sub-species ID. The result was read in the BIOLOG Micro Station reader after 22 hours incubation outside GEN III incubation then printed (Annex III).
Antimicrobial Susceptibility Testing For Isolated Salmonella Enterica
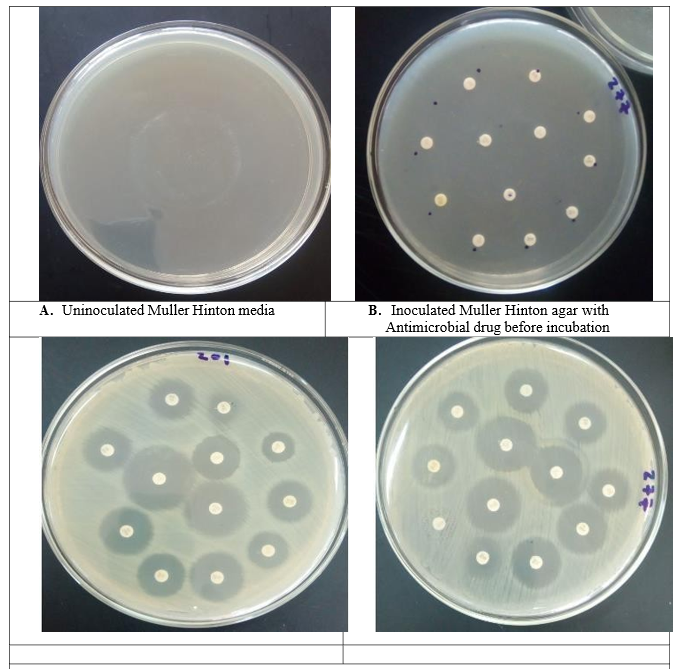
Antimicrobial susceptibility testing of Sal- monella enterica isolates was performed using disc diffusion method on Muller Hinton agar medium according to Office of International Epizootics Terrestrial Manual (OIE, 2012). Results, sizes of the zones of inhibition were interpreted in accordance with Clinical and Laboratory Standards Institute guidelines (CLSI, 2016). The following 12 commonly antibiotic discs were used: ampicillin (10μg), amoxicillin+clavulanicacid (20/10μg), amik- acin (30μg), chloramphenicol (30μg), cephalothin (30μg), ceftri- axone (30μg), ciprofloxacin (5μg), tetracycline(30μg), sulfame- thoxazole+trimethoprim(23.75/1.25μg),sulfisozazole(0.25m g), streptomycin (10μg) and gentamycin (10μg). The antimicrobial susceptibility was determined according to the standard diameter of inhibition for each antibiotic used according to guidelines set by the Clinical and Laboratory Standards Institute (22).
|
Antimicrobial agent |
Disk content |
Zone diameter interpretive criteria (nearest whole mm) |
||
|
S |
I |
R |
||
|
Ampicillin(AM) |
10μg |
≥ 17 |
14–16 |
≤ 13 |
|
Amoxicillin clavulanic |
|
|
|
|
|
acid(AMC) |
20/10 μg |
≥ 18 |
14–17 |
≤ 13 |
|
Amikacin(AN) |
30 μg |
≥ 17 |
15–16 |
≤ 14 |
|
Chloramphenicol(C) |
30μg |
≥ 18 |
13–17 |
≤ 12 |
|
Cephalothin(CF) |
30μg |
≥ 18 |
15–17 |
≤ 14 |
|
Ceftriaxone(CRO) |
30μg |
≥ 23 |
20–22 |
≤ 19 |
|
Ciprofloxacin (CIP) |
5 μg |
≥ 21 |
16–20 |
≤ 15 |
|
Tetracycline(TE) |
30μg |
≥ 15 |
12–14 |
≤ 11 |
|
Trimethoprim(1.25μg)- sulfamethoxazole (SXT) (23.75μg) |
25μg |
≥ 16 |
11–15 |
≤ 10 |
|
Sulfisoxazole (G) |
0.25mg |
≥ 17 |
13–16 |
≤ 12 |
|
Streptomycin(S) |
10μg |
≥ 15 |
12–14 |
≤ 11 |
|
Gentamycin(GM) |
10μg |
≥ 15 |
13–14 |
≤ 1 |
Legend: μg = Microgram, S= Susceptible, I= Intermediate, R= Resistant strains, source (CLSI, 20
Table 2: Antimicrobial Discs, Concentration and Interpretation Standards of their Action on Salmonella Isolate
Salmonella Enterica Serotyping
Salmonella enterica isolates identified by Biolog GEN III microplate were sent to Public Health Agency of Canada, National Microbiology Laboratory at Guelph Reference Service and Laboratory OIE Salmonella Reference Laboratory 110 Stone Rd. West, Guelph, Ontario for Salmonella typing. Serotyping of Salmonella enterica was done from National Microbiology Laboratory at Guelph Reference Service and Laboratory OIE Salmonella Reference Laboratory 110 Stone Rd. West, Guelph, Ontario, and NIG3W4 (519)822-3300 on the basis of their antigen.
Data Management and Analysis
Data which describing the diarrheagenic conditions suggestive of E.coli K99 and Salmonella enterica infection observed on calves along with age, sex, herd size, time of birth, type of delivery, colostrum fed within 24 hours and parity were classified, filtered and coded. Data after laboratory test result was recorded in Microsoft excel® 2010 from prepared result recording format paper. The data was then imported to the software STATA windows version 14 (StataCorp LP, College Station, Texas, USA) for appropriate statistical analysis. Pearson Chi-square (X2) test followed by multivariable logistic regression analysis was utilised to assess the degree of association between each risk factor and calf diarrhea causing E.coli K99 and Salmonella enterica. An association was regarded as significant if the p-value is <0.05.
Ethical Clearance
This study was conducted after obtaining approval from Addis University Aklilu Lemma Institute of Pathobiology “Animal Experiments Ethics Committee”. In the study area there was a proper oral consent with the animal owner or household to collect fecal sample from their animal with the cooperation of veterinarians who give service to the pastoral community in the study site.
Results
Isolation and Identification of Escherichia Coli K99
A total of 188 fecal samples were examined in laboratory for the isolation and identification of bacterium, of which 39 (20.74%) samples were positive for E. coli K99 by antigen test kit (ELISA). Highest percentage of E. coli K99 was found in Diho (29.6%) followed by Diduba (17.9%) and Kebena (13.6%) (Table 6). There was no significant difference (P>0.05) (P= 0.147) among three location for the prevalence of E. coli K99. Among 188 samples 3 of them were infected with both E.coli K99 and Salmonella enterica 1.6 %( 3/188) (Table 3).
|
Number of samples examined |
Both E.coli K99 and Salmonella enterica positive |
|
|
|
No. of positive |
% positive |
|
188 |
3 |
1.6 |
Table 3: Presence of Mixed Infection with both E.coli K99 and Salmonella Enterica in Diarrheic Calves
|
S/No. |
Kebele |
E.coli K99 |
Percentage of positive E.coli K99 |
|
|
Negative |
Positive |
|||
|
1. |
Diho |
38 |
16 |
29.6 |
|
2. |
Diduba |
92 |
20 |
17.9 |
|
3. |
Kebena |
19 |
3 |
13.6 |
|
Total |
|
149 |
39 |
20.7% |
Pearson chi2 (2) = 3.8369, P value= 0.147
Table 4: Frequency Distribution of E.coli K99 Isolated from Fecal Sample of Diarrheic Calves within three Kebeles in Awash Fentale Distric
Isolation and Identification of Salmonella Enterica
Salmonella species were identified using Biolog GEN III microplates, 10 (5.32%) were Salmonella enterica. Among these, Salmonella enterica subspecies enterica, Salmonella enterica subspecies salamae and Salmonella enterica subspecies diarizonae 5(38.46%), 4(30.78%) and 1(7.69%) were isolated respectively (Table 7).
Highest prevalence of Salmonella enterica was found in Diduba (7.1%) followed by Diho (3.7%) (Table 8). No samples were found positive in Kebena. There was no statistically significant difference (P>0.05) (P=0.324) in prevalence of Salmonella enterica subspecies for different locations within three Kebeles in Awash Fentale district. After culturing on media, Salmonella species revealed red colony with black center on MacConkey agar and pale, colorless, transparent, with a black center H2S produced on Salmonella Shigella (SS) Agar. A typical Salmonella colony on XLD agar has a slightly transparent zone of reddish colour and a black centre; a pink-red zone was seen in the medium surrounding the colonies. On the basis of culture and morphological properties all suspected colonies of Salmonella were subjected to selected biochemical tests such as ureases test, citrate utilization test, lysine iron agar and triple sugar iron agar test.
Identification of Salmonella Enterica by Using Biolog GEN III Microplate
After incubation of 13 Salmonella isolates aerobically at 33°C for 24 hours, then Omni Log identification system was automatically read each microplate and provided identification species ID. The result was read in the BIOLOG Micro Station reader then it was printed. Out of 13 Salmonella isolates 10 were positive for Salmonella enterica (Table 5).
|
ated Salmonella enterica |
of positive samples |
centage |
|
nella enterica ss enterica |
|
38.46% |
|
nella enterica ss salamae |
|
30.78% |
|
nella enterica ss diarizonae |
|
7.69% |
|
bactor koseri/youngae |
|
23.8% |
|
otal |
3 |
100% |
Legend: Salmonella enterica ss enterica =Salmonella enterica subspecies enterica, Salmonella enterica ss salamae=Salmonella enterica subspecies salamae and Salmonella enterica ss diarizonae =Salmonella enterica subspecies diarizonae
Table 5: Result of Biolog isolated Salmonella enterica
|
S/No. |
Kebele |
Salmonella enterica |
Percentage of positive Salmonella enterica |
|
|
Negative |
Positive |
|||
|
1. |
Diho |
52 |
2 |
3.7% |
|
2. |
Diduba |
104 |
8 |
7.1% |
|
3. |
Kebena |
22 |
0 |
0 |
|
|
Total |
178 |
10 |
5.32% |
Pearson chi2 (2) =2.2554; P= 0.324
Table 6: Frequency Distribution of Salmonella Enterica Isolated from Fecal Samples of Diarrheic Calves within three Kebeles in Awash Fentale District
|
S/No. |
Kebele |
Salmonella enterica |
Percentage of positive Salmonella enterica |
|
|
Negative |
Positive |
|||
|
1. |
Diho |
52 |
2 |
3.7% |
|
2. |
Diduba |
104 |
8 |
7.1% |
|
3. |
Kebena |
22 |
0 |
0 |
|
|
Total |
178 |
10 |
5.32% |
Pearson chi2 (2) =2.2554; P= 0.324
Salmonella Enterica Serotyping
Serotyping of Salmonella enterica was done from National Microbiology Laboratory at Guelph Reference Service and Laboratory OIE Salmonella Reference Laboratory 110 Stone Rd.West, Guelph, Ontario, and NIG3W4 (519)822-3300 on the basis of their “K” antigen. The 10 Salmonella enterica isolates were serotyped as 8 S. ZANZIBAR (80%) and 2 (20%) unclassified (Table 8).
|
S/No. |
Serotype |
Antigens |
No. |
Percentage |
|
1 |
II:58:k:1,6( not classified) |
58:K:1,6 |
2 |
20 |
|
2 |
ZANZIBAR |
10:K:1,5 |
8 |
80 |
|
|
Total |
|
10 |
100 |
Table 8: Serotyping Frequency Distribution of Salmonella Enterica
Antimicrobial Susceptibility Testing of Salmonella Enterica Isolates
Out of total 10 isolates screened for antimicrobial susceptibility test against twelve antibiotics discs, all isolates from calves were susceptible(100%) to ampicillin, amoxicillin+clavulanic acid, amikacin, chloramphenicol, cephalothin, ceftriaxone, ciprofloxacin, tetracycline, gentamycin and trimethoprim+sulfamethoxazole. Three isolates (30%) and one isolate (10%) were found to be susceptible to streptomycin and sulfisoxazole respectively. On the other hand, six isolates (60%) were found to be resistant for sulfisoxazole, whereas 7 isolates (70%) were intermediate against streptomycin (Table 9).
|
Antibiotics |
Susceptible (n (%)) |
Intermediate (n (%)) |
Resistant (n (%)) |
|
Ampicillin(AM) |
10(100) |
- |
- |
|
Amoxicillin clavulanic acid(AMC) |
10(100) |
- |
- |
|
Amikacin(AN) |
10(100) |
- |
- |
|
Chloramphenicol(C) |
10(100) |
- |
- |
|
Cephalothin(CF) |
10(100) |
- |
- |
|
Ceftriaxone(CRO) |
10(100) |
- |
- |
|
Ciprofloxacin (CIP) |
10(100) |
- |
- |
|
Tetracycline(TE) |
10(100) |
- |
- |
|
Trimethoprim sulfamethoxazole (SXT) |
10(100) |
- |
- |
|
Sulfisoxazole (G) |
1(10) |
3(30) |
6(60) |
|
Streptomycin(S) |
3(30) |
7(70) |
0 |
|
Gentamycin(GM) |
10(100) |
- |
- |
Table 9: Antimicrobial Susceptibility Testing Results of Salmonella Enterica Isolates from Diarrheic Calves (n= 10)
Risk Factors Associated with the Presence of E.coli K99 and Salmonella Enterica
Due to similar management of households in pastoral production system, no statistical comparisons were done for most of the household factors and no relevant association was found to various variables. However, some risk factors associated to the exposure of E.coli K99 and Salmonella enterica are presented in Table 11 and Table 12 below. The positive samples of E. coli K99 and Salmonella enterica in 1-60 days, 61-120 days and 121 -180 days age groups were 31, 5, 3 for E. coli K99 and 4, 4, 2 for Salmonella enterica respectively. Thus the prevalence of E. coli K99 and Salmonella enterica in 1-60 days, 61-120 days and 121- 180 were 22.14%, 17.24% and 15.79 for E. coli K99 and 2.86%, 13.79, 10.53% for Salmonella enterica respectively within the age groups. There was statically significant difference within age groups in the presence of Salmonella enterica infection (P=0.033).
The total number of positive samples in male calves were 23,7 and the prevalence were 25.27%,7.69% and in the case of female calves, the total number of positive samples were 16,3 and the prevalence were 16.49%,3.09% for E. coli K99 and Salmonella enterica respectively. No significant difference between two sexes was found for both bacteria (P>0.05). The prevalence of E. coli K99 was 18.81% (n=19/101) in day time of birth and 20.99% (n=17/81) in night time of birth. The prevalence of E. coli K99 was similar in both time of birth. The p-value was calculated as 0.187 (P>0.05) that mean the results were statistically not significant. In the case of type of delivery, the occurrence of E. coli K99 was 20.93 %( 36/172) in normal delivery and 16.67% (1/6) in assisted delivery. There was no association between type of delivery and occurrence of E.coli K99 in calves (P=0.967).
Colostrum fed within 24 hours was found to affect significantly the occurrence of E.coli K99 (P< 0.05; P=0.000). Thus, 73.33 %( 11/15) of the cases were found to be not fed colostrum within 24 hours whereas 16.18 %( 28/173) were fed colostrum within 24 hours. Continuous access of calf to dam during the first 24 hours significantly reduce the infection rate with E.coli K99 (P < 0.05; P=0.000). Thus, 14 out of 21(66.67%) infected calves did not get continuous access of colostrum to dam during the first 24 hours, whereas 25 of 167(14.97%) calves were have got continuous access of colostrum to dam during the first 24 hours.
Infection with Salmonella enterica was found to be significantly affected by colostrum fed within 24 hours (P< 0.05; P=0.008) (Table 11). Thus, 3 cases (20.00%) were found not fed colostrum within 24 hours, whereas 7 cases (4.05%) were fed colostrum within 24 hours. Significant association was not found between continuous access of calf to dam during the first 24 hours and infection with Salmonella enterica (P>0.05; P=0.362); two calves (9.52%) infected with Salmonella enterica were found not to be accessed the calf to the dam during the first 24 hours, while eight (4.79%) were found to be accessed the calf to the dam during the first 24 hours.
Host associated risk factors with the presence of E.coli K99 and Salmonella enterica in diarrheic calves such as sex, time of birth and type delivery did not affect significantly E.coli K99 and Salmonella enterica infection in the present investigation. On the other hand, dam and environmental associated risk factors in diarrheic calves including age of dam at parturition, parity, herd size and birth location did not affect E.coli K99 and Salmonella enterica infection significantly in the current study.
|
Risk factors |
Identification for |
Category |
No. examined |
Positive |
Percentage positive (%) |
P-value |
|
|
E.coli K99 |
1-60 days |
140 |
31 |
22.14 |
|
|
|
|
61-120 days |
29 |
5 |
17.24 |
|
|
Age |
|
121-180 days |
19 |
3 |
15.79 |
0.17 |
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
1-60 days |
140 |
4 |
2.86 |
|
|
|
|
61-120 days |
29 |
4 |
13.79 |
0.033* |
|
|
|
121-180 days |
19 |
2 |
10.53 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
|
E.coli K99 |
Male |
91 |
23 |
25.27 |
|
|
|
|
Female |
97 |
16 |
16.49 |
0.138 |
|
|
|
Total |
188 |
39 |
20.74 |
|
|
Sex |
Salmonella enterica |
Male |
91 |
7 |
7.69 |
|
|
|
|
Female |
97 |
3 |
3.09 |
0.160 |
|
|
|
Total |
188 |
10 |
5.32 |
|
|
|
E.coli K99 |
Day |
101 |
19 |
18.81 |
|
|
|
|
Night |
81 |
17 |
20.99 |
0.187 |
|
Time of Birth |
|
Donâ??t know |
6 |
3 |
50.00 |
|
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
Day |
101 |
6 |
5.94 |
|
|
|
|
Night |
81 |
3 |
3.70 |
0.362 |
|
|
|
Donâ??t know |
6 |
1 |
16.67 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
|
E.coli K99 |
Normal |
172 |
36 |
20.93 |
|
|
|
|
Assisted |
6 |
1 |
16.67 |
0.967 |
|
Type of delivery |
|
Donâ??t know |
10 |
2 |
20.00 |
|
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
Normal |
172 |
8 |
4.65 |
|
|
|
|
Assisted |
6 |
1 |
16.67 |
0.346 |
|
|
|
Donâ??t know |
10 |
1 |
10.00 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
|
E.coli K99 |
Yes |
173 |
28 |
16.18 |
0.000* |
|
Colostrum fed within 24 hours |
|
No |
15 |
11 |
73.33 |
|
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
Yes |
173 |
7 |
4.05 |
0.008* |
|
|
|
No |
15 |
3 |
20.00 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
Continuous access to dam |
E.coli K99 |
Yes |
167 |
25 |
14.97 |
|
|
during the first 24 hours |
|
|
|
|
|
|
|
|
|
No |
21 |
14 |
66.67 |
0.000* |
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
Yes |
167 |
8 |
4.79 |
|
|
|
|
No |
21 |
2 |
9.52 |
0.362 |
|
|
|
Total |
188 |
10 |
5.32 |
|
*P-value <0.05
Table 10: Host Associated Risk Factors with the Presence of E.coli K99 and Salmonella Enterica
|
Risk factor |
Identification for |
Category |
No. examined |
Positive |
Percentage positive (%) |
P-value |
|
Age of dam at parturition |
E.coli K99 |
3-7 years |
125 |
27 |
21.60 |
|
|
|
|
8-12 years |
58 |
1 |
18.97 |
0.19 |
|
|
|
13-16 years |
5 |
1 |
20.00 |
|
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
3-7 years |
125 |
8 |
6.40 |
|
|
|
|
8-12 years |
58 |
2 |
3.45 |
0.614 |
|
|
|
13-16 years |
5 |
0 |
0.00 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
Parity |
E.coli K99 |
< 3 |
96 |
21 |
21.88 |
|
|
|
|
3 |
37 |
6 |
16.22 |
0.750 |
|
|
|
>3 |
55 |
12 |
21.82 |
|
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
< 3 |
96 |
8 |
8.33 |
|
|
|
|
3 |
37 |
0 |
0.00 |
0.127 |
|
|
|
>3 |
55 |
2 |
3.64 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
Herd size |
E.coli K99 |
4-26 head |
104 |
20 |
19.23 |
|
|
|
|
27-49 head |
57 |
15 |
26.32 |
0.407 |
|
|
|
50-70 head |
27 |
4 |
14.81 |
|
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
4-26 head |
104 |
7 |
6.73 |
|
|
|
|
27-49 head |
57 |
1 |
1.75 |
0.353 |
|
|
|
50-70 head |
27 |
2 |
7.41 |
|
|
|
|
Total |
188 |
10 |
5.32 |
|
|
Birth location |
E.coli K99 |
In barn |
115 |
27 |
23.48 |
|
|
|
|
Outside |
73 |
12 |
16.44 |
0.246 |
|
|
|
Total |
188 |
39 |
20.74 |
|
|
|
Salmonella enterica |
In barn |
115 |
6 |
5.22 |
|
|
|
|
Outside |
73 |
4 |
5.48 |
0.938 |
|
|
|
Total |
188 |
10 |
5.32 |
|
Table 11: Distribution of E.coli K99 and Salmonella Enterica with Dam and Environmental Associated Risk Factors
Multivariable Logistic Regression Analysis for the Association of Different Risk Factors with Presence of E. coli K99 and Salmonella Enterica
The results described in Table 13 and 14 showed multivariable logistic regression analysis that was performed to know the association of different risk factors with the occurrence of E. coli K99 and Salmonella enterica. In multivariable logistic regression analysis, sex was not significantly (P=0.289) associated with occurrence of E. coli K99 in which it was not significantly associated with occurrence of E.coli K99 in chi square test. The occurrence of E.coli K99 was 1.58(95% CI=0.683-3.598) times more likely to occur in male caves than female calves. Here was statistically significant difference observed between the occurrence E.coli K99 and different risk factors like colostrum fed within 24 hours (P=0.033) and continuous access of calf to dam during the first 24 hours (P=0.002). With regard to colostrum feeding within 24 hours, the prevalence of E.coli K99 was 0.15(95% CI=0.028- 0.861) times more likely to occur in calves fed colostrum within 24 hours than calves did not fed colostrum within 24 hours. There was no statistically significant difference observed (P >0.05) between the occurrence of the E. coli K99 and age, time of birth, type of delivery, birth location, herd size, age of dam at parturition and dam gestation number similarly with chi-square test.
Similarly colostrum fed within 24 hours (P=0.044) and age (P= 0.014) were significantly (P<0.05) associated with occurrence of Salmonella enterica. With regard to colostrum fed within 24 hours, the presence of Salmonella enterica was 0.15(95% CI=0.028- 0.861) times more likely to occur in calves fed colostrum within 24 hours than calves not fed colostrum within 24 hours. Also, the occurrence of Salmonella enterica was 13.15(95% CI=1.677- 103.127) times more likely to occur in 61-120 days age group than 1-60 days age group of the calves. Other hand, there was no statistically significant difference observed (P>0.05) between the occurrence of the Salmonella enterica with continuous access of calf to dam during the first 24 hours, age, time of birth, type of delivery, birth location, herd size, age of dam at parturition and dam gestation number.
|
Risk factors |
Category |
OR |
95% CI |
P-value |
|
|
1-60 days |
Ref. |
|
|
|
Age |
61-120 days |
0.43 |
0.100-1.872 |
0.260 |
|
|
121-180 days |
0.48 |
0.105-2.155 |
0.335 |
|
Sex |
Male |
1.58 |
0.683-3.598 |
0.289 |
|
|
Female |
Ref. |
|
|
|
Time of Birth |
Day |
0.163 |
0.010-2.711 |
0.206 |
|
|
Night |
0.20 |
0.0123-3.120 |
0.251 |
|
Type of delivery |
Normal |
3.96 |
.438-35.808 |
0.221 |
|
|
Assisted |
0.87 |
0.016-48.152 |
0.946 |
|
Birth location |
In barn |
Ref. |
|
|
|
|
Outside |
0.67 |
0.261-1.718 |
0.404 |
|
Colostrum fed within 24 |
|
|
|
|
|
hours |
Yes |
0.15 |
0.028-0.861 |
0.033* |
|
|
No |
Ref. |
|
|
|
Continuous access of calf to dam during the first 24 hours |
Yes |
0.10 |
0.022-0.426 |
0.002* |
|
|
No |
Ref. |
|
|
|
Herd size |
4-26 head |
Ref. |
|
|
|
|
27-49 head |
0.73 |
0.263-2.050 |
0.555 |
|
|
50-70 head |
0.32 |
0.067-1.480 |
0.144 |
|
Age of dam at parturition |
3-7 years |
Ref. |
|
|
|
|
8-12 years |
0.33 |
0.040- 2.674 |
0.297 |
|
|
13-16 years |
0.11 |
0.002-4.728 |
0.246 |
|
Dam gestation number |
< 3 |
Ref. |
|
|
|
|
3 |
1.52 |
0.438-5.254 |
0.511 |
|
|
>3 |
3.85 |
0.432-34.392 |
0.22 |
Legend: *statistically significant; OR-odds ratio; CI- confidence interval; Ref-reference category
Table 12: Multivariable Logistic Regression Analysis Showing the Association of Different Risk Factors with the Presence of E. coli K99
|
Risk factors |
Category |
OR |
95% CI |
P-value |
|
|
1-60 days |
Ref. |
|
|
|
Age |
61-120 days |
13.15 |
1.677-103.127 |
0.014* |
|
|
121-180 days |
4.36 |
0.494-38.476 |
0.185 |
|
Sex |
Male |
3.49 |
0.5293-23.057 |
0.194 |
|
|
Female |
Ref. |
|
|
|
Time of Birth |
Day |
0.33 |
0.009-11.873 |
0.546 |
|
|
Night |
0.19 |
0.005-7.606 |
0.374 |
|
Type of delivery |
Normal |
0.27 |
0.013-5.712 |
0.401 |
|
|
Assisted |
Ref. |
|
|
|
Birth location |
In barn |
Ref. |
|
|
|
|
outside |
0.86 |
0.160-4.650 |
0.864 |
|
Colostrum fed within |
|
|
|
|
|
24 hours |
Yes |
0.02 |
0.001-0.913 |
0.044* |
|
|
No |
Ref. |
|
|
|
Continuous access of |
|
|
|
|
|
calf to dam during the first 24 hours. |
Yes |
28.94 |
0.423- 1979.635 |
0.119 |
|
|
No |
Ref. |
|
|
|
Herd size |
4-26 head |
Ref. |
|
|
|
|
27-49 head |
0.48 |
0.0433-5.398 |
0.555 |
|
|
50-70 head |
5.70 |
0.512-63.377 |
0.157 |
|
Age of dam at parturition |
3-7 years |
Ref. |
|
|
|
|
8-12 years |
.15 |
.0001-248.459 |
0.619 |
|
|
13-16 years |
1 |
|
|
|
Dam gestation number |
< 3 |
Ref. |
|
|
|
|
3 |
1 |
|
|
|
|
>3 |
1.80 |
.001- 2908.273 |
0.876 |
Legend: *statistically significant; OR-odds ratio; CI- confidence interval; Ref- reference category
Table 14: Multivariable Logistic Regression Analysis Showing the Association of Different Risk Factors with the Presence of Salmonella Enterica
Discussion
E.coli K99 and Salmonella enterica are the main pathogen and most commonly associated with diarrhoea in calves[23,24]. The involvement of E.coli K99 and Salmonella enterica in calf diarrhea have not been well studied in Ethiopia. This current cross sectional study was aimed to isolate and identify E.coli K99 and Salmonella enterica from diarrheic calves at the age of one day to six months old. In the present study, the prevalence of E. coli K99 in diarrheic calf was 20.74%. This finding had nearly coincide with the find- ings of in India (18.75%), (28.4%) and (8) in Algeria(16 %) in di- arrheic calves [25,26]. A higher prevalence of E. coli K99 was re- ported by in Mozambique (40%) and in Brazil (35 %) [27,28]. On the contrary, many previous studies mentioned a lower prevalence 5.3%, 11.3%, 11.42%, 12.9 %. of E.coli K99 were reported by and respectively [29-32]. The differences in prevalence and frequency rates among the studies may be attributed to regional variations, management and hygienic conditions where E.coli K99 infection occurs mainly through ingestion of contaminated food and water, the age of the animals, vaccination of dam, colostrum intake, or different diagnostic methods [32].
Frequency distribution of E.coli K99 isolated from fecal sample of diarrheic calves within three Kebeles was 29.6%, 17.9% and 13.6% in Diho, Diduba and Kebena respectively. There is no sig- nificant difference within the three Kebeles in the frequency dis- tribution of E.coli K99 (P>0.05) that was P=0.147. This may be due to the similarity of management; location and the production system within the study area. There was no previous report in this study area for comparation to current result. Salmonella enterica is an important animal and human pathogen that can cause enteritis and septicaemia in calves [33]. Furthermore, Salmonella enterica serotypes are one of the most important foodborne pathogens, re- sulting in enteric disease, hospitalization, and deaths worldwide [3].
The occurrence of Salmonella enterica infection in diarrheic calves was 5.32% in present study. This finding is agreement with the re- sult reported by in India (5.0%) and in Uruguay (6.1%) [33,34]. Thus obtained finding is higher than that previously reported re- sults by (27) in Mozambique (2%). The detection of Salmonella enterica in calves is lower than those findings reported by (13%) and (16.25%) in Egypt [35,36]. The differences of the occurrence rates of Salmonella in calves than the previous reports may be at- tributed to the small age of calves which is highly susceptible to infection with Salmonella the difference in breed, geographical locations, poor management practices and standards of hygienic measures as well as used laboratory diagnostic tests. There is no previous report in Ethiopia showing distribution of Salmonella en- terica in calves up six months of age. In the current study, serotyp- ing of Salmonella enterica isolates revealed that 8 S. ZANZIBAR (80%) and 2 unclassified (20%).
The intent of this study was also aimed to assess risk factors asso- ciated with E.coli K99 and Salmonella enterica infections in new- born diarrheic calves. Multivariable logistic regression analysis was done to identify the significant risk factors associated with examined bacteria. Colostrum feeding within 24 hours and con- tinuous access of calf to dam during the first 24 hours were found to affect significantly the occurrence of E.coli K99 in diarrheic calves. Thus study showed that the occurrence of E.coli K99 in male calves was higher (25.27%) than in female calves (16.49%). This might be the reason that more care is given to female calves than male calf. Because female calf is needed by the pastoralist where compared with male calf. Further investigation should be conducted to reach conclusion to this assumption. Calves in age group 1 to 60 days were more susceptible while that age group 61-120 days and 121-180 days were less susceptible. The high prevalence of E.coli K99 shown by calves in age group 1-60 days (22.14%). The probable explanation for this finding may be related to either a poor and / or no absorption of colostrum and weakening of passive immunity. There was no statistically significant differ- ence between age group in the occurrence of this bacterium in this finding.
It was evident that calves in household, which received colos- trum within 24 hours from their dames were less frequent to be infected with E.coli K99 (P=0.033; OR=0.15; 95% CI=0.028- 0.861) than those did not fed colostrum within 24 hours calves. Similarly, calves had continuous access of calf to dam during the first 24 hours were less frequent to be infected with the pathogen (P=0.002, OR=0.10, 95% CI=0.022-0.426) those did not have con- tinuous access of calf to dam during the first 24 hours. There was high occurrence of E.coli K99 in calves that did not fed colostrum within 24 hours (73.33%) as compared to calves fed colostrum within 24 hours (16.18%). The enterotoxigenic E. coli, bearing the K99 attachment factor, can be found in calves younger than three days, especially in animals that did not receive any colostrum or received colostrum without any specific antibodies against this pathogen. The antibodies produced by cows in response to natural infection or vaccination are transmitted to the calf at birth via the colostrum [37].
Final multivariable logistic regression analysis showed that age and colostrum fed within 24 hours were significantly associated with Salmonella enterica shedding. Calf age was significantly associated with Salmonella enterica (P=0.014, OR=13.15, 95% CI=1.677-103.127). This result agreed with that reported by You- nis et al. (2009) who recorded age was significantly associated with Salmonella enterica in diarrheic calves in Egypt.
The highest occurrence of this pathogen was recorded in the 61-120 days of age (13.79%) than 1-60 days of age (2.86%) and 121-180 days of age (10.53%) diarrheic calves. Similarly with previous reports by that Salmonella enteric infection was usually recorded in the first 3 months of age causing severe symptoms [38]. Colostrum fed within 24 hours was significantly associated with the infection of Salmonella enterica in diarrheic calves (P=0.044, OR=0.02, 95% CI=0.001-0.913). The higher prevalence Salmonella enterica in the present study might be attributed to the variations in environmental and man- agement conditions of the households such as insufficient and/ or poor-quality colostrum intake by the calves. In addition to that poor hygiene often allows buildup of pathogenic strains in the young animalâ??s environment.
Antimicrobial agents are indispensable for decreasing mortal- ity and morbidity associated with infectious diseases in animals and humans [39]. In veterinary medicine, they have been used for therapy, metaphylaxis, prophylaxis, and growth promotion, being the enteric diseases one of the main animal infections treated with antibiotics [40]. In the present study, Salmonella enterica isolat- ed from diarrheic feces of calves were examined for antimicrobi- al susceptibility using 12 antibiotic discs. All 10(100%) isolates showed susceptible to ampicillin, amoxicillin+clavulanic acid, amikacin, chloramphenicol, cephalothin, ceftriaxone, ciprofloxa- cin, gentamycin, tetracycline and trimethoprim+ sulfamethoxaz- ole. Six isolates (60%) were found to be resistant for sulfisoxazole. The susceptibility of these isolates to antimicrobials may be linked with infrequent use for in the study area. Likewise, similar result, a study conducted in Brazil, also showed that the most common sus- ceptibility profile of Salmonella enterica isolated from calves was to amikacin, amoxicillin, ampicillin, ciprofloxacin, gentamicin, tetracycline, and trimethoprim+sulfamethoxazole [41]. The sus- ceptibility of Salmonella isolates to ciprofloxacin (100%) agreed with the reports of [42]. On the other hand, all the isolates were resistant to tetracycline (100%), streptomycin (91.4%),trimetho- prim+sulfamethoxazole(74.3%),chloramphenicol(85.7%),amoxi-cillin +clavulanic (62.9%) and gentamycin (60%) reported by in Uruguay reported that the most widespread resistance was against tetracycline (87.8%), streptomycin (85.4%) and ampicillin (22%) [33]. The wide variation in the observed results for Salmonella enterica isolated from calves could be due to differences on the tested antimicrobials, geographical regions, number of isolates and age of animals [41].
Conclusion and Recommendations
This study was planned to isolate and identify E.coli K99 and Sal- monella enterica from diarrheic calves with their associated risk factors. Multivariable logistic regression analysis and chi-square test were framed to find the risk factors behind the occurrence of E.coli K99 and Salmonella enterica. The prevalence of E.coli K99 and Salmonella enterica was 20.74%, 5.32% that obtained from 188 diarrheic calves, respectively. Three cases were mixed infec- tion by Salmonella enterica and E.coli K99. In the present study, pathasure enteritis 4 (antigen test kit) and Biolog GEN III micro- plate enabled identification of E.coli K99 and Salmonella enterica in diarrheic calves from 1 day to 6 months of age respectively. Biolog GEN III microplate was found delivering a high level of accuracy that is comparable to molecular methods among avail- able diagnostic tests. Pathasure enteritis 4 tests (ELISA) was found to be quick identification method of E.coli K99 which is compared with convectional laboratory methods like bacterial culture. Cul- ture characteristics of E. coli K99, bright pink to red, yellow and distinct green metallic sheen colonies were observed on MacCon- key, XLD and EMB agar media respectively. On the other hand, Salmonella species revealed red, pale and colorless, a pink-red zone colony with black center on MacConkey, Salmonella Shigel- la (SS) and XLD agar respectively. Salmonella showed positive reaction to selected biochemical tests such as ureases test, citrate utilization test, lysine iron agar and triple sugar iron agar test. This study revealed that 8 were S. ZANZIBAR (80%) from 10 sero- typed Salmonella Enterica.
The results of antimicrobial susceptibility testing revealed that all Salmonella enterica isolated from diarrheic feces of calves were susceptible (100%) to ampicillin, amoxicillin+clavulanic acid, amikacin, chloramphenicol, cephalothin, ceftriaxone, ciproflox-acin, gentamycin, tetracycline and trimethoprim+sulfamethoxaz- ole. Colostrum feeding within 24 hours and continuous access of calf to dam during the first 24 hours were found significantly as- sociated with occurrence of E.coli K99 and Salmonella enterica. On the other hand there was no statistically significant difference observed between the occurrence of the E. coli K99 and Salmo- nella enterica with time of birth, type of delivery, birth location, herd size, age of dam at parturition and dam gestation number. The virulence factors of the organisms were not identified in present study. In general identification of the risk factors associated with the spreading of bacteria causing diarrhea may be helpful for con- struction of suitable methods for prevention and control. Based on the above conclusions, the current study led to the following recommendations:
• Molecular characterization of E.coli K99 and Salmonella enteri- ca should be conducted.
• Awareness of the pastoral community should be done about the impotence of colostrum feeding to their calves and other manage- ment practices.
• Further study also should be made in this area to get more infor- mation about E.coli
• K99, Salmonella enterica and associated risk factors.
• Antimicrobial susceptibility testing should be conducted regular- ly on commonly used drugs.
Limitations of the Study
• Failure to conduct antimicrobial susceptibility of E.coli K99 iso- lates in order to know drug resistance of the pathogen in calves due to lack of testing antibiotic disc. Because the objective of the project is addressing causes of young stock mortality likes infec- tious disease, such as E. coli K99, Salmonella and management problem.
• Failure to molecular characterization of the isolated bacteria.
• Lack of some chemicals and reagent which are important for isolation and identification of Salmonella using biochemical tests supplied by the project.
• Failure to present the result at the given schedule due to the proj- ect time management, shortage of transport vehicle and availabili- ty of laboratory resources.
Annexes
Some Pictures that have been taken during Field and Laboratory Works




References
- Ferreira, J. P., Gochez, D., Jeannin, M., Magongo, M. W., Loi, C., Bucher, K., ... & Erlacher-Vindel, E. (2022). JAC- Antimicrobial Resistance.
- Hajipour, M. J., Fromm, K. M.,Ashkarran,A.A., deAberasturi,D. J., de Larramendi, I. R., Rojo, T., ... & Mahmoudi, M. (2012). Antibacterial properties of nanoparticles. Trends in biotechnology, 30(10), 499-511.
- Hur, J., Jawale, C., & Lee, J. H. (2012). Antimicrobial resistance of Salmonella isolated from food animals: A review. Food Research International, 45(2), 819-830.
- Beyene, G., Nair, S., Asrat, D., Mengistu, Y., Engers, H., & Wain, J. (2011). Multidrug resistant Salmonella Concord is a major cause of salmonellosis in children in Ethiopia. The Journal of Infection in Developing Countries, 5(01), 023-033.
- Keelara, S., Scott, H. M., Morrow, W. M., Gebreyes, W. A., Correa, M., Nayak, R., ... & Thakur, S. (2013). Longitudinal study of distributions of similar antimicrobial-resistant Salmonella serovars in pigs and their environment in two distinct swine production systems. Applied and environmental microbiology, 79(17), 5167-5178.
- Ferede, Y., Mazengia, H., Bimrew, T., Bitew, A., Nega, M., Kebede, A., ... & Bitew, A. (2014). Pre-weaning morbidity and mortality of crossbred calves in Bahir Dar Zuria and Gozamen districts of Amhara region, northwest Ethiopia. Open Access Library Journal, 1(3), 1-8.
- Wudu, T., Kelay, B., Mekonnen, H. M., & Tesfu, K. (2008). Calf morbidity and mortality in smallholder dairy farms in Ada’a Liben district of Oromia, Ethiopia. Tropical Animal Health and Production, 40, 369-376.
- Selles, S. M. A., Kouidri, M., Belhamiti, B. T., Amrane, A. A., Benia, A. R., Hammoudi, S. M., ... & Boukraa, L. (2018). Enteropathogens associated with neonatal calves diarrhea in Tiaret area (Western Algeria). Veterinaria, 67(2).
- El-Seedy, F. R., Abed, A. H., Yanni, H. A., & Abd El-Rahman,S. A. A. (2016). Prevalence of Salmonella and E. coli in neonatal diarrheic calves. Beni-Suef University journal of basic and applied sciences, 5(1), 45-51.
- Alemu, S., & Zewde, B. M. (2012). Prevalence and antimicrobial resistance profiles of Salmonella enterica serovars isolated from slaughtered cattle in Bahir Dar, Ethiopia. Tropical animal health and production, 44, 595-600.
- Klein-Jöbstl, D., Iwersen, M., & Drillich, M. (2014). Farm characteristics and calf management practices on dairy farms with and without diarrhea: A case-control study to investigate risk factors for calf diarrhea. Journal of dairy science, 97(8), 5110-5119.
- Shaaban, S. I., Ayoub, M. A., Ghorbal, S. H., & Nossair, M. (2018). Calves as a Reservoir of Some Diarrheagenic Agents for Human Contacts in El-Behira Province. Alexandria Journal for Veterinary Sciences, 56(2).
- Humphries, R. M., & Linscott, A. J. (2015). Practical guidancefor clinical microbiology laboratories: diagnosis of bacterialgastroenteritis. Clinical microbiology reviews, 28(1), 3-31.
- Kubota, K. A., Wolfgang, W. J., Baker, D. J., Boxrud, D., Turner, L., Trees, E., ... & Gerner-Smidt, P. (2019). PulseNet and the changing paradigm of laboratory-based surveillance for foodborne diseases. Public Health Reports, 134(2_suppl), 22S-28S.
- Manual, O. I. E. (2016). Office International des Epizootiasis (OIE) Terrestrial (World Organisation for Animal Health) Cryptosporidiosis Review, 2008: 1-55.
- Mengistu, S., Abayneh, E., & Shiferaw, D. (2017). E. coli O157: H7 and Salmonella species: public health importance and microbial safety in beef at selected slaughter houses and retail shops in eastern Ethiopia. J Vet Sci Technol, 8(468), 2.
- Beyi, A. F., Fite, A. T., Tora, E., Tafese, A., Genu, T., Kaba, T., ... & Cox, E. (2017). Prevalence and antimicrobial susceptibility of Escherichia coli O157 in beef at butcher shops and restaurants in central Ethiopia. BMC microbiology, 17, 1-6.
- Assefa, A., & Bihon, A. (2018). A systematic review and meta-analysis of prevalence of Escherichia coli in foods of animal origin in Ethiopia. Heliyon, 4(8).
- Bibiso, M. (2017). Assessment of soil properties in some selected parts of Ethiopia. American-Eurasian Journal of Agriculture and Environmental Science, 17 (2), 143, 147.
- Quinn, P. J., Markey, B. K., Leonard, F. C., Hartigan, P., Fanning, S., & Fitzpatrick, E. (2011). Veterinary microbiology and microbial disease. John Wiley & Sons.
- Annex, I. (2013). MEETING OF THE OIE TERRESTRIAL ANIMAL HEALTH STANDARDS COMMISSION. Work.
- Hasoon, N. A., & Hamed, S. L. (2019). Molecular characterization of carbapenemase-producing Gram-negative bacteria isolated from clinical specimens in Baghdad. Iraq. J Pure Appl Microbiol, 13(2), 1031-40.
- Prescott, J. F., MacInnes, J. I., Van Immerseel, F., Boyce,J. D., Rycroft, A. N., & Vázquez-Boland, J. A. (2022). Pathogenesis of bacterial infections in animals. John Wiley & Sons, Incorporated.
- Tegegne, F. M. (2019). Epidemiology of Salmonella and its serotypes in human, food animals, foods of animal origin, animal feed and environment. J Food Nutr Health, 2(1), 7-14.
- Sharieff, S. (2015). Microbiological Study of Pathological Vaginal Discharge in Rural Women (Doctoral dissertation, Rajiv Gandhi University of Health Sciences (India)).
- Shahrani, M., Dehkordi, F. S., & Momtaz, H. (2014). Characterization of Escherichia coli virulence genes, pathotypes and antibiotic resistance properties in diarrheic calves in Iran. Biological Research, 47, 1-13.
- Acha, S. J., Kühn, I., Jonsson, P., Mbazima, G., Katouli, M., & Möllby, R. (2004). Studies on calf diarrhoea in Mozambique: prevalence of bacterial pathogens. Acta Veterinaria Scandinavica, 45(1), 1-10.
- Langoni, H., Linhares, A. C., Avila, F. A. D., Da Silva, A. V., & Elias, A. O. (2004). Contribution to the study of diarrhea etiology in neonate dairy calves in São Paulo state, Brazil. Brazilian Journal of Veterinary Research and Animal Science, 41, 313-319.
- Kumar, S., Suvidhi, D. K., Milind, M., & Bhati, T. (2022). Occurrence of E. coli and associated risk factors in neonatal calf diarrhea from Bikaner. The Pharma Innovation Journal, 11, 1030-1033.
- Pourtaghi, H., & Sodagari, H. R. (2016). Antimicrobial resistance of entrotoxigenic and non-entrotoxigenic Escherichia coli isolated from diarrheic calves in Iran. Int J Enteric Pathog, 4(2), e34557.
- Lacroute, H. (2016). Place de la colibacillose chez le jeune veau et antibiorésistance dans l’Allier (2011-2013). Bulletin de l'Académie Vétérinaire de France, 169(2), 100-109.
- Picco, N. Y., Alustiza, F. E., Bellingeri, R. V., Grosso, M. C., Motta, C. E., Larriestra, A. J., ... & Vivas, A. B. (2015). Molecular screening of pathogenic Escherichia coli strains isolated from dairy neonatal calves in Cordoba province, Argentina. Revista Argentina de microbiologia, 47(2), 95-102.
- Casaux, M. L., Caffarena, R. D., Schild, C. O., Giannitti, F., Riet-Correa, F., & Fraga, M. (2019). Antibiotic resistance in Salmonella enterica isolated from dairy calves in Uruguay. Brazilian Journal of Microbiology, 50, 1139-1144.
- Nethra, R., Ramesh, P. T., Veeregowda, B. M., Kamran, C. A., Muralidhara, A., & Sathyanarayana, M. L. (2019). Prevalence of Salmonella in Bovine Fecal Samples, Slaughter House Samples and Environmental Samples. Int. J. Curr. Microbiol. App. Sci, 8(10), 352-360.
- El-Rahman, A. M. A., Mahmoud, A. E. K. A., Khadr, A. M., & El-Shemy, T. M. (2016). Some Studies on Salmonella Enterica Associated with Diarrhea in Cattle. Alexandria Journal for Veterinary Sciences, 48(2).
- Gharieb, R. M., Fawzi, E. M., Attia, N. E., & Bayoumi, Y. H.(2015). Calf diarrhea in Sharkia province, Egypt: diagnosis; prevalence, virulence profiles and zoonotic potential of the causal bacterial agents. International Journal of Agriculture Science and Veterinary Medicine, 3, 71-87.
- Gharieb, R. M., Fawzi, E. M., Attia, N. E., & Bayoumi, Y. H. (2015). Calf diarrhea in Sharkia province, Egypt: diagnosis; prevalence, virulence profiles and zoonotic potential of the causal bacterial agents. International Journal of Agriculture Science and Veterinary Medicine, 3, 71-87.
- Fossler, C. P., Wells, S. J., Kaneene, J. B., Ruegg, P. L.,Warnick, L. D., Bender, J. B., ... & Halbert, L. W. (2005). Herd-level factors associated with isolation of Salmonella in a multi-state study of conventional and organic dairy farms: I. Salmonella shedding in cows. Preventive veterinary medicine, 70(3-4), 257-277.
- Tadesse, D. A., Zhao, S., Tong, E., Ayers, S., Singh, A., Bartholomew, M. J., & McDermott, P. F. (2012). Antimicrobial drug resistance in Escherichia coli from humans and food animals, United States, 1950–2002. Emerging infectious diseases, 18(5), 741.
- Teuber, M. (2001). Veterinary use and antibiotic resistance.Current opinion in microbiology, 4(5), 493-499.
- Souto, M. S., Coura, F. M., Dorneles, E. M., Stynen, A. P.R., Alves, T. M., Santana, J. A., ... & Lage, A. P. (2017).Antimicrobial susceptibility and phylotyping profile of pathogenic Escherichia coli and Salmonella enterica isolates from calves and pigs in Minas Gerais, Brazil. Tropical animal health and production, 49, 13-23.
- Guesh, M. (2017). Prevalence and antimicrobial susceptibility of Salmonella species from lactating cows in dairy farm of bahir dar town, Ethiopia. African Journal of Microbiology Research, 11(43), 1578-1585.