Research Article - (2025) Volume 3, Issue 2
Antimicrobial Resistance Patterns of Isolates from Bloodstream Infections at Jinja Regional Referral Hospital in Uganda
2Jinja Regional Referral Hospital, Jinja, Uganda
3Centers for Antimicrobial Optimization Network (CAMONet), Infectious Diseases Institute, College of Health Sciences, Makerere University, Kampala, Uganda
Received Date: Apr 25, 2025 / Accepted Date: Jun 16, 2025 / Published Date: Jul 25, 2025
Copyright: ©2025 Fahad Lwigale, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Lwigale, F., Akello, G., Mayito, J., Magembe, J., Wanjala, B. W., Bakabweyaka, A., et al. (2025). Antimicrobial Resistance Patterns of Isolates from Bloodstream Infections at Jinja Regional Referral Hospital in Uganda. J Emerg Med OA, 3(2), 01-12.
Abstract
Introduction Bloodstream infections (BSIs) are common acute syndromes of variable origin with high chances for extreme health effects including death. Empirical therapy is usually employed to manage the BSIs despite the high resistance to the antibacterial agents used including cefriaxone and/or metronindazole. This study determined the aetiologic agents responsible for bloodstream infections, their antimicrobial susceptibility patterns, and infection sources in Eastern-central Uganda.
Methods This retrospective cross-sectional study involved analysis of data from all non-duplicate blood culture reports from 2019 to 2021. These were earlier generated using single set blood culture, conventional microbiology techniques and disc diffusion-based antimicrobial susceptibility testing (AST) guided by the Clinical and Laboratory Standards Institute (CLSI) guidelines. World Health Organization Network (WHONET) 2022 desktop software and Microsoft Excel were utilized for data analysis.
Results The 1364 reports showed a diagnostic yield of 6.8% (93/1364) and a 1.5% (21/1364) contamination rate. Overall, 37% and 13% of infections were hospital-acquired and community-acquired respectively. Most etiologic agents were Gram-positive bacteria dominated by Staphylococcus aureus 39/114 (34.2%). Polymicrobial growth existed in 4/1364(0.3%) cases. Staphylococcus aureus was mainly resistant to penicillin G (100%) and most susceptible to gentamicin (%S=74.1, 95% C.I:53.4-88.1). Escherichia coli and unspecified Coliforms together showed high resistance to ampicillin (87.5%) and third-generation cephalosporins (100%). The least resistance was to chloramphenicol and carbapenems.
Conclusion Staphylococcus aureus, which is highly sensitive to gentamicin, was the commonest cause of BSIs in this setting. Therefore, empirical management of suspected BSI cases in the region should include gentamicin. To strengthen antimicrobial stewardship, microbiology services, including culture and sensitivity, are required to guide targeted antimicrobial use and monitor resistance trends.
Keywords
Antimicrobial Resistance, Blood Culture, Bloodstream Infections, Healthcare-Associated Infections, Jinja, Microbiology
Introduction
Bloodstream Infections (BSIs) are some of the most commonly encountered and serious healthcare-associated infections (HAIs) of public health concern [1-3]. They result from microorganisms accessing the bloodstream where they proliferate and excrete toxins during septicemia [4]. They are characterized by a number of symptoms including fever (>38°C), chills, hypotension, tachypnea, hypothermia (<36°C), apnea, or bradycardia depending on different factors including the age group [2,5]. A bloodstream infection (BSI) whether primary or secondary, is confirmed by a positive blood culture in the microbiology laboratory [1,2,6]. A primary BSI is a laboratory-confirmed positive blood culture without any link to another infected body site [5]. A secondary BSI on the other hand involves the isolation of an organism within a 14-17-day period from the bloodstream with the same identified as one responsible for another existing body site-specific infection or contributes to the syndrome [5]. This can be a urinary tract infection (UTI), respiratory tract infection (RTI), or skin and soft- tissue infection (SSTI) among others.
Among all ages, BSIs are one of the major causes of morbidity and mortality in both developed and developing countries [7-9]. As an emergency condition, sepsis requires immediate detection and identification of the causative agent for proper treatment given the emergence of antimicrobial resistance (AMR) [1,10,11]. For this matter, blood culture testing is an important tool for the diagnosis of BSIs and AMR surveillance [4,6,12]. Majority of reported BSI cases are in the 18-88 year age group, of which 10% to 81.5% cases are of hospital origin [2,4,8]. Geographic and epidemiological factors also result in differences in the BSIs profiles [2,8-10,13]. The “ESKAPE” group composed of Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species, altogether contributes up to 70% of the observed BSIs [15]. Variable resistance levels have been reported for multiple antibiotics including cephalosporins, gentamicin, and carbapenems among Enterobacterales and Acinetobacter baumannii [9,13,16].
These infections are commonly managed empirically using a combination of broad-spectrum intravenously administered antibiotics mainly ceftriaxone and metronidazole despite the reported high resistance against the third generation cephalosporins. The Uganda clinical guidelines also recommend use of gentamicin as part of the therapeutic combination for septicaemia of unknown aetiology [17]. However, only 30% [18] compliance to the guidelines has been reported with increased inappropriate use of ceftriaxone for majority of cases [19,20]. This creates more room for the emergence of AMR among bacteria and consequences including therapeutic failure, longer hospital-stay periods, higher chances for nosocomial infections, increased healthcare costs, and higher risk of death [13,21,22].
Despite the setup of the facility medicines and therapeutic committee (MTC), there is a shortage of congregated up-to-date information to guide the choice of antibiotics for empirical therapy to ensure better control of antimicrobial use in the region. The ever- changing microbiological profile of BSIs and the threat of AMR makes it essential to routinely generate, analyze, and utilize the local microbiology data to develop and/or update facility-based treatment guidelines to manage infectious syndromes. This study aimed to determine the burden of bloodstream infections, common etiological agents; the percentage resistance and/or sensitivity of the causative agents to common antibiotics; and the proportions due to community and healthcare-associated infections at Jinja Regional Referral Hospital (Jinja RRH).
Materials & Methods
Operational Definitions
1. A Bloodstream Infection (BSI) refers to a positive blood culture test due to the isolation of a significant pathogenic microorganism/s from the suspected patient’s blood sample. The organism can be bacterial or fungal in nature.
2. Organisms commonly found on the skin such as Corynebacterium species, and Bacillus species other than B. anthracis were treated as contaminants. Coagulase-negative staphylococci (CoNS) were also regarded as contaminants since only one blood culture set was collected for analysis during primary data generation. The rest of the organisms were considered significant aetiologic agents (Significant growth) when calculating the frequencies and respective percentages.
3. Methicilin-resistant Staphylococcus aureus (MRSA) were isolates of Staphylococcus aureus resistant to cefoxitin [23].
4. Hospital and community-acquired infections were defined as a proxy based on the duration of patient hospitalization at the facility before being symptomatic for infection and sample collected for laboratory analysis [24]. The infections diagnosed based on samples collected from patients admitted and exposed to the hospital premises for more than 48 hours (two days) were treated as hospital-acquired infections [24]. These are also referred to as healthcare-associated infections (HAI) or nosocomial infections [25].
5. Community-acquired infections on the other hand involved diagnosis of infection among patients exposed to the premises of the hospital for not more than 48 hours [24].
Study Design
This was a retrospective cross-sectional study conducted in 2022 using data generated over a two-and-a-half-year period from January 2019 through June 2021. This period was shortly after the stabilization of microbiology service delivery in the region with support from the Infectious Diseases Institute (IDI), Makerere University College of Health Sciences.
Study Setting and Population
The data was collected at Jinja RRH, which is a 500-bed capacity hospital located along Bax street within the central town of Jinja city in the Eastern-central part of Uganda. This facility serves a population of approximately 4.5 million people from within Jinja and the surrounding areas of over 11 districts including Iganga, Mayuge, Kamuli, Buikwe, Lugazi, Kayunga, and Mukono districts. The facility is equipped with a microbiology laboratory where examinations of various samples are conducted for routine patient care and management. Blood culture was carried out using the BD-BACTEC™ FX40 automated system. Organism identification was carried out using conventional methods including Gram staining, microscopy, sugar utilization and biochemical testing up to the genus and species levels. Antimicrobial susceptibility testing(AST) was carried out using the Kirby-Bauer disk diffusion method following the Clinical and Laboratory Standards Institute (CLSI) guidelines; CLSI-2018-M100-S28 [26], CLSI-2020- M100-S30 [27] and CLSI-2021-M100-S31 [23]. The laboratory is accredited, performs internal quality control and routinely engages in external quality assessments plus inter-laboratory comparison with the national reference laboratory.
Data from non-duplicate single set blood culture tests carried out for all patients with suspected BSI during the study period was considered. Same identity isolates from the same patient relative to different hospital visits were excluded.
Data Collection
The data necessary for the study was accessed on February 1, 2022, and extracted from the African Laboratory Information System (ALIS) into an Excel sheet. This was cleaned and anonymized by removing specific identifiers such as patient names and numbers before being analyzed.
Inclusion and Exclusion Criteria
All sex types and age groups such as pediatrics, adolescents, and adults were included in the analysis. Only the first isolate of every patient in the study period was included in the analysis to generate the antimicrobial susceptibility patterns as guided by CLSI-M39A4E-2014 [28]. A minimum of thirty (30) isolates were required for an isolate to be analyzed individually to generate the antimicrobial resistance profile. Otherwise, they were grouped. Patient data without final laboratory analysis results and those outside the study interval were not considered for analysis.
Ethics Statement
The Jinja Hospital Research and Ethics Committee (JREC) approved the study with registration number JREC 159/2022. The data was obtained from records of personnel who seek medical services at the facility. This is kept in medical records including patient files, laboratory request forms, and information systems for proper care and later reference. The Microbiology Laboratory records were anonymously analyzed for this study and the Jinja Hospital Research and Ethics Committee waived the need to obtain consent from individual patients. All methods were executed in accordance with relevant guidelines and regulations.
Data Analysis
Microsoft Excel (2016) was used for data cleaning and descriptive analysis. World Health Organization Network (WHO-NET 2022) desktop software was used to analyze the antimicrobial susceptibility data. Only the first isolate of every participant was involved in the generation of antimicrobial susceptibility profiles. These were presented as percentage resistance and/or sensitivity/ susceptibility. Results including isolate distributions, possible infection sources, organisms/aetiologic agents, and AST data were presented in the form of frequencies and percentages in different tables and figures. There was a description of their distribution in socio-demographics including age, sex, and admission location (wards).
Results
The Existing Level of Bloodstream Infections
A total of 1364 participants were involved in the analysis. 800(58.7%) were male. The majority (86.2%) of participants were less than 12 years of age. A total of 114(8.4%) blood samples were positive. Of these, 93 had significant growth representing a true diagnostic yield of 6.8% as shown in Figure 1. The contamination rate observed was 1.5%(21/1364). Positivity was observed more among the males 67/114(58.8%) than the females 47/114(41.2%) (Table 1).
Figure 1: Levels of Positivity for Blood Cultures. The Observed Prevalence of Bloodstream Infections is 6.8%.
Table 1: Outcomes from Blood Culture in the Different Population Groups
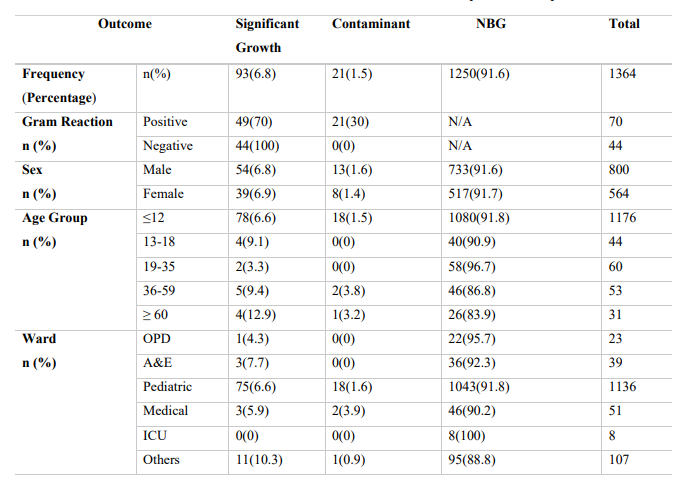
A&E, Accidents and Emergency; ICU, Intensive Care Unit; NBG, No Bacterial Growth; N/A, Not Applicable; OPD, Outpatient Department; n, Frequency; %, Percentage
Frequency and Percentage of Isolated Organisms
Of the 114 positive cases, 68(60.0%) were Gram-positive bacteria while 44(38.6%) were Gram-negative. Candida species represented the remainder 2/114(1.7%) (Figure 2).
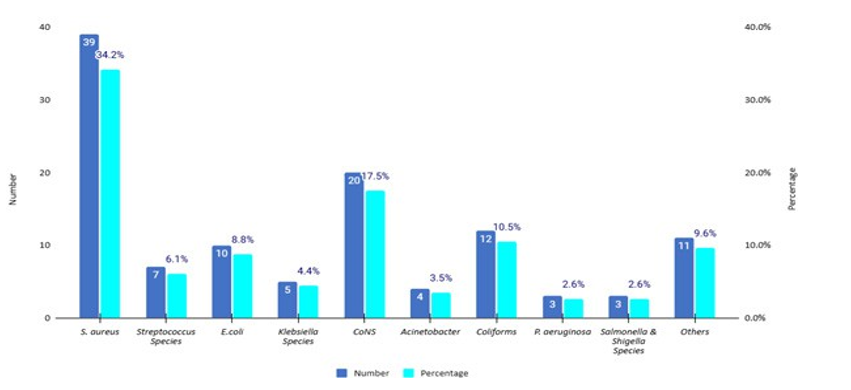
Figure 2: Frequency and Percentage of Isolated Organisms (n=114).
Gram positive bacteria majorly included, Staphylococcus aureus 39(34.2%), Coagulase negative staphylococcus 20(17.5%) then Streptococcus species, 7(6.1%). Gram-negative bacteria on the other hand were made of general unspecified Coliforms 12(10.5%), E. coli 10(8.8%), Klebsiella species 5(4.4%), Acinetobacter species 4(3.5%), Pseudomonas aeruginosa 3(2.6%), then Salmonella species 2(1.8%). Other organisms included Candida albicans, Enterobacter, Serratia, Citrobacter, and Shigella species. Most cases were observed in the pediatric unit among children less than 13 years of age. Polymicrobial growth was observed in 4(0.3%) cases of which two involved a Serratia species and one had a Bacillus species in addition to the co-isolates.
Antimicrobial Susceptibility Profile of Isolated Organisms
Staphylococcus aureus was majorly susceptible to gentamicin 21/27(74.1%) (Table 2). Among Escherichia coli and unspecified Coliforms isolates combined(n=22), percentage resistance to ampicillin, cefuroxime, ceftazidime, cefotaxime, gentamicin, ciprofloxacin, imipenem, ertapenem, co-trimoxazole, tetracycline, and chloramphenicol was 87.5%, 100%, 100%, 100%, 50%, 50%, 20%, 33.3%, 66.7%,60%, and 28.6% respectively. Non- Enterobacterales (Pseudomonas aeruginosa and Acinetobacter baumannii) together (n=7) had the following resistance profile; ceftazidime (20%), aztreonam (50%), imipenem (0%), meropenem (50%), gentamicin (40%), ciprofloxacin (33.3%) and tetracycline (0%).
Table 2: Antimicrobial Susceptibility Test Response for S. aureus Isolates
|
Code |
Antibiotic name |
Antibiotic class |
Breakpoints |
Number |
%R |
%I |
%S |
%R 95%C.I. |
%S 95%C.I. |
|
FOX_ND30 |
Cefoxitin |
Cephems |
S >= 22 |
5 |
60.0 |
0.0 |
40.0 |
|
7.3-83.0 |
|
CHL_ND30 |
Chloramphenicol |
Phenicols |
13 - 17 |
6 |
33.3 |
0.0 |
66.7 |
6.0-75.9 |
24.1-94.0 |
|
CIP_ND5 |
Ciprofloxacin |
Quinolones |
16 - 20 |
29 |
20.7 |
24.1 |
55.2 |
8.7-40.3 |
36.0-73.0 |
|
CLI_ND2 |
Clindamycin |
Lincosamides |
15 - 20 |
22 |
13.6 |
9.1 |
77.3 |
3.6-36.0 |
54.2-91.3 |
|
ERY_ND15 |
Erythromycin |
Macrolides |
14 - 22 |
23 |
52.2 |
8.7 |
39.1 |
31.1-72.6 |
20.5-61.2 |
|
GEN_ND10 |
Gentamicin |
Aminoglycosides |
13 - 14 |
27 |
14.8 |
11.1 |
74.1 |
4.9-34.6 |
53.4-88.1 |
|
PEN_ND10 |
Penicillin G |
Penicillins |
S >= 29 |
6 |
100.0 |
0.0 |
0.0 |
51.7-100 |
0.0-48.3 |
|
TCY_ND30 |
Tetracycline |
Tetracyclines |
15 - 18 |
20 |
25.0 |
20.0 |
55.0 |
9.6-49.4 |
|
R-Resistant; S-Susceptible; I-Intermediate; C.I-Confidence Interval; %-Percentage
Supplementary Table S1 shows the total number of different isolates and respective proportions tested against the antibiotics. Further, respective frequencies for the Resistant(R), Intermediate(I), and Susceptible(S) categories as outcomes of the antimicrobial susceptibility test (AST) reporting format have been revealed. This details the ratio of organism to antibiotic tested with respective outcomes compared with the number of organisms isolated.
Sources of Confirmed Bloodstream Infections
Altogether, 36.8% and 13.2% of the positive cases were hospital- acquired and community-acquired infections respectively. On a sad note, over 50% of the cases were of unknown origin (Table 3).
Table 3: Possible Infection Sources in The Different Population Groups.
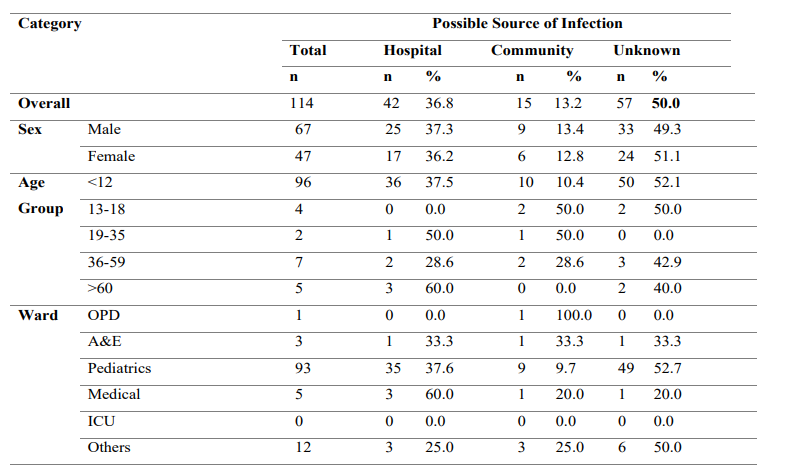
A&E, Accidents and Emergency; ICU, Intensive Care Unit; OPD, Outpatient Department; n, Number; %, Percentage
Discussion
This study obtained a significant positivity rate of BSIs of 6.8% which is greater than 4.6% and 2.6% previously observed in Uganda and Jinja RRH respectively [12]. This indicates an improvement in pathogen recovery from blood cultures. Similar rates have been obtained in other studies [16,29-31]. Variations in BSI prevalence in different units of health facilities have also been reported[3]. Another study in Ethiopia obtained a much much higher positivity of up to 71% [32]. This could be due to the study population of cancer patients, majority of whom were neutropenic and hence at a higher risk for BSIs [32,33]. The highest positivity in this study was observed among the pediatrics. This could be explained by the low immune status of the children, making them more vulnerable to the BSIs. This calls for improvement in the infection prevention measures in these units. Blood culture is also recommended among adults whose presentation fits the definition of sepsis to enable timely case management and limit possible mortality. Up to 91.6% of the blood cultures had no bacterial growth. This could be due to the prior use of antibiotics between 1-28 days before blood culture testing which was observed in many participants, especially in the period starting early 2020. More males (58.8%) than females had BSIs, which is in agreement with the majority of the previous studies [2,4,13,14].
Gram-positive bacteria (61.4%) were more dominant compared to their Gram-negative counterparts (38.6%) as previously reported in other studies [3,4,12,14,32]. However, there have also been differing observations elsewhere where Gram-negatives were the majority [9,10,16,30,34,35]. This is only a tip of the iceberg regarding the variation of dominant organism distribution, which calls for local data analysis and utilization.
Overall, Staphylococcus aureus (34.2%) was the most isolated organism. This finding was also observed elsewhere [4,12,32]. Coagulase-Negative Staphylococcus (CoNS) were also more frequently isolated in the pediatric samples compared to Streptococcus species, contributing to a contamination rate of 1.5% (21/1364). This was less than 13.6% and 4.8% observed at Princess Marina Hospital (PMH) and Tygerberg Hospital (TBH) neonatal units in Botswana and South Africa respectively [36]. Other studies have also reported higher rates [2,9,14,15,22,31]. This could be due to the high likelihood of contamination in children’s blood samples during sample collection. However, the observed contamination rate in this study falls within acceptable limits ranging from 1-3% [6, 37]. These isolates were not treated as true pathogens and no AST was done on them. This is a good practice to avoid unnecessary use of antibiotics in these children and their possible effects. Notably, only a single blood culture set was commonly carried out. This is most likely due to the limited availability of testing materials such as blood culture bottles among other factors. This on the other hand limits the capability to confirm the fact that common commensals such as CoNS could be responsible for some of the true infections observed especially among immunocompromised individuals and children. This in turn leaves them vulnerable to the BSIs of that origin. It is recommended to employ appropriate policies to monitor and reduce the contamination levels to less than 1% and avoid the associated consequences [37].
AST was variably done and the ratio of the isolates to antibiotic tested was quite low (Refer to supplementary Table S1), similar to what was observed at Mulago Hospital [38]. This shows the need for Uganda to improve supply management to ensure stable availability of logistics for laboratory testing to enhance AMR surveillance. For the number of Staphylococcus aureus isolates tested against cefoxitin(n=5), 60% turned out resistant indicating a possibility of high prevalence of MRSA in circulation. These being β-Lactamase producers, they had already shown no response to penicillin G (%R=100%). However, gentamicin 74.1% (%S), ciprofloxacin 55.2% (%S), and tetracycline 55% (%S) showed better activity against Staphylococcus aureus compared to other agents.
Escherichia coli and unspecified Coliforms together (n=22) showed relatively high levels of resistance to commonly used antibiotics for empiric and targeted treatment such as the third- generation cephalosporins. They also showed 20% and 33.3% resistance to imipenem and ertapenem respectively among the isolates tested. This indicates increasing resistance to the reserved antibiotics, limiting the availability of alternatives when faced with MDR pathogens. A similar feature was observed among the non-Enterobacterales, although they were susceptible to tetracycline. However, there were fewer than 30 isolates for these two groups of organisms, making it impossible to make appropriate conclusions about their susceptibility patterns as guided by CLSI- M39A4E-2014 [28].
Although over 50% of the patients lacked date of hospitalization prior to sample collection, at least 37% had nosocomial BSI. Similar observations were made elsewhere[2]. This makes it necessary to strengthen infection prevention and control measures observed in the facility to impede outbreaks and spread of resistant bugs. Up to 13% of the positive cases could be linked to community-acquired infections. This is less than what has been reported by other studies [29, 36]. The shortage of data on some variables calls for the need to improve information management systems at the surveillance sites including stronger linkage of the laboratory and clinical units.
Limitations
This study had limitations. The first is that there was quite a lot of missing data from the utilized information system especially relative to the hospitalization history. This led to insufficient categorization of infections as HAI or CAI. Secondly, very few isolates had been tested against each antibiotic among the positive cultures which hindered deeper analysis per isolate species. This could be due to the frequent stockouts of antibiotic discs at the study facility. Otherwise, higher isolate-antibiotic test ratios would provide better information to guide empirical therapy locally. Last but not least, the analyzed data was generated from a single site despite patients originating from a wide geographical area. This facility being the only microbiology testing site in the region limited comparability options of culture and sensitivity testing outcomes.
Conclusions
Staphylococcus aureus was the most common organism responsible for bloodstream infections at this site. There is a relatively high level of resistance to commonly used antibiotics such as ceftriaxone among all organisms, indicating the need for their reduced empirical use. Currently, gentamicin can be considered a priority when selecting combination agents to empirically manage BSIs suspected to be due to Staphylococcus aureus. Use of microbiology services especially culture and sensitivity needs to be increased to guide targeted antimicrobial use. Further investigation is required to determine the proportion of BSIs due to CoNS to inform proper management, antimicrobial stewardship and surveillance policies.
Acknowledgements
Much appreciation for contributions from the administration and staff of Jinja Regional Referral Hospital, especially the Children’s ward, Nalufenya for the great work together with Ms. Kasuswa Sophia, Mr. Kasibante Samuel, and similarly Ms. Matinyi Sandra. The Infectious Diseases Institute, Makerere University College of Health Sciences is applauded for facilitating the provision of microbiology services at this site. Research writing was conducted with support from the Centers for Antimicrobial Optimization Network (CAMO-Net) Uganda.
Data Summary
Data are available on reasonable request.Antimicrobial Susceptibility Test (AST) results are summarized in Table S1.
Competing Interest
The author declares no competing interests.
Author Contributions
Conceptualization, F.L, J.M, R.W, F.K, M.L; methodology, F.L, G.A, J.M, B.W.W; data curation, F.L, M.J, G.A, M.N, R.M, A.B, H.M; formal analysis, F.L, M.N, R.M, H.M; investigation, F.L, J.M, M.J, G.A, A.B, B.W.W; writing—original draft preparation, F.L.; writing—review and editing, J.M, G.A, M.J, R.K, A.B, B.W.W, F.K, R.W, H.M, M.H, M.J, G.A, M.L; All authors have read and agreed to the published version of the manuscript.
References
- Timsit, J. F., Ruppé, E., Barbier, F., Tabah, A., & Bassetti, M. (2020). Bloodstream infections in critically ill patients: an expert statement. Intensive care medicine, 46(2), 266-284.
- Santoro, A., Franceschini, E., Meschiari, M., Menozzi, M., Zona, S., Venturelli, C., ... & Mussini, C. (2020, November). Epidemiology and risk factors associated with mortality inconsecutive patients with bacterial bloodstream infection: impact of MDR and XDR bacteria. In Open forum infectious diseases (Vol. 7, No. 11, p. ofaa461). US: Oxford University Press.
- Duan, N., Sun, L., Huang, C., Li, H., & Cheng, B. (2021). Microbial distribution and antibiotic susceptibility of bloodstream infections in different intensive care units. Frontiers in microbiology, 12, 792282.
- Hussain, M. S., Basher, H., Lone M. S., Wani, K. (2019). Bloodstream Infections in Pediatric Population in a Tertiary Care Hospital.
- Bloodstream Infection Event (Central Line-Associated Bloodstream Infection and Non-central Line Associated Bloodstream Infection). ( 2023). Accessed: 2022, December 29, 2022:
- CLSI: Principles and Procedures for Blood Cultures; Aproved Guideline.CLSI document M47-A. Wayne, PA: Clinical and Laboratory Standards Institute. 2007:110.
- Sarkar, S. K., Bhattacharyya, A., Paria, K., & Mandal, S.M. (2018). A retrospective study on bacteria causing blood stream infection: antibiotics resistance and management. Indian Journal of Pharmaceutical Sciences, 80(3).
- Tian, L., Zhang, Z., Sun, Z. (2019). Antimicrobial resistance trends in bloodstream infections at a large teaching hospital in China: a 20-year surveillance study (1998-2017). Antimicrobial resistance & infection control, 8, 1-8.
- Lubwama, M., Phipps, W., Najjuka, C. F., Kajumbula, H., Ddungu, H., Kambugu, J. B., & Bwanga, F. (2019). Bacteremia in febrile cancer patients in Uganda. BMC research notes, 12(1), 464.
- Zou, Q., Zou, H., Shen, Y., Yu, L., Zhou, W., Sheng, C., ...& Li, C. (2021). Pathogenic spectrum and resistance pattern of bloodstream infections isolated from postpartum women: a multicenter retrospective study. Infection and drug resistance, 2387-2395.
- Lwigale, F., Kibombo, D., Kasango, S. D., Tabajjwa, D., Atuheire, C., Kungu, J., ... & Rwego, I. B. (2024). Prevalence, resistance profiles and factors associated with skin and soft-tissue infections at Jinja regional referral hospital: A retrospective study. PLOS Global Public Health, 4(8), e0003582.
- Kisame, R., Najjemba, R., van Griensven, J., Kitutu, F. E., Takarinda, K., Thekkur, P., ... & Lamorde, M. (2021). Blood culture testing outcomes among non-malarial febrile children at antimicrobial resistance surveillance sites in Uganda, 2017– 2018. Tropical Medicine and Infectious Disease, 6(2), 71.
- Bandy, A., & Almaeen, A. H. (2020). Pathogenic spectrum of blood stream infections and resistance pattern in Gram- negative bacteria from Aljouf region of Saudi Arabia. PLOS one, 15(6), e0233704.
- Abu,Taha. S. A., Al-Kharraz, T., Belkebir, S., Abu, Taha. A., Zyoud, she. (2022). Patterns of microbial resistance in bloodstream infections of hemodialysis patients: a cross- sectional study from Palestine. Scientific Reports.12,1-10.
- Yang, S., Xu, H., Sun, J., & Sun, S. (2019). Shifting trends and age distribution of ESKAPEEc resistance in bloodstream infection, Southwest China, 2012–2017. Antimicrobial Resistance & Infection Control, 8(1), 61.
- Seni, J., Mwakyoma, A. A., Mashuda, F., Marando, R., Ahmed, M., DeVinney, R., ... & Mshana, S. E. (2019). Deciphering risk factors for blood stream infections, bacteria species and antimicrobial resistance profiles among children under five years of age in North-Western Tanzania: a multicentre study in a cascade of referral health care system. BMC pediatrics, 19(1), 32.
- Ministry of Health U. (2023). Uganda Clinical Guidelines. In National Guidelines for Management of Common Health Conditions. Edited by Ministry of Health U. Ministry of Health, Uganda, Ministry of Health, Uganda website, 1095.
- Kiggundu, R., Wittenauer, R., Waswa, J. P., Nakambale, H. N., Kitutu, F. E., Murungi, M., ... & Konduri, N. (2022). Point prevalence survey of antibiotic use across 13 hospitals in Uganda. Antibiotics, 11(2), 199.
- Kutyabami, P., Munanura, E. I., Kalidi, R., Balikuna, S., Ndagire, M., Kaggwa, B., ... & Ndagije, H. B. (2021). Evaluation of the clinical use of ceftriaxone among in-patients in selected health facilities in Uganda. Antibiotics, 10(7), 779.
- Kizito, M., Lalitha, R., Kajumbula, H., Ssenyonga, R., Muyanja, D., & Byakika-Kibwika, P. (2021). Antibiotic prevalence study and factors influencing prescription of WHO watch category antibiotic ceftriaxone in a tertiary care private not for profit hospital in Uganda. Antibiotics, 10(10), 1167.
- Winsley, T. J., Snape, I., McKinlay, J., et al. (2014). The ecological controls on the prevalence of candidate division TM7 in polar regions. Frontiers in microbiology. 5, 345.
- Paul, M., Bhatia, M., Rekha, U. S., Omar, B. J., & Gupta, P. (2020). Microbiological profile of blood stream infections in febrile neutropenic patients at a tertiary care teaching hospital in Rishikesh, Uttarakhand. Journal of Laboratory Physicians, 12(02), 147-153.
- Wayne, P. (2020). CLSI performance standards for antimicrobial susceptibility testing. CLSI supplements M, 100, 20-30.
- Organization WH. (2021). Global Antimicrobial Resistance Surveillance System (GLASS). Edited by Organization WH. 67.
- Organization WH: 29 April 2010 The burden of health care-associated infection worldwide. https://www.who. int/news-room/feature-stories/detail/the-burden-of-health-care-associated-infection-worldwide#:~:text=Health%20 care%2Dassociated%20infection%20(HAI,have%20 solved%20the%20problem%20yet. Accessed: 20-11-2024.
- CLSI: Performance Standards for Antimicrobial Susceptibility Testing.28th ed. CLSI guideline M100. Wayne, PA: Clinical and Laboratory Standards Institute 2018.
- CLSI: Performance Standards for Antimicrobial Susceptibility Testing.30th ed. CLSI guideline M100. Wayne, PA: Clinical and Laboratory Standards Institute. 2020.
- CLSI: Analysis and Presentation of Cumulative Antimicrobial Susceptibility Data; Approved Guideline-Fourth Edition. CLSI guideline M39-A4. Wayne, PA: Clinical and Laboratory Standards Institute. 2014.
- Myat, T. O., Oo, K. M., Mone, H. K., Htike, W. W., Biswas, A., Hannaway, R. F., ... & Crump, J. A. (2020). A prospective study of bloodstream infections among febrile adolescents and adults attending Yangon General Hospital, Yangon, Myanmar. PLoS neglected tropical diseases, 14(4), e0008268.
- Marchello, C. S., Dale, A. P., Pisharody, S., Rubach, M. P., Crump, J. A. (2019). A systematic review and meta-analysis of the prevalence of community-onset bloodstream infections among hospitalized patients in Africa and Asia. Antimicrobial agents and chemotherapy. 64, e01974-01919.
- Ombelet, S., Kpossou, G., Kotchare, C., Agbobli, E., Sogbo, F., Massou, F., ... & Jacobs, J. (2022). Blood culture surveillance in a secondary care hospital in Benin: epidemiology of bloodstream infection pathogens and antimicrobial resistance. BMC infectious diseases, 22(1), 119.
- Arega, B., Woldeamanuel, Y., Adane, K., Sherif, A. A., & Asrat, D. (2018). Microbial spectrum and drug-resistance profile of isolates causing bloodstream infections in febrile cancer patients at a referral hospital in Addis Ababa, Ethiopia. Infection and drug resistance, 1511-1519.
- Li, Z., Zhuang, H., Wang, G., Wang, H., & Dong, Y. (2021). Prevalence, predictors, and mortality of bloodstream infections due to methicillin-resistant Staphylococcus aureus in patients with malignancy: systemic review and meta-analysis. BMCinfectious diseases, 21(1), 74.
- Habyarimana, T., Murenzi, D., Musoni, E., Yadufashije, C., & N Niyonzima, F. (2021). Bacteriological profile and antimicrobial susceptibility patterns of bloodstream infection at Kigali University Teaching Hospital. Infection and drug resistance, 699-707.
- Kokkayil, P., Agarwal, R., Mohapatra, S., Bakshi, S., Das, B., Sood, S., ... & Kapil, A. (2018). Bacterial profile and antibiogram of blood stream infections in febrile neutropenic patients with haematological malignancies. The Journal of Infection in Developing Countries, 12(06), 442-447.
- Gezmu, A. M., Bulabula, A. N., Dramowski, A., Bekker, A., Aucamp, M., Souda, S., & Nakstad, B. (2021). Laboratory- confirmed bloodstream infections in two large neonatal units in sub-Saharan Africa. International Journal of Infectious Diseases, 103, 201-207.
- Doern, G. V., Carroll, K. C., Diekema, D. J., Garey, K.W., Rupp, M. E., Weinstein, M. P., & Sexton, D. J. (2019). Practical guidance for clinical microbiology laboratories: a comprehensive update on the problem of blood culture contamination and a discussion of methods for addressing the problem. Clinical microbiology reviews, 33(1), 10-1128.
- Chaplain, D., Asutaku, B. B., Mona, M., Bulafu, D., & Aruhomukama, D. (2022). The need to improve antimicrobial susceptibility testing capacity in Ugandan health facilities: insights from a surveillance primer. Antimicrobial Resistance & Infection Control, 11(1), 23.
Supplementary Table S1: Frequency and Percentage of Organisms Tested Against Different Antibiotics
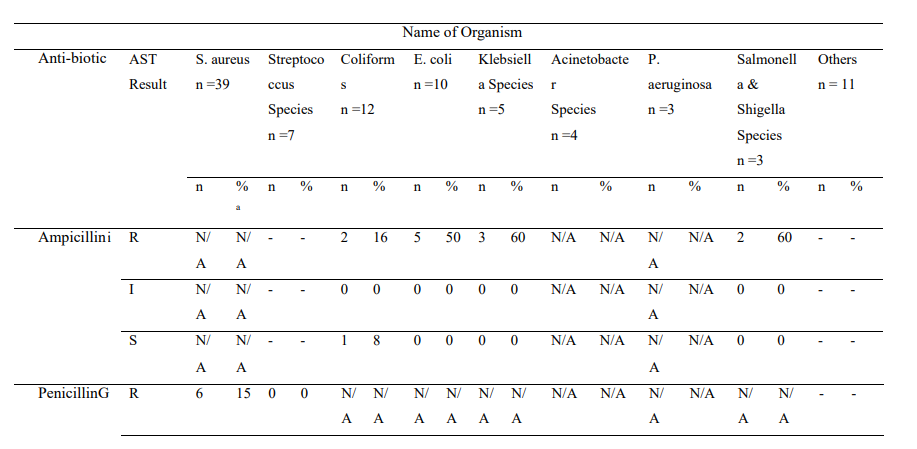


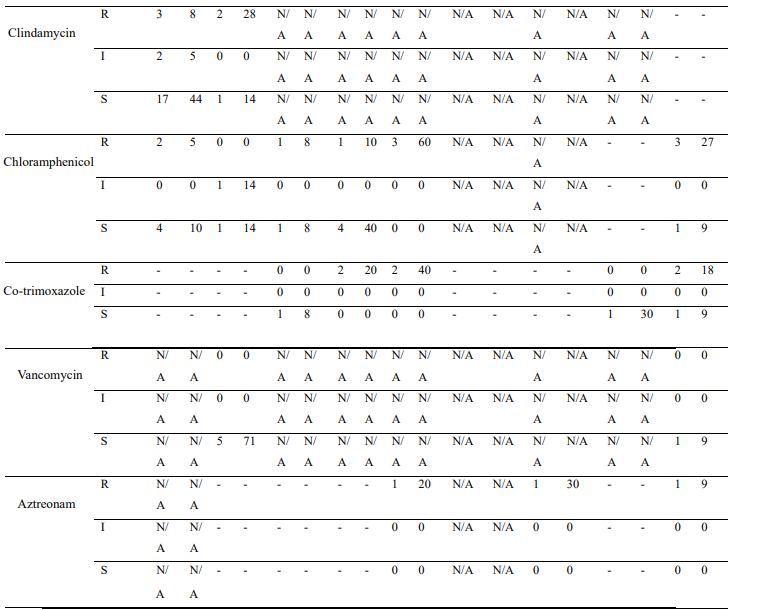
AST, Antimicrobial Susceptibility Test; I, Intermediate; R, Resistant; S, Susceptible; n, Number of isolates; -, Data was unavailable aPercentages were calculated for isolates (per AST result) relative to the number of isolated organisms
N/A, Not Applicable (The organism-antibiotic combination is not recommended. Others cannot be reliably tested by disk diffusion method.)



