Research Article - (2025) Volume 4, Issue 2
Antibacterial-Resistant Genes (Abrg) Associated with Bloodstream Infections in Patients Receiving Treatment at Nigeria Airforce Medical Center Onikan, Lagos State
2Department of Public Health, Faculty of Allied Health Sciences, David Umahi Federal University of He, Nigeria
3Liverpool John Moores University, James Parsons Building, 3 Byrom St. Liverpool L3 3AF, Nigeria
4Department of microbiology, University of Agriculture and Environmental Sciences, Umuagwo, Imo State, Nigeria
5Department of Biological Sciences, University of Africa, Toru Orua, Bayelsa State, Nigeria
Received Date: Apr 08, 2025 / Accepted Date: May 20, 2025 / Published Date: May 31, 2025
Copyright: ©Â©2025 Uzochukwu G. Ekeleme, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Ukachukwu, C. C., Ekeleme, G. U., Iwuala, C. C., Dozie, U. W., Okereke, C. C. A., et al. (2025). Antibacterial-Resistant Genes (Abrg) Associated With Bloodstream Infections In Patients Receiving Treatment At Nigeria Airforce Medical Center Onikan, Lagos State. Archives Clin Med Microbiol, 4(2), 01-13.
Abstract
Background/Objectives: The antibacterial-resistant genes (ABRG) associated with bloodstream infections in patients (BSI-PAP) receiving treatment at the Nigerian Air Force Medical Centre, Onikan, Lagos State, were studied from October 2023 to June 2024. BSI-PAP was defined as bloodstream infections diagnosed within 48 hours of hospitalization.
Methods: Blood samples were analyzed for bacterial contamination and antibiotic resistance, adhering to standard microbiological protocols.
Results: The prevalence of BSI-PAP was found to be at 9.5%. The likelihood of being admitted with BSI-PAP was highest among patients with diabetes mellitus, previous hospitalization, renal failure, and chronic dermatitis. The odds ratios (OR) with 95% confidence intervals (CI) were as follows: Diabetes mellitus: OR = 4.96 (95% CI = 1.37–7.32), previous hospitalization: OR = 2.59 (95% CI = 0.29–4.9), renal failure: OR = 2.39 (95% CI = −0.17–4.89), chronic dermatitis: OR = 1.32 (95% CI = 0.37–4.29). Among the patients admitted with BSI-PAP, the following bacterial species were identified: S. scuri ssp. lentus (35.3%), S. gallinarum (17.6%), S. eqourum (17.6%), E. cloacae ssp (11.8%), B. capacia complex (5.9%), S. xylulosus (5.9%), C. freundi (5.9%). Gram-negative bacteria accounted for a greater proportion of BSI cases (56.7%) compared to gram-positive bacteria (43.3%). Approximately 76% of BSI-PAP cases with these organisms exhibited antibiotic resistance. Resistance to erythromycin, sulfamethoxazole, ciprofloxacin, and ampicillin was most prevalent in patients aged 30 years. A high prevalence of resistant genes was observed, including TEM, CTX-M, SHV, and VEB types. The bacterial isolates (S. scuri ssp. lentus, S. gallinarum, S. eqourum, and S. xylulosus) demonstrated high antimicrobial resistance to commonly prescribed antibiotics, particularly gentamicin, erythromycin, sulfamethoxazole, and tetracycline.
Keywords
Bloodstream Infections, Bacteria, Prevalence, Susceptibility, Resistant Genes
Introduction
The effectiveness of antimicrobial agents in fighting bloodstream infectious diseases is limited by antimicrobial resistance (AMR) a growing global health concern [1]. The ranks AMR among the top ten global public health threats [2]. Overuse and misuse of antibiotics in healthcare and other industries have led to the emergence and spread of antimicrobial-resistant pathogens. In 2019, antimicrobial resistance directly caused about 1 in 27 million of the 495 million deaths linked to the disease. Sub-Saharan Africa and South Asia had the highest rates of antimicrobial resistancerelated deaths, with 23.5 deaths per 100,000 and 21.5 deaths per 100,000 population, respectively, according to the Global Burden of Disease analysis [3].
Bloodstream infections (BSIs) remain a significant global public health threat, despite significant advancements in management and therapy [4]. In the United States, they have a high mortality rate, resulting in an estimated 200,000 deaths annually [5]. Early treatment of significant bacteremia and sepsis is crucial for achieving good clinical outcomes [6]. The prevalence of antimicrobial-resistant pattern-related bloodstream infections in admitted patients (BSI-PAP) has increased recently, and it is a potentially fatal illness that consumes substantial resources and incurs significant costs [7]. The overall population-based incidence of BSI-PAP in Canada, Europe, and the US ranges from 81.6 to 189 per 100,000 person-years, with a reported incidence of 18.6 per 1000 discharges in 2013 [8]. Concerns have been raised about the increasing incidence of BSI-PAP, particularly concerning multiple-resistant organisms [9]. Although BSI has been the subject of several population-based, the rise of antimicrobial resistance jeopardizes the effectiveness of healthcare systems, highlighting the urgent need for comprehensive research and interventions [10,11]. Of particular concern are bloodstream infections caused by antimicrobial-resistant patterns in patients receiving medical care in hospitals. Treating these infections can be challenging due to the limited availability of effective antimicrobial agents that target antimicrobial-resistant strains. This study aims to identify the antibacterial-resistant genes associated with bloodstream infections in patients receiving treatment at the Nigeria Airforce Medical Center Onikan, Lagos State.
Materials and Methods
Study Design
This study adopted the use of a cross-sectional study design approach.
Study Area
The study was conducted at the Nigerian Air Force Medical Centre, Onikan. The medical centre is a military health care organisation located in Lagos Island. The medical centre attends to the medical needs of residents of Lagos Island and its surroundings.
Study Population
The population of this study was made up of patients receiving treatment from October 2023 to June 2024 at the Nigerian Air Force Medical Centre, Onikan.
Sample Size and Sampling Method
Determination of Sample Size
The sample size was determined using the formula (n= (Z/2)2 P (1-P)/d2), as was reviewed in similar studies reported by [12]. The maximum sample size was obtained from a study conducted in southern Nigeria where the prevalence/proportion of bloodstream infections was 28.09% (0.281) [12]. As a result, n= (Z/2)2 P (1- P) /d2 with a margin of error (d=0.05) and a 95% confidence interval. p=0.281, d=0.05, Z=0.05=Z/2=0.025=1.96. As a result, n= (1.96)2 x 0.281(1-0.281)/ (0.05)2 =310. Then, adjusting for a 5% non-response rate, the minimum sample size was 315. 3.2.
Inclusion and Exclusion Criteria
This study included patients of both sexes and all ages with suspected and proven bloodstream infections. Patients who were critically ill and unable to provide a blood sample during the data collection period were excluded from the study. The study also excluded patients who were released from the hospital and then readmitted with a BSI within 30 days [13].
Sampling Method
A convenience sampling technique was used. This was achieved when the physician attending to the patients suspects bloodstream infection in any of the hospital wards.
Method of Data Collection, Culture and Identification of Antibacterial-Resistant Genes
Data Collection Using a Semi-Structured Questionnaire
The sociodemographic data, and information on risk factors such as recent invasive procedures, immunosuppression, and underlying health conditions were collected from the hospitalized patients and/or their caregivers using a semi-structured questionnaire with the assistance of trained nursing staff and laboratory personnel of the hospital. Also, some of the patient information was collected from the health records, including age, gender, comorbidities, length of hospital stay, and any prior antibiotic exposure.
Blood Samples Collection and Culture
Based on the National Healthcare Safety Network (NHSN) definitions, as provided by the Centre for Disease Control and Prevention (CDC) in 2016. Patients with BSI-PAP were defined as those who had a BSI within 48 hours of admission. The patient's blood collection was performed aseptically, ensuring strict adherence to sterile techniques to avoid contamination. The venipuncture site was properly disinfected with 70% alcohol and 2% iodine tincture. Within a 30-minute interval, two bottles of blood for each patient (5 ml of blood for patients over five years of age and 2 ml of blood for patients under five years of age) were aseptically collected from any of the hands as described by [14]. The collected sample was inoculated into a blood culture bottle containing Brain Heart Infusion (BHI) broth as a suitable growth medium. The culture flask was incubated at 37°C for 24 to 48 hours to promote microbial growth. The culture was monitored regularly to detect signs of microbial proliferation by looking for turbidity and or colour changes in the medium. When microbial growth was observed, the sample was sub-cultured on solid agar media including MacConkey agar, blood agar, and chocolate agar for isolation and identification of the bacteria.
Identification of the Bacterial Isolates
Gram staining was used to identify the gram reaction of the bacterial isolates that showed positive blood cultures. Traditional biochemical assays and other various methods including, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-MS), and molecular techniques such as polymerase chain reaction (PCR) were used to further identify the bacterial isolates [15].
Antimicrobial Susceptibility Testing (AST)
The disk diffusion method as described by was used to perform the AST [16]. The antimicrobial susceptibility of each of the blood culture isolates was assessed following the guidelines of the Clinical and Laboratory Standards Institute [17]. On nutrient agar, a new subculture of every isolate was initially created. Each isolate was incubated overnight at 37ºC after five colonies were touched with a sterile straight wire and suspended in a sterile Bijou bottle with five millilitres of peptone water (Lab M). Using sterile saline, the overnight broth cultures were diluted to 106 colony-forming units per millilitre by comparing the inoculum's turbidity to the 0.5 McFarland turbidity standards. Mueller-Hinton agar (Oxoid, England) plates were dried and inoculated using a sterile cottontipped applicator mixed with a standardized inoculum. Each plate had sterile antibiotic discs, which were then incubated aerobically at 37ºC for 24 hours. Following the standardized CLSI 2021 guidelines, the zone diameter of inhibition of each isolate to the disc was measured using a calibrated ruler [17,18].
On Mueller-Hinton agar supplemented with 2 per cent NaCl, staphylococcal isolates exhibiting gentamicin resistance were identified, as previously reported [19]. Gentamicin 10μg (Gen), erythromycin 10μg (Er), sulfamethoxazole 10μg (Sxt), ciprofloxacin 10μg (Cip), ampicillin 10μg (Amp), tetracycline 10μg (Tet), and cefotaxime 10μg (Ctx) were the antibiotics used for Gram-positive isolates, whereas erythromycin 10μg (Er), sulfamethoxazole 10μg (Sxt), ciprofloxacin 10μg (Cip), ampicillin 10μg (Amp), ampicillin 10μg (Cip), ampicillin 10μg (Amp), tetracycline 10μg (Tet), cefotaxime 5μg (Ctx), and ampicillinsubitan 30μg (Sam) were the antibiotics for Gram-negative isolates. To identify infections and patterns of antimicrobial susceptibility, the algorithms used microbiology results and ICD-9-CM billing codes for the following seven organisms: Citrobacter freundi, Burkholderia cepacia-komplex, Staphylococcus eqourum, Staphylococcus gallinarum, Staphylococcus Scurissp lentus, Staphylococcus Xylulosus, Enterobacter cloacae ssp, and Staphylococcus eqourum. These organisms were chosen because they were among the most common causes of BSI and frequently develop resistance to one or more antimicrobials [11].
Extended-Spectrum Beta-Lactamase (ESBL) Screening
The isolates were screened by one of these drugs (30 μg of erythromycin, sulfamethoxazole, ciprofloxacin or gentamicin, tetracycline or ampicillin-subitan) on a standard disk diffusion procedure. The zones 29 mm for erythromycin, 20 mm for tetracycline or 10 mm for ampicillin-subitan, 24 mm for gentamicin, or 30 mm for ciprofloxacin were considered as probable ESBL producers which were further confirmed by double disk diffusion testing.
Extended-Spectrum β Lactamase Confirmation by Double Disk Diffusion Testing
The disks containing erythromycin with and without absolute ethanol (8ml) were placed 20mm apart on the surface of a Kigler’s Iron Agar (KIA) plate previously inoculated with the test organism and incubated at 56°C for 10 minutes. A test result was considered positive if there was an increase of 5mm or higher in the zone diameter for the combination of antimicrobial agent and absolute ethanol compared with the antibiotic alone. The test was done in cognisance with the Clinical and Laboratory Standards Institute (CLSI) document M100-S21 [20]. Staphylococcus scurissp lentus was used as the ESBL-positive control, and Barkholderia cepaciakomplex was used as the negative control. The disks containing erythromycin with and without absolute ethanol (8ml) were placed 20mm apart on the surface of a Kigler’s Iron Agar (KIA) plate previously inoculated with the test organism and incubated at 56°C for 10 minutes. A test result was considered positive if there was an increase of 5mm or higher in the zone diameter for the combination of antimicrobial agent and absolute ethanol compared with the antibiotic alone. The test was done in cognisance with the Clinical and Laboratory Standards Institute (CLSI) document M100-S21 [20]. Staphylococcus scurissp lentus was used as the ESBL-positive control, and Barkholderia cepaciakomplex was used as the negative control.
The Genomic DNA Extraction
The specimen (100 µl) was added into a micro-centrifuge tube, followed by the addition of 500 µl of Lysis Buffer. The mixture was then vortexed and incubated at 56ºC for 10 minutes. After incubation, it was centrifuged at 10,000 rpm for 1 minute. Subsequently, 200 µl of absolute ethanol was added to the tube, and the mixture was transferred into a spin column.
The spin column was centrifuged at 10,000 rpm for 30 seconds, after which the flow-through was discarded, and the collection tube was blotted on tissue paper. Next, 500 µl of Wash Buffer 1 was added to the spin column, followed by centrifugation at 10,000 rpm for 1 minute. The flow-through was discarded again, and the collection tube was blotted on tissue paper. This was repeated with 500 µl of Wash Buffer 2, centrifuged at 10,000 rpm for 1 minute, and the flow-through was discarded as before.
All traces of ethanol were removed, and the spin column was centrifuged once more at 12,000 to 14,000 rpm for 3 minutes. The spin column was then placed into another micro-centrifuge tube, and 50 µl of Elution Buffer or nuclease-free water was added to the centre of the column. The setup was incubated at room temperature for 1 to 2 minutes and centrifuged at 10,000 rpm for 1 minute to elute the DNA. Finally, the DNA was stored at -20ºC or -80ºC.
Detection of Antibacterial Resistance Genes
The genotypic analysis for erythromycin ribosome methylation gene F (ermF), erythromycin ribosome methylation gene X (ermX), erythromycin ribosome methylation gene A (ereA), beta-lactamase methicillin-resistance Staphylococcus scurissp lentus (blaMsrS), vietnamese extended-spectrum beta-lactamase (VEB), cefotaximase-type extended-spectrum beta-lactamase (CTX-M), betal-lactamase Klebsiella pneumonia (blaSHV), and beta-lactamase and enzyme (blaTEM), were performed to confirm the resistant isolates. The antimicrobial-resistant pathogens as described by (2021) were determined by selecting specific genes associated with antimicrobial resistance, and amplifying them through a series of temperature changes using DNA primers and DNA polymerase [21]. The amplified DNA was then analyzed for the presence of antimicrobial resistance genes using techniques such as gel electrophoresis. Finally, the results were compared to known resistance gene sequences to confirm the presence of specific resistance genes. The primers of the genes used are described in Table 1.
|
Targets |
Forward Sequence |
Reverse Sequence |
Band size |
|
ermF |
CGA CAC AGC TTT GGT TGA AC |
QQA CCT ACC TCA TAG ACA AG |
309 |
|
ermX |
GAG ATC GGR CCA GGA AGC |
GTG TGC ACC ATC GCC TGA |
488 |
|
ereA |
GCC GGT GCT CAT CAT GAA CTT GAG |
CGA CTC TAT TCG ATC AGA GGC |
420 |
|
MsrS |
GCA CTT ATT GGG GGT AAT GG |
GTC TAT AAG TGC TCT ATC GTG |
384 |
|
VEB |
GAT GGT GTT TGG TCG CAT ATC GCA AC |
CAT CGC TGT TGG GGT TGC CCA ATT TT |
391 |
|
TEM |
TCG CCG CAT ACA CTA TTC TCA AGA ATGAC |
CAG CAA TAA ACC AGC CAG CCG GAA G |
422 |
|
CTX-M |
ATG TGC AGY ACC AGT AAR GTK ATGGC |
GGT RAA RTA RGT SACC AGA AYC AGC GG |
590 |
|
SHV |
TGT ATT ATCTC(C/T) CTG TTA GCC(A/G) CCCTG |
GCT CTG CTT TGT TAT TCG GGC CAA GC |
739 |
Table 1: Primer Sequence used for Resistance Gene Detection
Method of Data Analysis
After coding and properly sorting the data in a Microsoft Excel spreadsheet, we transferred it to SPSS version 25 software for analysis. To ensure the data's consistency and completeness, we used a cross-tabulation of all variables. We then calculated the descriptive statistics, including percentages, frequencies, means, and standard deviations (SD), for all variables. Next, we performed bivariate analyses to determine the relationships between BSI-PAP and potential predictor variables, such as source of admission, age, gender, previous hospitalization, renal failure, diabetes mellitus, malignancy, and chronic dermatitis. We used chi-square tests for categorical variables and the Wilcoxon rank sum test for confidence class interval (CCI). Multiple logistic regression was then used to include the variables significantly associated with BSI-HOA in the bivariate analyses (P < 0.05). Due to the small sample size (n = 2), we excluded malignancies from the multivariable analysis. Additionally, we excluded CCI from the multivariable analysis since some of the predictors in our model were included in the CCI composite score.
Results
Socio-Demographic Information of the Study Participants
A total of 315 patients were sampled in this study on antibacterialresistant genes associated with bloodstream infections present in admitted patients at Nigeria Air Force Medical Centre, Onikan Lagos State. All patients had received at least one antimicrobial agent during the study period. Out of these, 30 patients (10%) had bloodstream infections. Among these 30 patients, 21 (71%) were male and 9 (29%) were female as shown in table 2. The participants' educational backgrounds were as follows: 14 (47%) had a secondary school certificate, 4 (24%) had technical/ vocational training, and 9 (29%) had a Bachelor's degree. The lifestyle and habits of the participants were as follows: 26 (88%) did not smoke, while 4 (12%) admitted to smoking.
|
Variables |
Frequency |
Percentage (%) |
|
Age (years) |
|
|
|
18- 24 |
11 |
35 |
|
25 – 29 |
9 |
29 |
|
30 – 34 |
5 |
18 |
|
35 - 39 |
1 |
6 |
|
40 – 44 |
4 |
12 |
|
Total |
17 |
|
|
Mean |
29.29 |
|
|
Standard Deviation |
7.447 |
|
|
Gender |
|
|
|
Male |
21 |
71 |
|
Female |
9 |
29 |
|
Mean |
1.29 |
|
|
Standard Deviation |
0.469 |
|
|
Educational Background |
|
|
|
Secondary |
14 |
47 |
|
Technical/Vocational Training |
7 |
24 |
|
Bachelor’s Degree |
9 |
29 |
|
Mean |
3.79 |
|
|
Standard Deviation |
0.893 |
|
|
Health History |
|
|
|
Diagnosed with Chronic Medical Condition |
|
|
|
YES |
30 |
100 |
|
NO |
0 |
0 |
|
Mean |
2.00 |
|
|
Standard Deviation |
0.000 |
|
|
Specify the Chronic Disease |
|
|
|
Diabetes Mellitus |
4 |
12 |
|
Malignancies |
2 |
6 |
|
Renal Failure |
9 |
29 |
|
Chronic Dermatitis |
5 |
18 |
|
Urinary tract infection |
11 |
35 |
|
Currently Taking Antibiotics |
|
|
|
YES |
30 |
100 |
|
NO |
0 |
0 |
|
Mean |
2.00 |
|
|
Standard Deviation |
0.000 |
|
|
Antibiotics Intake |
|
|
|
Within the last 3 months |
9 |
29 |
|
Within the last 6 months |
16 |
53 |
|
Within a year |
5 |
18 |
|
Mean |
4.00 |
|
|
Standard Deviation |
0.679 |
|
|
Surgeries for last 6 Months |
|
|
|
YES |
0 |
0 |
|
NO |
30 |
100 |
|
Lifestyle and Habits |
|
|
|
Smoking |
|
|
|
YES |
4 |
12 |
|
NO |
26 |
88 |
|
Mean |
1.93 |
|
|
Standard Deviation |
0.267 |
|
|
Consume Alcoholic Beverages |
|
|
|
Never |
12 |
35 |
|
Occasionally |
14 |
47 |
|
Regularly |
4 |
18 |
|
Mean |
1.71 |
|
|
Standard Deviation |
0.726 |
|
|
Do you engage in regular exercise? |
|
|
|
YES |
5 |
15 |
|
NO |
25 |
85 |
|
Mean |
1.79 |
|
|
Standard Deviation |
0.426 |
|
Table 2: Socio-Demographic Information of the Study Participants
Figure 1 below revealed the ages of the participants with ages of 18 to 24 years having the highest frequency. The mean was 29.29 and the SD was 7.447 which indicates less variability.
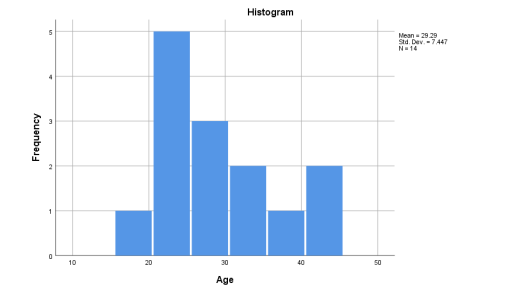
Figure 1: The Participants’ Age With Febrile Illness.
Prevalence and Co-Morbid Conditions of Bloodstream Infections Among the Admitted Patients (BSI-PAP)
The patients were 315, out of which 30 (9.5%) cases were of bloodstream infection admitted. The male patients were the most admitted with a BSI (71%) than the female (29%). The urinary tract infection, diabetes mellitus, renal failure, chronic dermatitis and malignancies were significantly associated with BSI-PAP in bivariate analyses. The mean score (mean = 3.8 ± SD = 3.5) of patients with BSI-PAP was significantly higher, compared with those patients without BSI-PAP (mean = 2.8 ± SD = 3.3), P < 0.001 (Table 3).
|
|
BSI-PAP (n = 30) (%) |
Non- BSI-PAP (n = 285) (%) |
P-value |
|
Laboratory Culture Media Source Eosin Methylene Blue (EMB) Salmonella-Shigella Agar (SSA) MacConkey Agar (MAC) |
14 (47) 12 (41) 4 (12) |
137 (48) 131 (46) 17 (6) |
< 0.001 |
|
Age Category (years) 18 – 24 25 – 29 30 – 34 35 – 39 40 – 44 |
11 (35) 9 (29) 5 (18) 1 (6) 4 (12) |
66 (23) 51 (18) 100 (35) 23 (8) 46 (16) |
< 0.001 |
|
Gender Male Female |
21 (71) 9 (29) |
202 (71) 83 (29) |
< 0.001 |
|
Co-Morbid Conditions Urinary tract infection Diabetes mellitus Renal failure Chronic Dermatitis Malignancies CCI Scores: Mean [ Standard Deviation] |
11 (53) 4 (29) 9 (35) 5 (18) 2 (12) 3.8 [ 3.5] |
94 (33) 51 (18) 68 (24) 48 (17) 26 (9) 2.8 [ 3.3] |
< 0.001 < 0.001 < 0.001 < 0.001 < 0.001 |
Table 3: Prevalence and Co-Morbid Conditions of Bloodstream Infections Among the Admitted Patients (Bsi-Pap)
The Risk Factors and Predictors Contributing to the Development of AMRP-Associated Bloodstream Infections Present in Admitted Patients (BSI-PAP).
Table 4 shows the multivariable analysis of the association between Patients’ characteristics and BSI-PAP, the odds of being admitted with BSI-PAP were greatest among patients admitted with diabetes mellitus, urinary tract infection, renal failure and chronic dermatitis. The odds ratios 95% confidence interval (CI) were 4.96 (95% CI = 1.37-7.32), 2.59 (95% CI = 0.29-4.9), 2.39 (95% CI = -0.17-4.89) and 1.32 (95% CI = 0.37-4.29) and, respectively. There was no association between ages from 30 to 44 years, female gender and EMB culture media.
|
Predictors |
Odds Ratio (OR) |
95% Confidence Interval (C.I) |
|
|
Lower |
Upper |
||
|
Co-Morbid Conditions |
|
|
|
|
Urinary tract infections |
2.5899 |
0.2868 |
4.9367 |
|
Diabetes Mellitus |
4.9566 |
1.3728 |
7.3295 |
|
Renal Failure |
2.3855 |
-0.1677 |
4.8943 |
|
Chronic Dermatitis |
1.3193 |
0.3668 |
4.2941 |
|
Age (In years) |
|
|
|
|
18 – 24 |
1.8777 |
0.6950 |
5.0723 |
|
25 – 29 |
1.8613 |
0.6274 |
5.5220 |
|
30 – 34 |
0.3137 |
0.0718 |
1.3698 |
|
35 – 39 |
0.8341 |
0.1073 |
6.4851 |
|
40 – 44 |
0.8536 |
0.1949 |
3.7393 |
|
Gender |
|
|
|
|
Male |
1.0213 |
0.4257 |
2.4500 |
|
Female |
0.9792 |
0.1273 |
2.4213 |
|
Laboratory Culture Media Source |
|
|
|
|
Eosin Methylene Blue (EMB) |
0.6192 |
0.2296 |
1.6698 |
|
Salmonella-Shigella Agar (SSA) |
1.2158 |
0.5003 |
2.9547 |
|
MacConkey Agaar (MAC) |
2.0354 |
0.4612 |
8.9836 |
Table 4: Association Between Patients’ Features and Bsi-Pap in Multivariable Logistic Regression Analysis.
Bacterial Isolates Associated with BSI-PAP
Table 5 shows the isolates associated with BSI-PAP among the patients admitted with BSI-PAP (n = 30), the largest proportion presented with S. scurissp Lentus (n = 6, 35.3%), followed by S. gallinarum (n = 3, 17.6%), S. eqourum (n = 3, 17.6%), E.cloacae SSP (n = 5, 11.8%), B. capacia-komplex (n = 7, 5.9%), S. xylulosus (n = 1, 5.9%), and C. freundi (n = 5, 5.9%). About 76% (13) of the BSI-PAP cases with these organisms were antibiotic-resistant.
|
Isolates |
Total |
EMB Resistant (n, %) |
SSA Resistant (n, %) |
MAC Resistant (n, %) |
P-Value |
|
S.gallinarum |
3 |
1(95) |
1 (92) |
1 (100) |
0.545* |
|
S.scuri SSP lentus |
6 |
3 (78) |
2 (79) |
1( 49) |
0.001* |
|
S.xylulosus |
1 |
0 (0) |
1 (95) |
0 (0) |
0.720f |
|
S.eqourum |
3 |
2 (62) |
1 (78) |
0 (0) |
0.003f |
|
E. cloacae SSP |
5 |
0 (0) |
0 (0) |
0 (0) |
0.842* |
|
B.cepacia-konplex |
7 |
0 (0) |
0 (0) |
0 (0) |
0.725* |
|
C.freundi |
5 |
0 (0) |
0 (0) |
0 (0) |
0.360f |
|
Total |
30 |
6 (78) |
5 (86) |
2 (75) |
0.001* |
|
*-chi-square, f-Fisher’s exact test |
|||||
Table 5: Bacterial Isolates Associated With Bsi-Pap Among the Patients Admitted With BSI-PAP
The AMRP of Bacterial Isolates from BSI-PAP Among Commonly Prescribed Antibiotics
Out of which 30 (34.7%) documented BSI, gram-negative bacteria had the highest frequency (56.7%), which are E. cloacae ssp, B. cepacia-komplex, and C. freundi. The gram-positive bacteria had 43.3% in BSI, which S. scuri ssp lentus, S. gallinarum. S. eqourum and S.xylulosus were present. The most frequently isolated bacterium was B. cepacia-komplex (23.3%) and others occurred in descending order as follows: S. scuri ssp lentus (20%), E. cloacae ssp and C. freundi (16.7%), S. gallinarum and S. eqourum (10%) and S. xylulosus (3.3%). A high level of resistance was obtained among both the gram-positive (4–100%) and gram-negative bacterial isolates (3–100%) (Table 6).
|
Bacterial isolates |
No of Isolates (%) |
|
Gram Positive |
13 (43.3) |
|
i. S.scurissp.lentus |
6 (20) GENRSS = 5 (83%) |
|
ii. S.gallinarum |
3 (10) GENRSG = 3 (100%) |
|
iii. S.xylulosus |
1 (3.3) |
|
iv. S.eqourum |
3 (10) ERSE = 3 (100%) |
|
Gram Negative |
17 (56.7) |
|
i.E.cloacae ssp |
5 (16.7) |
|
ii.B.cepacia-komplex |
7 (23.3) |
|
iii.C.freundi |
5 (16.7) |
|
Total Isolates |
30 |
|
GENRSS = Gentamicin-resistant Staphylococcus scuri ssp lentus GENRSG = Gentamicin resistant gallinarum ERSE = Erythromicin resistant Streptococcus eqourum |
|
Table 6: The Bacterial Isolates of Bloodstream Infection (BSI)
The Efficacy of the Antibiotic Combination used as first- line Empirical Treatment of BSI-PAP
Table 7 displayed the pattern of the drug resistance for both gram- positive and gram-negative isolates, this revealed single and multiple drug resistance patterns with the number of antibiotics ranging from 2 to 5 and from 2 to 6 for Gram positive and Gram negative respectively. For the drug resistance pattern for Gram- positive isolates; gentamicin erythromycin and erythromycin sulfamethoxazole tetracycline resistance phenotypes occurred most frequently among the patients aged 18-29 years (54%), while for the gram-negative isolates, the resistance phenotypes erythromycin sulfamethoxazole and erythromycin sulfamethoxazole ciprofloxacin ampicillin occurred most frequently in the age group of 30–44 years (72%).
|
Gram Positive Isolates |
||||
|
No of Antibiotics |
Resistance (R) Pattern |
No of Isolates |
Bacteria |
% |
|
1 |
Gentamicin |
1 |
S.scuri ssp |
8 |
|
2 |
Gentamicin Erythromycin |
4 |
S.xylulosus; |
31 |
|
|
|
|
S.equorum; |
|
|
|
|
|
S.gallinarum; |
|
|
|
|
|
S.scuri ssp |
|
|
|
Erythromicin Sulfamethoxazole |
2 |
S.gallinarum; |
15 |
|
|
|
|
S.scuri ssp |
|
|
3 |
Erythromicin Sulfamethoxazole Tetracycline |
3 |
S.xylulosus; |
23 |
|
|
|
|
S.equorum; |
|
|
|
|
|
S.gallinarum |
|
|
4 |
Gentamicin Erythromycin Sulfamethoxazole |
1 |
S.scuri ssp |
8 |
|
|
Ciprofloxacin |
|
|
|
|
|
Sulfamethoxazole Ciprofloxacin Ampicillin Cefotoxine |
1 |
S.scuri ssp |
8 |
|
|
Gentamicin Erythromycin Sulfamethoxazole |
|
|
|
|
5 |
Ciprofloxacin Ampicillin |
1 |
S.scuri ssp |
8 |
|
|
|
13 |
|
|
|
Gram Negative Isolates |
||||
|
1 |
Erythromicin |
3 |
B.cepacia-komplex, |
18 |
|
|
|
|
E.cleacole ssp |
|
|
|
|
|
C.freuodii |
|
|
2 |
Erythromicin Sulfamethoxazole |
3 |
B.cepacia-komplex, |
18 |
|
|
|
|
E.cleacole ssp |
|
|
|
|
|
C.freuodii |
|
|
3 |
Erythromicin Sulfamethoxazole Ciprofloxacin |
1 |
B.cepacia-komplex |
6 |
|
4 |
Erythromicin Sulfamethoxazole Ciprofloxacin |
3 |
B.cepacia-komplex, |
18 |
|
|
Ampicillin |
|
E.cleacole ssp |
|
|
|
|
|
C.freuodii |
|
|
5 |
Erythromicin Sulfamethoxazole Ciprofloxacin |
3 |
B.cepacia-komplex, |
18 |
|
|
Ampicillin Tetracycline |
|
E.cleacole ssp |
|
|
|
|
|
C.freuodii |
|
|
6 |
Erythromicin Sulfamethoxazole Ciprofloxacin |
1 |
B.cepacia-komplex |
6 |
|
|
Ampicillin Tetracycline Cefotoxime |
|
|
|
|
7 |
Erythromicin Sulfamethoxazole Ciprofloxacin |
3 |
B.cepacia-komplex, |
18 |
|
|
Ampicillin Tetracycline Cefotoxime Ampicillin-Subitan |
|
E.cleacole ssp |
|
|
|
|
|
C.freuodii A |
|
|
|
|
17 |
|
|
Table 7: The Antibiotics Resistance Pattern of Gram Positive and Gram-Negative Isolates
Extended-Spectrum Beta-Lactamase (ESBL) Screening and Confirmatory Testing
In this study, 7 bacterial isolates of E. coli (B. cepacia-komplex) and 5 bacterial isolates of K.pnemonia (E. cloacae ssp) were recovered from the hospitalized patients. Generally, 29% (2 out of 7) isolates of E. coli (B. cepacia-komplex) and 20% (1 out of 5) K. pneumonia (E. cloacae ssp) isolates showed resistance to the third- generation cephalosporin by phenotypic screening tests (Table 8).
|
Bacterial Isolates |
ESBL Screening by the disk diffusion method |
ESBL confirmed by double disk diffusion testing. |
|||
|
|
Erythromycin 30(µg) N (%) |
Tetracycline 30(µg) N (%) |
Sulfamethoxazole 30(µg) N (%) |
Ciproflaxin 30(µg) N (%) |
Erythromycin (30µg) with absolute ethanol (10 µg) N (%) |
|
E.coli.(B. cepacia- komplex) (7) |
7 (100) |
3 (43) |
6 (86) |
5 (71) |
7 (100) |
|
K.pneumonia (E.cloacae ssp) (5) |
5 (100) |
2 (40) |
4 (80) |
3 (60) |
5 (100) |
|
Total (12) |
12 (100) |
5 (42) |
10 (83) |
8 (67) |
12 (100) |
Table 8: Phenotypic Detection of ESBL Isolates.
Molecular Detection of Resistant Genes
The overall incidence of βeta-lactamase genes was found to be 30.7% (12 out of 39) which included 100% (7 out of 7) of Escherichia coli (B. cepacia-komplex) and 100% (5 of 5) of Klebsiella pneumonia (E. cloacae ssp), respectively. Molecular characterization showed that out of the 12 phenotypically positive ESBL isolates, 33% (4 out of 12) were positive for the blaCTX-M (Figure 4.3, lane 3, 4, 5, 6, 8). It consisted of 29% (2 out of 7) of Escherichia coli (B. cepacia-komplex) and 40% (2 out of 5) of Klebsiella pneumonia (E.cloacae ssp). Generally, TEM (42%) was the most common genotype followed by CTX-M (33%), SHV (17%) and VEB (8%) both in single or when combined. No isolates harbored ermF, ermX, ereA, and MsrS genes (Table 9). In total, 35% of the total ESBL-positive isolates harboured the three ESBL genes, while 61% carried two of the tested ESBL genes.
|
|
Escherichia coli (B. cepacia-komplex) (N = 7) (%) |
Klebsiella pneumonia (E.cloacae ssp) (N = 5) (%) |
Total (N = 12) (%) |
|
ermF |
0 (0) |
0 (0) |
0 (0) |
|
ermX |
0 (0) |
0 (0) |
0 (0) |
|
ereA |
0 (0) |
0 (0) |
0 (0) |
|
MsrS |
0 (0) |
0 (0) |
0 (0) |
|
VEB |
1 (14) |
0 (0) |
1 (8) |
|
CTX-M |
2 (29) |
2 (40) |
4 (33) |
|
SHV |
1 (14) |
1 (20) |
2 (17) |
|
TEM |
3 (43) |
2 (40) |
5 (42) |
Table 9: Distribution of Different Genes in Esbl-Producing Escherichia Coli (B. cepacia-komplex) and Klebsiella Pneumoniae (E. cloacae ssp) Isolates
Discussion
The Prevalence and Distribution of AntimicrobialResistant Pathogen (AMRP) causing Bloodstream Infections present in Admitted Patients (BSI-PAP).
The patients receiving healthcare treatment in Nigeria Airforce Medical Center Onikan, Lagos State from January to June 2024 were 315, out of which 30 were cases of BSI-PAP that led to a prevalence rate of 9.5%. Consistently, investigated BSI among nursing home (NH) and non-nursing home (NNH) admissions and ascertained that NH exposure was a risk factor for BSI-PAP caused by gram-positive and gram-negative organisms and reported that patients admitted for a long-term in a medical care facility was associated with gram-negative BSI-PAP [22,23]. One of the limitations of these studies was that the patients were from a single hospital, which is also applicable to the current study.
The Risk Factors and Predictors contributing to the Development of AMRP-Associated Bloodstream Infections present in Admitted Patients (BSI-PAP).
The multivariable analysis of the association between Patients’ characteristics and BSI-PAP shows that younger ages 18 to 34 years and male gender were the significant predictors of BSIPAP. This is contrary to the results of who found older age and male gender to be significant predictors of BSI-PAP (Sequel to the patient characteristics studied, patients with diabetes mellitus, prior hospitalisation, renal failure and chronic dermatitis had the highest odds ratios of being admitted with a BSI [24,25]. These findings are consistent with two previous studies of that reported renal failure and prior hospitalisation to be associated with BSI-PAP [26,27]. Generally, reported that patients with these conditions have multiple co-morbidities which include changes in immune function which influence their risk of BSI acquisition and dermatitis which menaced the skin barrier [28]. However,this finding follows the study of who reported diabetes mellitus increased the odds of BSI-PAP in chronically sick patients (OR = 1.42, 95% CI = 1.10– 1.82) () [29].
The isolate S. scuri ssp lentus was associated with BSI-PAP which accounted for almost half of the isolates among the organisms studied and had a higher proportion of resistant strains. This finding is in contrast with the reports of which reports that S. aureus accounted for most of the isolates with a higher proportion of resistant strains among the patients admitted from nursing homes [27].
The AMRP of Bacterial Isolates of BSI-PAP among Commonly Prescribed Antibiotics
During the period of this investigation, 30 (34.7%) were microbiologically documented BSI as being mono-microbial. This value follows the range reported by in Nigeria and other countries [27,30]. The Gram-negative bacteria were more frequently involved (56.7%), than the gram-positive bacteria (43.3%) in BSI. The high level of resistance to commonly available antibiotics that were used for the treatment of BSI was obtained among both the Gram-positive (4–100%) and Gram-negative bacterial isolates (3–100%). The study of has shown an association with increased mortality rates, overstay in hospital admission and costs due to BSI-PAP-resistant organisms [31].
The Efficacy of the Antibiotic Combination used as First Line Treatment of Bsi-Pap
The pattern of the drug resistance for both Gram-positive and Gram-negative isolates revealed single and multiple drug resistance patterns with the number of antibiotics ranging from two to five and from two to six for Gram-positive and Gram-negative respectively. This is consistent with the study conducted by, where the drug resistance pattern of both the Gram-positive and Gram-negative isolates revealed single and multiple drug resistance patterns with the number of antibiotics ranging from two to seven [32]. For the drug resistance occurred most frequently among the patients aged 18-29 years (54%), while for the Gram-negative isolates, the resistance phenotypes E Sxt and E Sxt Cip Amp occurred most frequently in the age group 30–44 years (72%).
Extended-Spectrum Beta-Lactamase (ESBL) Screening and Confirmatory Testing and the Molecular Analysis of the Resistant Genes
The highly prevalent co-morbidity in this study is the major issue among the patients that were hospitalized, and multidrug-resistant E. coli (B. cepacia-komplex) and K. pneumonia (E. cloacae ssp), were the most important pathogens (Prasada et al., 2019). Generally, 29% isolates of E. coli (B. cepacia-komplex) and 20% of K. pneumoniae (E. cloacae ssp) isolates showed resistance to the third-generation cephalosporin by phenotypic screening tests. The reports of had lower prevalence of 60.80% [33,34].
The overall incidence of βeta-lactamase genes was found to be 30.7%. Molecular characterization showed that out of the 12 phenotypically positive ESBL isolates, 33% was positive for the blaCTX-M, TEM (42%), SHV (17%) and VEB (8%). No isolates harbored ermF, ermX, ereA, and MsrS genes. There is a probability of community-based dissemination of ESBL-producing isolates because this is a serious issue. There is a significant implication with the high rate of various ESBL-producing genes for patient treatment and outline the benefits that strengthens the antimicrobial surveillance, antibiotic stewardship, and continuous monitoring of the rate of ESBL production along with multidrug resistance among nosocomial isolates. This aligned with a study conducted in North East districts in India, where cefotaximase-type extendedspectrum beta-lactamase (CTX-M) was found in E. coli (88.67%) and beta-lactamase and enzyme (blaTEM) in K. pneumoniae (77.58%) as the genotype that predominates the most. [35]. However, the study of identified a higher rate of cefotaximasetype extended-spectrum beta-lactamase 15 (blaCTX-M15) (100%) compared with this current study (33%) [36].
Conclusion
This study highlights a 9.5% prevalence of bloodstream infections (BSI) and identifies key risk factors, including younger adults, males, diabetes, prior hospitalization, renal failure, and chronic dermatitis. High antimicrobial resistance was noted among clinical isolates, particularly to gentamicin, erythromycin, sulfamethoxazole, and tetracycline. The study underscores the increasing complexity of healthcare delivery and its role in disseminating resistant organisms. Extended-spectrum betalactamase (ESBL) prevalence in UTIs caused by Escherichia coli and Klebsiella pneumoniae is significant, with resistance linked to TEM, CTX-M, SHV, and VEB beta-lactamases. Findings emphasize stricter control measures and the cautious use of ceftazidime in managing ESBL infections.
Declarations
Ethnical Consideration/Informed Consent
Informed written consent was sought and obtained from all study participants. Additionally, a letter of introduction (FUT/SOHT/ PUH/CS.0012/VOL.2) to carry out the study was obtained from the Department of Public Health Technology, FUTO, to the ethics committee of the Nigerian Air Force Medical Centre, Onikan, Lagos State. The study was approved by the 055 NAF Medical Centre Ethics Committee of Nigerian Air Force on 11/10/2023 with the reference number 055 CAMP/405/MEDCEN.
Consent for the publication:
Not applicable.
Availability of Data and Materials:
All data generated or analysed during this study are included in this published article.
Competing Interests:
There is no competing interest among the authors.
Funding:
This research was self-funded by the authors.
Authors' Contributions:
UGE and BCA designed research; QCO, CCI, IGC, CCO, CCU, CLO, ABN conducted research; CCAO, NIW and ECN analyzed data; COA, QCO, IGC, and CLO wrote the paper; UMC, UMD, VNU, BCA and UGE had primary responsibility for the final content. All authors read and approved the final manuscript.
Acknowledgements:
Not applicable
Authors' information:
Not applicable
Clinical trial number:
Not applicable
References
- Silveira, M. C., Marcos, C., Cassiano, I., Pontes, L. D.,Oliveira, T. R., Bastos, C. et al. (2021) Genetic basis of antimicrobial resistant Gram-negative bacteria isolated from bloodstream in Brazil. Front Med 8:635206.
- World Health Organization (WHO) (2021) Antimicrobial resistance.
- Murray, C. J., Ikuta, K. S., Sharara, F., Swetschinski, L., Aguilar, G. R., Gray, A., ... & Tasak, N. (2022). Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The lancet, 399(10325), 629-655.
- Donkor, E. S., Muhsen, K., Johnson, S. A., Kotey, F. C.,Dayie, N. T., Tetteh-Quarcoo, P. B., ... & Cohen, D. (2023). Multicenter surveillance of antimicrobial resistance among Gram-negative bacteria isolated from bloodstream infections in Ghana. Antibiotics, 12(2), 255.
- Sautter, R. L., Parrott, J. S., Nachamkin, I., Diel, C., Tom,R. J., Bobenchik, A. M., ... & Baselski, V. (2024). American Society for Microbiology evidence-based laboratory medicine practice guidelines to reduce blood culture contamination rates: a systematic review and meta-analysis. Clinical microbiology reviews, 37(4), e00087-24.
- Spellberg, B. (2020). Principles and practice of infectious diseases. 9th ed. Elsevier, Philadelphia, PA, pp 211-220.
- Søgaard, M., Nørgaard, M., Dethlefsen, C., & Schønheyder,H. C. (2011). Temporal changes in the incidence and 30- day mortality associated with bacteremia in hospitalized patients from 1992 through 2006: a population-based cohort study. Clinical infectious diseases, 52(1), 61-69.
- Laupland, K. B., Pasquill, K., Steele, L., & Parfitt, E. C. (2021). Burden of bloodstream infection in older persons: a populationâ?based study. BMC geriatrics, 21, 1-7.
- Orsini, J., Mainardi, C., Muzylo, E., Karki, N., Cohen, N., & Sakoulas, G. (2012). Microbiological profile of organisms causing bloodstream infection in critically ill patients. Journal of clinical medicine research, 4(6), 371.
- Artero, A., Zaragoza, R., Camarena, J. J., Sancho, S., González, R., & Nogueira, J. M. (2010). Prognostic factors of mortality in patients with community-acquired bloodstream infection with severe sepsis and septic shock. Journal of critical care, 25(2), 276-281.
- World Health Organization (WHO). (2015). Global action plan on antimicrobial resistance. WHO, Geneva, Switzerland.
- Ekeng, B. E., Emanghe, U. E., Monjol, B. E., Iwuafor, A. A., Ochang, E. A., & Ereh, S. E. (2021). Susceptibility Pattern of Pathogens Causing Blood Stream Infections in a Tertiary Care Hospital: A Two-year Retrospective Study from Southern Nigeria. Microbiology Research Journal International, 31(3), 53-60.
- Horan, T. C., Andrus, M., & Dudeck, M. A. (2008). CDC/ NHSN surveillance definition of health care–associated infection and criteria for specific types of infections in the acute care setting. American journal of infection control, 36(5), 309-332.
- Wyss, G., Berger, S., Haubitz, S., Fankhauser, H., Buergi, U., Mueller, B., ... & Conen, A. (2020). The Shapiro–Procalcitonin algorithm (SPA) as a decision tool for blood culture sampling: validation in a prospective cohort study. Infection, 48, 523- 533.
- Clark, A. E., Kaleta, E. J., Arora, A., & Wolk, D. M. (2013). Matrix-assisted laser desorption ionization–time of flight mass spectrometry: a fundamental shift in the routine practice of clinical microbiology. Clinical microbiology reviews, 26(3), 547-603.
- Khan, Z. A., Siddiqui, M. F., & Park, S. (2019). Current and emerging methods of antibiotic susceptibility testing. Diagnostics, 9(2), 49.
- Clinical and Laboratory Standards Institute (CLSI). (2021). Performance standards for antimicrobial disc susceptibility tests. CLSI document M100-S17, 17th Informational Supplement
- Humphries, R., Bobenchik, A. M., Hindler, J. A., & Schuetz, A.N. (2021). Overview of changes to the clinical and laboratory standards institute performance standards for antimicrobial susceptibility testing, M100. Journal of clinical microbiology, 59(12), 10-1128.
- Taiwo, S. S., Onile, B. A., & Akanbi II, A. A. (2004). Methicillin-resistant Staphylococcus aureus (MRSA) isolates in Ilorin, Nigeria. African Journal of Clinical and Experimental Microbiology, 5(2), 189-197.
- Cockerill, F. R., Wikler, M. A., & Alder, J. (2013). Twenty- Second Informational Supplement: Performance Standards for Antimicrobial Susceptibility Testing; M100–S21. Clinical and Laboratory Standards Institute, Wayne, PA.
- Zhang, S., Li, X., Wu, J., Coin, L., O’Brien, J., Hai, F., & Jiang, G. (2021). Molecular methods for pathogenic bacteria detection and recent advances in wastewater analysis. Water, 13(24), 3551.
- Jubeh, B., Breijyeh, Z., & Karaman, R. (2020). Resistance of gram-positive bacteria to current antibacterial agents and overcoming approaches. Molecules, 25(12), 2888.
- Algammal, A. M., Hetta, H. F., Elkelish, A., Alkhalifah, D.H. H., Hozzein, W. N., Batiha, G. E. S., ... & Mabrok, M.A. (2020). Methicillin-Resistant Staphylococcus aureus (MRSA): one health perspective approach to the bacterium epidemiology, virulence factors, antibiotic-resistance, and zoonotic impact. Infection and drug resistance, 3255-3265.
- Uslan, D. Z., Crane, S. J., Steckelberg, J. M., Cockerill,F. R., Sauver, J. L. S., Wilson, W. R., & Baddour, L. M. (2007). Age-and sex-associated trends in bloodstream infection: a population-based study in Olmsted County, Minnesota. Archives of internal medicine, 167(8), 834-839.
- Akoua-Koffi, C., Tia, H., Plo, J. K., Monemo, P., Cissé,A., Yao, C., ... & Becker, S. L. (2015). Epidemiology of community-onset bloodstream infections in Bouaké, central Côte d’Ivoire. New microbes and new infections, 7, 100-104.
- Rodríguezâ?Baño, J., Lópezâ?Prieto, M. D., Portillo, M. M., Retamar, P., Natera, C., Nuño, E., ... & SAEI/SAMPAC Bacteraemia Group. (2010). Epidemiology and clinical features of communityâ?acquired, healthcareâ?associated and nosocomial bloodstream infections in tertiaryâ?care and community hospitals. Clinical Microbiology and Infection, 16(9), 1408-1413.
- Sainfer, A., Berin, C., Jianfang, L., Elani, L. (2018). Prevalence and risk factors for bloodstream infections present on hospital admission. J Infect Prev 19(1):37–42.
- Stewart, J. H., Vajdic, C. M., Van Leeuwen, M. T., Amin, J., Webster, A. C., Chapman, J. R., ... & McCredie, M.R. (2009). The pattern of excess cancer in dialysis and transplantation. Nephrology Dialysis Transplantation, 24(10), 3225-3231.
- McKane, C. K., Marmarelis, M., Mendu, M. L., Moromizato, T., Gibbons, F. K., & Christopher, K. B. (2014). Diabetes mellitus and community-acquired bloodstream infections in the critically ill. Journal of critical care, 29(1), 70-76.
- Hugonnet, S., Sax, H., Eggimann, P., Chevrolet, J. C., & Pittet,D. (2004). Nosocomial bloodstream infection and clinical sepsis. Emerging infectious diseases, 10(1), 76.
- Rello, J., Ochagavia, A., Sabanes, E., Roque, M., Mariscal, D., Reynaga, E., & Valles, J. (2000). Evaluation of outcome of intravenous catheter-related infections in critically ill patients. American journal of respiratory and critical care medicine, 162(3), 1027-1030.
- Majumder, M. A. A., Rahman, S., Cohall, D., Bharatha, A., Singh, K., Haque, M., & Gittens-St Hilaire, M. (2020). Antimicrobial stewardship: fighting antimicrobial resistance and protecting global public health. Infection and drug resistance, 4713-4738.
- Veeraraghavan, B., Jesudason, M. R., Prakasah, J. A. J.(2018). Antimicrobial susceptibility profiles of gram-negative bacteria causing infections collected across India during 2014–2016: Study for monitoring antimicrobial resistance trend report. Indian journal of medical microbiology, 36(1), 32-36.
- Gautam, V., Thakur, A., Sharma, M., Singh, A., Bansal, S., Sharma, A., ... & Ray, P. (2019). Molecular characterization of extended-spectrum β-lactamases among clinical isolates of Escherichia coli & Klebsiella pneumoniae: a multi-centric study from tertiary care hospitals in India. Indian Journal of Medical Research, 149(2), 208-215.
- Bora, A., Hazarika, N. K., Shukla, S. K., Prasad, K. N., Sarma,J. B., & Ahmed, G. (2014). Prevalence of blaTEM, blaSHV and blaCTX-M genes in clinical isolates of Escherichia coli and Klebsiella pneumoniae from Northeast India. Indian journal of Pathology and Microbiology, 57(2), 249-254.
- Roy, S., Gaind, R., Chellani, H., Mohanty, S., Datta, S., Singh, A. K., & Basu, S. (2013). Neonatal septicaemia caused by diverse clones of Klebsiella pneumoniae & Escherichia coli harbouring blaCTX-M-15. Indian Journal of Medical Research, 137(4), 791-799.
- Clinical and Laboratory Standards Institute (CLSI). (2021). Performance standards for antimicrobial disc susceptibility tests. CLSI document M100-S17, 17th Informational Supplement.
- Karkada, U. H., Adamic, L. A., Kahn, J. M., & Iwashyna, T.J. (2011). Limiting the spread of highly resistant hospital- acquired microorganisms via critical care transfers: a simulation study. Intensive care medicine, 37, 1633-1640.
- Prasada, S., Bhat, A., Bhat, S., Shenoy Mulki, S., & Tulasidas,S. (2019). Changing antibiotic susceptibility pattern in uropathogenic Escherichia coli over a period of 5 years in a tertiary care center. Infection and Drug Resistance, 1439- 1443.
- Zhang, Z., Sun, Z., & Tian, L. (2022). Antimicrobial resistance among pathogens causing bloodstream infections: a multicenter surveillance report over 20 years (1998– 2017). Infection and Drug Resistance, 249-260.
Appendices
Appendix A: Ethical Consent from Nigeria Air-force Medical Centre, Onikan Lagos State.




